

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC136507.18 - phase: 1 /pseudo
(305 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
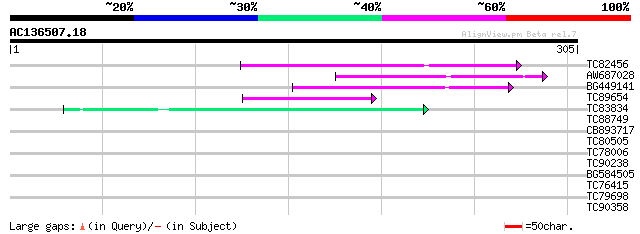
Score E
Sequences producing significant alignments: (bits) Value
TC82456 weakly similar to GP|18057159|gb|AAL58182.1 putative far... 62 3e-10
AW687028 weakly similar to GP|9909176|dbj| hypothetical protein~... 62 4e-10
BG449141 similar to GP|15983442|gb At1g10240/F14N23_12 {Arabidop... 57 9e-09
TC89654 weakly similar to PIR|T05644|T05644 hypothetical protein... 42 3e-04
TC83834 similar to PIR|G96565|G96565 F6D8.26 [imported] - Arabid... 42 4e-04
TC88749 40 0.001
CB893717 similar to SP|O04933|SPS Sucrose-phosphate synthase 2 (... 30 1.2
TC80505 29 2.0
TC78006 similar to GP|9758963|dbj|BAB09406.1 gb|AAF19567.1~gene_... 28 3.4
TC90238 similar to PIR|T07126|T07126 magnesium chelatase (EC 4.9... 28 5.7
BG584505 similar to GP|22831651|gb| CG32780-PA {Drosophila melan... 27 7.5
TC76415 similar to PIR|T05479|T05479 hypothetical protein T8O5.1... 27 7.5
TC79698 similar to GP|10176933|dbj|BAB10177. gene_id:MTG10.18~un... 27 9.8
TC90358 similar to PIR|D96799|D96799 hypothetical protein F22K20... 27 9.8
>TC82456 weakly similar to GP|18057159|gb|AAL58182.1 putative far-red
impaired response protein {Oryza sativa}, partial (7%)
Length = 1244
Score = 62.0 bits (149), Expect = 3e-10
Identities = 34/152 (22%), Positives = 69/152 (45%), Gaps = 1/152 (0%)
Frame = +2
Query: 125 RGDKKPLQYLISKLEEHKYTYYSRTQLESN-TIEDIFWAHPTSIKLFNNFPTVLVMDSTY 183
RGD + ++++ +Y ++ I+++FWA + F V+ +D+TY
Sbjct: 710 RGDVDAIHSFFHRMQKQNSQFYCAMDMDDKRNIQNLFWADARCRAAYEYFGEVITLDTTY 889
Query: 184 KTNIYRMPMFEVVGVTSTDLTYSVGFGFMTHEKEENFVWVLKMLRKLLSSKMNMPKVIVT 243
T+ Y +P+ VGV T +G +++ + W+ R L + P I+T
Sbjct: 890 LTSKYDLPLVPFVGVNHHGQTILLGCAILSNLDAKTLTWL--FTRWLECMHGHAPNGIIT 1063
Query: 244 DRDMSLMKAVAHVFPESYAMNCYFHVQANVKQ 275
+ D ++ A+ FP++ C +H+ V +
Sbjct: 1064EEDKAMKNAIEVAFPKARHRWCLWHIMKKVPE 1159
>AW687028 weakly similar to GP|9909176|dbj| hypothetical protein~similar to
Arabidopsis thaliana F20D10.300, partial (11%)
Length = 662
Score = 61.6 bits (148), Expect = 4e-10
Identities = 35/114 (30%), Positives = 56/114 (48%)
Frame = +1
Query: 176 VLVMDSTYKTNIYRMPMFEVVGVTSTDLTYSVGFGFMTHEKEENFVWVLKMLRKLLSSKM 235
VL D+TYK N Y P+ G T G + E E++ WVL + + +K
Sbjct: 4 VLAFDTTYKKNKYNYPLCIFSGCNHHSQTIIFGVALLEDETIESYKWVLNRFLECMENKF 183
Query: 236 NMPKVIVTDRDMSLMKAVAHVFPESYAMNCYFHVQANVKQRCVLDCKYPLGFKK 289
PK +VTD D S+ +A+ VFP++ C +H+ N ++ + + GF+K
Sbjct: 184 --PKAVVTDGDGSMREAIKQVFPDASHRLCAWHLHKNAQEN-IKKTPFLEGFRK 336
>BG449141 similar to GP|15983442|gb At1g10240/F14N23_12 {Arabidopsis
thaliana}, partial (33%)
Length = 690
Score = 57.0 bits (136), Expect = 9e-09
Identities = 30/119 (25%), Positives = 64/119 (53%)
Frame = +3
Query: 153 SNTIEDIFWAHPTSIKLFNNFPTVLVMDSTYKTNIYRMPMFEVVGVTSTDLTYSVGFGFM 212
+N +E+I W++ +SI+L++ F +V D+T++ + MP+ VG+ + + G +
