

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC136507.1 - phase: 0
(236 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
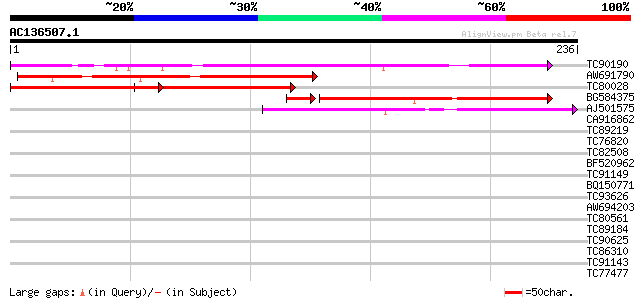
Score E
Sequences producing significant alignments: (bits) Value
TC90190 similar to GP|3426041|gb|AAC32240.1| unknown protein {Ar... 159 1e-39
AW691790 similar to GP|18491203|gb| unknown protein {Arabidopsis... 142 1e-34
TC80028 132 2e-31
BG584375 similar to GP|6498442|dbj ESTs AU081301(E20138) C99280(... 117 5e-29
AJ501575 homologue to GP|18491203|gb| unknown protein {Arabidops... 86 1e-17
CA916862 GP|12805231|gb Similar to mesenchyme homeobox 2 {Mus mu... 35 0.019
TC89219 weakly similar to PIR|C96656|C96656 unknown protein 438... 35 0.025
TC76820 weakly similar to GP|18175688|gb|AAL59911.1 unknown prot... 34 0.043
TC82508 similar to PIR|T50527|T50527 hypothetical protein T27I15... 33 0.095
BF520962 weakly similar to PIR|C96656|C96 unknown protein 43836... 32 0.16
TC91149 similar to GP|15081723|gb|AAK82516.1 T10F18_70/T10F18_70... 32 0.21
BQ150771 GP|4028979|emb glutamate permease {synthetic construct}... 31 0.36
TC93626 weakly similar to PIR|G84776|G84776 hypothetical protein... 30 0.62
AW694203 weakly similar to PIR|C96656|C96 unknown protein 43836... 30 0.62
TC80561 weakly similar to GP|10177856|dbj|BAB11208. phloem-speci... 30 0.81
TC89184 30 0.81
TC90625 weakly similar to GP|10177856|dbj|BAB11208. phloem-speci... 29 1.8
TC86310 similar to GP|16930469|gb|AAL31920.1 At2g26190/T1D16.17 ... 28 3.1
TC91143 similar to PIR|T47557|T47557 hypothetical protein F8J2.1... 28 3.1
TC77477 homologue to GP|5305740|gb|AAD41796.1| precursor monofun... 28 3.1
>TC90190 similar to GP|3426041|gb|AAC32240.1| unknown protein {Arabidopsis
thaliana}, partial (60%)
Length = 1626
Score = 159 bits (401), Expect = 1e-39
Identities = 101/241 (41%), Positives = 135/241 (55%), Gaps = 15/241 (6%)
Frame = +2
Query: 1 MLLYFITCVSFFLFLKPLLLLKPLPSWTNEVRLISLLFHKDFC--LKSFC--KLFRKTLV 56
M I+ VSF + L + KPL S N +L+S FC L SF +L +T+
Sbjct: 275 MFYILISFVSFVMILSKSFINKPLSS--NVKKLVS----NGFCGKLLSFLASRLKTRTIF 436
Query: 57 SRMSLV----------KKTLVSKVENVEEEVDSLQQNLSVLDLPELVLECILEKLPPSSL 106
S K LVSKVE EEE ++ S+LDL +L L+CILEKL PS L
Sbjct: 437 LPFSFFQFSLGMQLQNKSCLVSKVEYREEE----EEKASLLDLHDLPLDCILEKLSPSEL 604
Query: 107 CQMAGVCHSLRERCVSDHLWERHMKQKWGRVIGSVAYREWKWHVATKK-NIGSLRHGKPR 165
C +A VC SLRE+C SD+LWE+HMK KWG+++G AYR+WK +VA+K S +H +
Sbjct: 605 CNVAQVCKSLREKCRSDYLWEKHMKMKWGKILGDSAYRQWKCYVASKSIEKNSNQHKNQK 784
Query: 166 SLIKFMSLYWPFSWMRMKVDAAYSNSANKQNSYLPPDSFMTWYLALETGNFSFPAQVYNR 225
+++ F+ + PF W++ K + K P DS YL+LE G F FPAQVYNR