Sbjct: 312 NNRLENIAWSYASSIQLYDIFGDAVVFDTTHRLTAFDMPLGLWVGINNYGMPCFFGCVLL 491
Query: 213 THEKEENFVWVLKMLRKLLSSKMNMPKVIVTDRDMSLMKAVAHVFPESYAMNCYFHVQA 271
E +F W +K ++ K P+ I+TD+++ L +A++ P + C + + A
Sbjct: 492 RDETVRSFSWAIKAFLGFMNGK--APQTILTDQNICLKEALSAEMPMTKHAFCIWMIVA 662
>TC89654 weakly similar to PIR|T05644|T05644 hypothetical protein F20D10.290
- Arabidopsis thaliana, partial (65%)
Length = 1358
Score = 42.0 bits (97), Expect = 3e-04
Identities = 23/72 (31%), Positives = 34/72 (46%)
Frame = +2
Query: 126 GDKKPLQYLISKLEEHKYTYYSRTQLESNTIEDIFWAHPTSIKLFNNFPTVLVMDSTYKT 185
G + L YL E+ +Y+ N +IFWA T ++ F ++ D+TYKT
Sbjct: 119 GGHQVLDYLKRMRAENHAFFYAVQSDVDNAGGNIFWADETCRTNYSYFGDTVIFDTTYKT 298
Query: 186 NIYRMPMFEVVG 197
N YR+P G
Sbjct: 299 NQYRVPFASFTG 334
>TC83834 similar to PIR|G96565|G96565 F6D8.26 [imported] - Arabidopsis
thaliana, partial (24%)
Length = 1032
Score = 41.6 bits (96), Expect = 4e-04
Identities = 35/198 (17%), Positives = 79/198 (39%), Gaps = 2/198 (1%)
Frame = +3
Query: 30 KCGCLFMISGYQSKQTKEWRLNILNGVHNHPMEPALEGHILAGRLKEDDKKIVRDLTKNK 89
+ GC MI + +++ WR+ + HNH + + I L D + K
Sbjct: 414 RTGCPAMIR-MKLVESQRWRICEVTLEHNHVLGAKIHKSIKKNSLPSSDAE-----GKTI 575
Query: 90 MLPRNILIHLK-NQRPHCMTNVKQVYNERQQIWKANRGDKKPLQYLISKLE-EHKYTYYS 147
+ ++I + N + + +++ +GD + + + +++ + +Y
Sbjct: 576 KVYHALVIDTEGNDNLNSNARDDRAFSKYSNKLNLRKGDTQAIYNFLCRMQLTNPNFFYL 755
Query: 148 RTQLESNTIEDIFWAHPTSIKLFNNFPTVLVMDSTYKTNIYRMPMFEVVGVTSTDLTYSV 207
+ + + W S F V+ D+TY N Y +P+ +VG+ + +
Sbjct: 756 MDFNDEGHLRNALWVDAKSRAACGYFSDVIYFDNTYLVNKYEIPLVALVGINHHGQSVLL 935
Query: 208 GFGFMTHEKEENFVWVLK 225
G G + E E++ W+ +
Sbjct: 936 GCGLLAGEIIESYKWLFR 989
>TC88749
Length = 1221
Score = 39.7 bits (91), Expect = 0.001
Identities = 18/29 (62%), Positives = 20/29 (68%)
Frame = +2
Query: 1 NPMFRLVCERSGSHIVPKKKPKHAGTGSR 29
NPM L+CERSG H V KK+ KH T SR
Sbjct: 641 NPMLELMCERSGDHKVSKKRLKHEAT*SR 727
>CB893717 similar to SP|O04933|SPS Sucrose-phosphate synthase 2 (EC 2.4.1.14)
(UDP-glucose-fructose- phosphate glucosyltransferase
2)., partial (22%)
Length = 843
Score = 30.0 bits (66), Expect = 1.2
Identities = 18/41 (43%), Positives = 28/41 (67%), Gaps = 1/41 (2%)
Frame = -3
Query: 63 PALEGHILAGRLKEDDK-KIVRDLTKNKMLPRNILIHLKNQ 102
P LEGH+L +LK D K ++V+DL K+ + NI I L+++
Sbjct: 127 PPLEGHVLQPQLK*DQKAQLVQDL*KHPLSLPNIGICLEHR 5
>TC80505
Length = 675
Score = 29.3 bits (64), Expect = 2.0
Identities = 21/63 (33%), Positives = 27/63 (42%)
Frame = +1
Query: 152 ESNTIEDIFWAHPTSIKLFNNFPTVLVMDSTYKTNIYRMPMFEVVGVTSTDLTYSVGFGF 211
ESN F H ++ + NFPT ++S K NIY F V + VGFG
Sbjct: 316 ESNNFSFSFLFHK*KLQKYTNFPTPFSINSPPKPNIYFFLSFLV*DL-------RVGFGT 474
Query: 212 MTH 214
H
Sbjct: 475 CHH 483
>TC78006 similar to GP|9758963|dbj|BAB09406.1
gb|AAF19567.1~gene_id:MYH9.6~similar to unknown protein
{Arabidopsis thaliana}, partial (48%)
Length = 1808
Score = 28.5 bits (62), Expect = 3.4