Sbjct: 785 TVLAFLHAFLPFFWIKSK--------SEKYIKSRPDDSIAALYLSLENGKFWFPAQVYNR 940
Query: 226 E 226
E
Sbjct: 941 E 943
>AW691790 similar to GP|18491203|gb| unknown protein {Arabidopsis thaliana},
partial (15%)
Length = 456
Score = 142 bits (358), Expect = 1e-34
Identities = 76/135 (56%), Positives = 96/135 (70%), Gaps = 10/135 (7%)
Frame = +3
Query: 4 YFITCVSFFLFLK--PLLLLKPLPSWTNEVRLISLLFHKDFCLKSFCKLFRK-------- 53
+ ITC+SFF FL L LK PSWT+E + + H+ LK+F KLF+
Sbjct: 75 FIITCISFFFFLNLNSLSFLKSFPSWTSE----NEMKHQVLSLKTFSKLFKNHFWIGLLC 242
Query: 54 TLVSRMSLVKKTLVSKVENVEEEVDSLQQNLSVLDLPELVLECILEKLPPSSLCQMAGVC 113
+ +RMS+V+K+L S+VENVEE ++ LS+LDLPEL+LECILEKLPPSSLCQMAG+C
Sbjct: 243 EIHTRMSMVRKSLSSRVENVEE----IEDKLSLLDLPELILECILEKLPPSSLCQMAGIC 410
Query: 114 HSLRERCVSDHLWER 128
HSLRERCVSD+ WER
Sbjct: 411 HSLRERCVSDYFWER 455
>TC80028
Length = 1120
Score = 132 bits (331), Expect = 2e-31
Identities = 64/64 (100%), Positives = 64/64 (100%)
Frame = +2
Query: 1 MLLYFITCVSFFLFLKPLLLLKPLPSWTNEVRLISLLFHKDFCLKSFCKLFRKTLVSRMS 60
MLLYFITCVSFFLFLKPLLLLKPLPSWTNEVRLISLLFHKDFCLKSFCKLFRKTLVSRMS
Sbjct: 761 MLLYFITCVSFFLFLKPLLLLKPLPSWTNEVRLISLLFHKDFCLKSFCKLFRKTLVSRMS 940
Query: 61 LVKK 64
LVKK
Sbjct: 941 LVKK 952
Score = 109 bits (273), Expect = 8e-25
Identities = 56/67 (83%), Positives = 59/67 (87%)
Frame = +3
Query: 53 KTLVSRMSLVKKTLVSKVENVEEEVDSLQQNLSVLDLPELVLECILEKLPPSSLCQMAGV 112
K L L +KTLVSKVENVEEEVDSLQQNLSVLDLP LVLECILEKLPPSSLCQMAGV
Sbjct: 918 KHLFQECLL*RKTLVSKVENVEEEVDSLQQNLSVLDLPXLVLECILEKLPPSSLCQMAGV 1097
Query: 113 CHSLRER 119
CHSL+E+
Sbjct: 1098CHSLKEK 1118
>BG584375 similar to GP|6498442|dbj ESTs AU081301(E20138) C99280(E10593)
correspond to a region of the predicted gene.~Similar
to, partial (39%)
Length = 765
Score = 117 bits (292), Expect(2) = 5e-29
Identities = 56/98 (57%), Positives = 67/98 (68%), Gaps = 1/98 (1%)
Frame = +3
Query: 130 MKQKWGRVIGSVAYREWKWHVATKKNIGSLRHGKPRSL-IKFMSLYWPFSWMRMKVDAAY 188
MK+KWG VIG AYREWKWHVA+ +GSL+HGK L +K SL WPF M++KV
Sbjct: 42 MKKKWGGVIGQAAYREWKWHVASNMGVGSLKHGKQSGLWMKIFSLLWPFQGMKLKV--GD 215
Query: 189 SNSANKQNSYLPPDSFMTWYLALETGNFSFPAQVYNRE 226
N + S LP ++ M WYLA+ETG F FPAQVYNRE
Sbjct: 216 GNDSCNHRSSLPVEAVMNWYLAIETGRFWFPAQVYNRE 329
Score = 27.7 bits (60), Expect(2) = 5e-29
Identities = 10/12 (83%), Positives = 11/12 (91%)
Frame = +1
Query: 116 LRERCVSDHLWE 127
LRERCVSD+ WE
Sbjct: 1 LRERCVSDYFWE 36
>AJ501575 homologue to GP|18491203|gb| unknown protein {Arabidopsis
thaliana}, partial (3%)
Length = 461
Score = 85.9 bits (211), Expect = 1e-17
Identities = 48/133 (36%), Positives = 72/133 (54%), Gaps = 2/133 (1%)
Frame = +1
Query: 106 LCQMAGVCHSLRERCVSDHLWERHMKQKWGRVIGSVAYREWKWHVATKKN--IGSLRHGK 163
LC++A VC LR R +D+LWE+ + Q+WG ++G +AY EW+WH A N I L +
Sbjct: 34 LCKVAQVCSCLRNRSKNDYLWEKQVNQRWGELLGDIAYHEWQWHTAKMVNEEILLLHQNQ 213
Query: 164 PRSLIKFMSLYWPFSWMRMKVDAAYSNSANKQNSYLPPDSFMTWYLALETGNFSFPAQVY 223
S F + WP + +Y + S L M+ ++ALETG F FPAQVY
Sbjct: 214 SGSCGTFTGV-WPSLHLH-----SYLENFRDMISLLRNCPKMSLFIALETGQFWFPAQVY 375
Query: 224 NREVTNSLCVLSY 236
+ ++ ++SY
Sbjct: 376 KKALSCYDALISY 414
>CA916862 GP|12805231|gb Similar to mesenchyme homeobox 2 {Mus musculus},
partial (5%)
Length = 727
Score = 35.4 bits (80), Expect = 0.019
Identities = 16/43 (37%), Positives = 27/43 (62%)
Frame = +2
Query: 75 EEVDSLQQNLSVLDLPELVLECILEKLPPSSLCQMAGVCHSLR 117
EEV + +N+ +LD+P +L I+ KLP SL + + VC+ +
Sbjct: 50 EEVGFVDENIQILDIPIHILFDIMLKLPAVSLIRCSSVCYKFK 178
>TC89219 weakly similar to PIR|C96656|C96656 unknown protein 43836-42510
[imported] - Arabidopsis thaliana, partial (46%)
Length = 764
Score = 35.0 bits (79), Expect = 0.025
Identities = 21/79 (26%), Positives = 36/79 (44%)
Frame = +3
Query: 49 KLFRKTLVSRMSLVKKTLVSKVENVEEEVDSLQQNLSVLDLPELVLECILEKLPPSSLCQ 108
++F ++ S M + S E EE +S + D+PE + IL L P +C+
Sbjct: 162 QIFVFSINSFMGASISSTASIREKNNEESNSSSSKTGLGDIPESCISSILMHLDPHDICK 341
Query: 109 MAGVCHSLRERCVSDHLWE 127
+A V + +D +WE
Sbjct: 342 LARVNRAFHRASSADFVWE 398
>TC76820 weakly similar to GP|18175688|gb|AAL59911.1 unknown protein
{Arabidopsis thaliana}, partial (67%)
Length = 1388
Score = 34.3 bits (77), Expect = 0.043
Identities = 18/69 (26%), Positives = 33/69 (47%)
Frame = +2
Query: 88 DLPELVLECILEKLPPSSLCQMAGVCHSLRERCVSDHLWERHMKQKWGRVIGSVAYREWK 147
D+PE + I+ +L P +C++A V + D++WE + + ++ + E K
Sbjct: 194 DIPESCISSIMMRLDPQEICKLARVNKTFHRASSDDYVWESKLPPSYKFLVKKILGEE-K 370
Query: 148 WHVATKKNI 156
TKK I
Sbjct: 371 LCSMTKKEI 397
>TC82508 similar to PIR|T50527|T50527 hypothetical protein T27I15_150 -
Arabidopsis thaliana, partial (71%)
Length = 1761
Score = 33.1 bits (74), Expect = 0.095
Identities = 13/52 (25%), Positives = 28/52 (53%)
Frame = +2
Query: 88 DLPELVLECILEKLPPSSLCQMAGVCHSLRERCVSDHLWERHMKQKWGRVIG 139
D+PE + +L L P +C++A + + R+ +D +WE + + ++G
Sbjct: 716 DIPESCVALVLMYLDPPDICKLARLNRAFRDASFADFVWESKLPLNYEFIMG 871
>BF520962 weakly similar to PIR|C96656|C96 unknown protein 43836-42510
[imported] - Arabidopsis thaliana, partial (58%)
Length = 681
Score = 32.3 bits (72), Expect = 0.16
Identities = 14/63 (22%), Positives = 32/63 (50%)
Frame = +3
Query: 79 SLQQNLSVLDLPELVLECILEKLPPSSLCQMAGVCHSLRERCVSDHLWERHMKQKWGRVI 138
S N S+ +LP+ + I+E + P +C++A + + ++D +WE + + +I