Identities = 17/51 (33%), Positives = 26/51 (50%), Gaps = 2/51 (3%)
Frame = -2
Query: 135 ISKLEEHKYTYYS-RTQLESNTIEDIFWAHPTSIKLFNNFPTVL-VMDSTY 183
++K + KY Y + NT EDI++ +P KLF + + MDS Y
Sbjct: 1771 VNKNHKKKYVYSEGKKSRVMNTFEDIYYPNPPQHKLFLHLRKIA*TMDSLY 1619
>TC90238 similar to PIR|T07126|T07126 magnesium chelatase (EC 4.99.1.-)
chain chlH - soybean, partial (15%)
Length = 844
Score = 27.7 bits (60), Expect = 5.7
Identities = 10/40 (25%), Positives = 22/40 (55%)
Frame = -1
Query: 167 IKLFNNFPTVLVMDSTYKTNIYRMPMFEVVGVTSTDLTYS 206
+ +N +++M+ Y N+Y P+F + +T +L +S
Sbjct: 160 LSFYNKEKVLVMMERWYLRNLYTFPLFFQICLTKVELCFS 41
>BG584505 similar to GP|22831651|gb| CG32780-PA {Drosophila melanogaster},
partial (1%)
Length = 822
Score = 27.3 bits (59), Expect = 7.5
Identities = 21/86 (24%), Positives = 38/86 (43%)
Frame = +2
Query: 73 RLKEDDKKIVRDLTKNKMLPRNILIHLKNQRPHCMTNVKQVYNERQQIWKANRGDKKPLQ 132
R ED +K+ + K + R I ++QR + + + E++ I K +PLQ
Sbjct: 2 RKSEDREKVGLKQYQKKKMERQRQI-ARDQRE--LARQRNIKIEKEAIEKEKNNKLRPLQ 172
Query: 133 YLISKLEEHKYTYYSRTQLESNTIED 158
+ K+E+ Q++ IED
Sbjct: 173 FQKKKIEDDTEAKLRPLQIQKQKIED 250
>TC76415 similar to PIR|T05479|T05479 hypothetical protein T8O5.180 -
Arabidopsis thaliana, partial (30%)
Length = 1114
Score = 27.3 bits (59), Expect = 7.5
Identities = 9/19 (47%), Positives = 14/19 (73%)
Frame = +2
Query: 35 FMISGYQSKQTKEWRLNIL 53
F+ + SK+ KEWR+NI+
Sbjct: 188 FVFEAFDSKKVKEWRINIV 244
>TC79698 similar to GP|10176933|dbj|BAB10177. gene_id:MTG10.18~unknown
protein {Arabidopsis thaliana}, partial (44%)
Length = 681
Score = 26.9 bits (58), Expect = 9.8
Identities = 17/59 (28%), Positives = 27/59 (44%), Gaps = 4/59 (6%)
Frame = -1
Query: 152 ESNTIEDIFWAHPTSIKLFNNFPTVLVMDS----TYKTNIYRMPMFEVVGVTSTDLTYS 206
++ T E+I W I L+N T L D T+ NIY + F + ++L +S
Sbjct: 411 DNKTRENIIWIMDMWIPLYNKRVTTLFTDCVECFTFSHNIYLITCFVITKAIPSNL*FS 235
>TC90358 similar to PIR|D96799|D96799 hypothetical protein F22K20.13
[imported] - Arabidopsis thaliana, partial (13%)
Length = 846
Score = 26.9 bits (58), Expect = 9.8
Identities = 12/36 (33%), Positives = 20/36 (55%)
Frame = +3
Query: 85 LTKNKMLPRNILIHLKNQRPHCMTNVKQVYNERQQI 120
+ +NKM+ IL+HL Q+ + V+ N + QI
Sbjct: 570 MERNKMISTKILVHLYLQKNEGQSKVRNPSNRKTQI 677
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.321 0.136 0.412
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 10,406,335
Number of Sequences: 36976
Number of extensions: 145453
Number of successful extensions: 769
Number of sequences better than 10.0: 28
Number of HSP's better than 10.0 without gapping: 766
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 769
length of query: 305
length of database: 9,014,727
effective HSP length: 96
effective length of query: 209
effective length of database: 5,465,031
effective search space: 1142191479
effective search space used: 1142191479
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 58 (26.9 bits)
Medicago: description of AC136507.18