Sbjct: 150 SSSPNPSLGELPQSCIAGIMEYMDPPQICKLASLNRAFHGASLADFVWESKLPSNYHVII 329
Query: 139 GSV 141
+
Sbjct: 330 QKI 338
>TC91149 similar to GP|15081723|gb|AAK82516.1 T10F18_70/T10F18_70
{Arabidopsis thaliana}, partial (14%)
Length = 895
Score = 32.0 bits (71), Expect = 0.21
Identities = 16/51 (31%), Positives = 25/51 (48%)
Frame = +1
Query: 85 SVLDLPELVLECILEKLPPSSLCQMAGVCHSLRERCVSDHLWERHMKQKWG 135
S+ DLP +++ IL L P L ++ V LR H W+ ++WG
Sbjct: 505 SITDLPPVLISEILNCLDPKELGIVSCVSLILRSLASEHHAWKEFYCERWG 657
>BQ150771 GP|4028979|emb glutamate permease {synthetic construct}, partial
(3%)
Length = 1234
Score = 31.2 bits (69), Expect = 0.36
Identities = 38/151 (25%), Positives = 61/151 (40%), Gaps = 12/151 (7%)
Frame = -1
Query: 96 CILEKLPPSSLCQMAGVCHSLR-ERCVSDHLWERHMKQKWGRVIGSVAYREWKWHVATKK 154
C+L LP + G C R RCV LW + + + S+ + V +
Sbjct: 856 CLLRALPLTG-----GHCRRARGTRCV---LWVWYFSSHFLSALASLRVQS-ACVVLVSR 704
Query: 155 NIGSLRHGKPRSLIKFMSLYWPFSWM--RMKVDAAYSNSANKQ---------NSYLPPDS 203
+ R + F+S+ P S + R+ A YS++ + NS+LPP S
Sbjct: 703 RASRSDVSRLRVVAYFLSVSPPASAVCARLASSACYSSTLFARAYCCLCLIYNSHLPPPS 524
Query: 204 FMTWYLALETGNFSFPAQVYNREVTNSLCVL 234
+ +Y +L F FP +V +CVL
Sbjct: 523 HLAFYPSLSFYTFRFPVRVSWPRAVCFVCVL 431
>TC93626 weakly similar to PIR|G84776|G84776 hypothetical protein At2g36090
[imported] - Arabidopsis thaliana, partial (25%)
Length = 805
Score = 30.4 bits (67), Expect = 0.62
Identities = 15/45 (33%), Positives = 18/45 (39%)
Frame = +3
Query: 90 PELVLECILEKLPPSSLCQMAGVCHSLRERCVSDHLWERHMKQKW 134
P+++ IL L SSL A C L DHLW W
Sbjct: 63 PDIIQTHILTYLDGSSLASAASTCSQLNTLSSHDHLWTNICHSTW 197
>AW694203 weakly similar to PIR|C96656|C96 unknown protein 43836-42510
[imported] - Arabidopsis thaliana, partial (52%)
Length = 621
Score = 30.4 bits (67), Expect = 0.62
Identities = 12/54 (22%), Positives = 27/54 (49%)
Frame = +1
Query: 88 DLPELVLECILEKLPPSSLCQMAGVCHSLRERCVSDHLWERHMKQKWGRVIGSV 141
+LPE + I+ + P +CQ+A + + R +D +WE + + ++ +
Sbjct: 169 ELPESCVASIIGYMDPPQICQLATLNRAFRAASSADFVWESKLPPNYHLLLAKI 330
>TC80561 weakly similar to GP|10177856|dbj|BAB11208. phloem-specific
lectin-like protein {Arabidopsis thaliana}, partial
(52%)
Length = 1436
Score = 30.0 bits (66), Expect = 0.81
Identities = 13/58 (22%), Positives = 29/58 (49%)
Frame = +3
Query: 81 QQNLSVLDLPELVLECILEKLPPSSLCQMAGVCHSLRERCVSDHLWERHMKQKWGRVI 138
++ + DLPE +E +L P + +++ V + R SD +W++ + + +I
Sbjct: 147 EEGVEFHDLPEGCIENVLSFTTPRDVARLSLVSSTFRSAAESDSVWDKFLPSDYHSII 320
>TC89184
Length = 847
Score = 30.0 bits (66), Expect = 0.81
Identities = 22/66 (33%), Positives = 27/66 (40%), Gaps = 1/66 (1%)
Frame = +3
Query: 71 ENVEEEVDSLQQNLSVLDLPELVLECILEKLPPSSLCQMAGVCHSLRERCVSD-HLWERH 129
+NV EE S + S D PE V CIL L P + A C D LW
Sbjct: 189 KNVNEEFSS--SSPSFTDFPEDVQLCILSFLCPIEIASFACTSKQFGTLCSDDAKLWLSL 362
Query: 130 MKQKWG 135
++WG
Sbjct: 363 CDRRWG 380
>TC90625 weakly similar to GP|10177856|dbj|BAB11208. phloem-specific
lectin-like protein {Arabidopsis thaliana}, partial (9%)
Length = 892
Score = 28.9 bits (63), Expect = 1.8
Identities = 20/92 (21%), Positives = 39/92 (41%), Gaps = 1/92 (1%)
Frame = +2
Query: 40 KDFCLKSFCKLFRKTLVSRMSLVKKTLVS-KVENVEEEVDSLQQNLSVLDLPELVLECIL 98
++ C ++C + V +S + T++ K E + ++ L PE + IL
Sbjct: 110 RERCKTNWCFETKSHKVKFLSCTRSTVMDMKTEEAKSMIEVL---------PEGCIAKIL 262
Query: 99 EKLPPSSLCQMAGVCHSLRERCVSDHLWERHM 130
P C+++ VC SD +W+R +
Sbjct: 263 SHTAPVDSCRLSLVCKGFCSAAKSDTVWDRFL 358
>TC86310 similar to GP|16930469|gb|AAL31920.1 At2g26190/T1D16.17
{Arabidopsis thaliana}, partial (21%)
Length = 951
Score = 28.1 bits (61), Expect = 3.1
Identities = 11/27 (40%), Positives = 16/27 (58%)
Frame = -3
Query: 18 LLLLKPLPSWTNEVRLISLLFHKDFCL 44
LLLLK + W E+ + ++H D CL
Sbjct: 319 LLLLKIIEVWVEEINELRWVYHSDSCL 239
>TC91143 similar to PIR|T47557|T47557 hypothetical protein F8J2.170 -
Arabidopsis thaliana, partial (60%)
Length = 669
Score = 28.1 bits (61), Expect = 3.1
Identities = 12/54 (22%), Positives = 26/54 (47%)
Frame = +1
Query: 85 SVLDLPELVLECILEKLPPSSLCQMAGVCHSLRERCVSDHLWERHMKQKWGRVI 138
S+ D+PE + + L P +C +A + + R +D +WE + + ++
Sbjct: 145 SLGDIPENCVARVFLHLTPPEICNLARLNRAFRGAASADSVWESKLPSNYQHLL 306
>TC77477 homologue to GP|5305740|gb|AAD41796.1| precursor monofunctional
aspartokinase {Glycine max}, partial (84%)
Length = 2339
Score = 28.1 bits (61), Expect = 3.1
Identities = 15/45 (33%), Positives = 23/45 (50%), Gaps = 3/45 (6%)
Frame = +3
Query: 15 LKPLLLLKPLPSWTNEVRLISLL---FHKDFCLKSFCKLFRKTLV 56
+KP LLL PL +W + R +L F K +K C + K ++
Sbjct: 219 VKPNLLLPPLATWQQQQRTTNLFGYEFPKKAAVKVTCNVLEKHVI 353
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.325 0.137 0.436
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 9,956,338
Number of Sequences: 36976
Number of extensions: 168926
Number of successful extensions: 1442
Number of sequences better than 10.0: 56
Number of HSP's better than 10.0 without gapping: 1424
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1433
length of query: 236
length of database: 9,014,727
effective HSP length: 93
effective length of query: 143
effective length of database: 5,575,959
effective search space: 797362137
effective search space used: 797362137
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 57 (26.6 bits)
Medicago: description of AC136507.1


