

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC136472.16 - phase: 0
(212 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
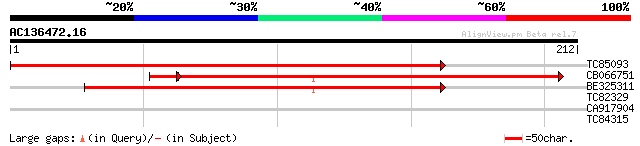
TC85093 similar to PIR|F86344|F86344 hypothetical protein AAF806... 315 8e-87
CB066751 similar to PIR|F86344|F86 hypothetical protein AAF80647... 194 1e-51
BE325311 similar to PIR|F86344|F86 hypothetical protein AAF80647... 196 6e-51
TC82329 similar to SP|Q01371|WC1_NEUCR White collar 1 protein (W... 28 3.4
CA917904 homologue to GP|19717681|gb| arsenite transport subunit... 27 4.4
TC84315 similar to GP|11876272|emb|CAC18917. unnamed protein pro... 27 7.5
>TC85093 similar to PIR|F86344|F86344 hypothetical protein AAF80647.1
[imported] - Arabidopsis thaliana, partial (59%)
Length = 548
Score = 315 bits (807), Expect = 8e-87
Identities = 161/163 (98%), Positives = 162/163 (98%)
Frame = +3
Query: 1 MAEIQHHHAATLARSESKVYNMDVEKKGEEENDEKDYTQRAQWLRAAVLGANDGLLSTAS 60
MAEIQHHHAATLARSESKVY+MDVEKKGEEENDEKDYTQRAQWLRAAVLGANDGLLSTAS
Sbjct: 30 MAEIQHHHAATLARSESKVYSMDVEKKGEEENDEKDYTQRAQWLRAAVLGANDGLLSTAS 209
Query: 61 LMMGVGAVTKDVKTMILTGIAGLVAGACSMAIGEFVSVYSQYDIEFAQMKRQGNISQKDK 120
LMMGVGAVTKDVKTMILTGIAGLVAGACSMAIGEFVSVYSQYDIEFAQMKRQGNISQKDK
Sbjct: 210 LMMGVGAVTKDVKTMILTGIAGLVAGACSMAIGEFVSVYSQYDIEFAQMKRQGNISQKDK 389
Query: 121 LPNPYYAAFASAIAFAVGAFVPLLGAAFVKDYKVRLGVVVGVV 163
LPNPYYAAFASAIAFAVGAFVPLLGAAFVKDYKVRLGV VGVV
Sbjct: 390 LPNPYYAAFASAIAFAVGAFVPLLGAAFVKDYKVRLGVXVGVV 518
>CB066751 similar to PIR|F86344|F86 hypothetical protein AAF80647.1
[imported] - Arabidopsis thaliana, partial (76%)
Length = 777
Score = 194 bits (494), Expect(2) = 1e-51
Identities = 101/151 (66%), Positives = 118/151 (77%), Gaps = 7/151 (4%)
Frame = -1
Query: 64 GVGAVTKDVKTMILTGIAGLVAGACSMAIGEFVSVYSQYDIEFAQMKRQ-------GNIS 116
GVGAV +D+K MILTG AGLVAGACSMAIGEFVSVYSQ DIE AQMKR+ +
Sbjct: 744 GVGAVKQDIKAMILTGFAGLVAGACSMAIGEFVSVYSQLDIEVAQMKREKERGREINDDE 565
Query: 117 QKDKLPNPYYAAFASAIAFAVGAFVPLLGAAFVKDYKVRLGVVVGVVSLALFGFGLLSAV 176
+K+ PNP AA ASA+AF++GA VPLL A F++DYKVR+GVVV VS AL FG L AV
Sbjct: 564 EKESFPNPLQAAAASALAFSIGAMVPLLAAFFIRDYKVRVGVVVAAVSFALLVFGWLGAV 385
Query: 177 LGKAPLVKSSLRVLIGGWLAMSLTFGLTKLV 207
LGKAP+ +S LRVL GGW+AMS+TFGLTKL+
Sbjct: 384 LGKAPIFRSVLRVLFGGWMAMSITFGLTKLI 292
Score = 25.4 bits (54), Expect(2) = 1e-51
Identities = 11/12 (91%), Positives = 12/12 (99%)
Frame = -2
Query: 53 DGLLSTASLMMG 64
DGL+STASLMMG
Sbjct: 776 DGLVSTASLMMG 741
>BE325311 similar to PIR|F86344|F86 hypothetical protein AAF80647.1
[imported] - Arabidopsis thaliana, partial (60%)
Length = 589
Score = 196 bits (498), Expect = 6e-51
Identities = 100/140 (71%), Positives = 118/140 (83%), Gaps = 5/140 (3%)
Frame = +2
Query: 29 EEENDEKDYTQRAQWLRAAVLGANDGLLSTASLMMGVGAVTKDVKTMILTGIAGLVAGAC 88
E E+DE DY++R+QWLRAAVLGANDGL+STASLMMGVGAV +D+K MIL+G AGLVAGAC
Sbjct: 149 EIESDEFDYSKRSQWLRAAVLGANDGLVSTASLMMGVGAVKQDIKAMILSGFAGLVAGAC 328
Query: 89 SMAIGEFVSVYSQYDIEFAQMKRQ-----GNISQKDKLPNPYYAAFASAIAFAVGAFVPL 143
SMAIGEFVSVYSQ DIE AQ+KR NI +K+ LPNP AA ASA+AF++GA VPL
Sbjct: 329 SMAIGEFVSVYSQLDIEIAQLKRDNIERGNNIEEKESLPNPLQAAAASALAFSIGAMVPL 508
Query: 144 LGAAFVKDYKVRLGVVVGVV 163
L A+F+KDYKVR+GVV+G V
Sbjct: 509 LAASFIKDYKVRVGVVLGAV 568
>TC82329 similar to SP|Q01371|WC1_NEUCR White collar 1 protein (WC1).
{Neurospora crassa}, partial (12%)
Length = 590
Score = 27.7 bits (60), Expect = 3.4
Identities = 22/72 (30%), Positives = 33/72 (45%), Gaps = 7/72 (9%)
Frame = +2
Query: 6 HHHAATLA-RSESKVYNMDVEKKGEEENDEKDYTQRAQWLRAAVLGANDGL------LST 58
H HA T A R +V++ +KG+E D + +Q R GAND L S
Sbjct: 128 HKHAGTHATRKPGRVWSR--HRKGQEGQDSNMQARGSQPQRTGAPGANDSLPGGRCRRSK 301
Query: 59 ASLMMGVGAVTK 70
A +++G +K
Sbjct: 302 AHVLVGTDEASK 337
>CA917904 homologue to GP|19717681|gb| arsenite transport subunit B
{Dictyostelium discoideum}, partial (2%)
Length = 750
Score = 27.3 bits (59), Expect = 4.4
Identities = 10/29 (34%), Positives = 18/29 (61%)
Frame = +3
Query: 6 HHHAATLARSESKVYNMDVEKKGEEENDE 34
HHH + +++ + ++ + KGEEE DE
Sbjct: 240 HHHHRIIKQNKKERIHIKIVDKGEEEKDE 326
>TC84315 similar to GP|11876272|emb|CAC18917. unnamed protein product
{Glycine max}, partial (49%)
Length = 811
Score = 26.6 bits (57), Expect = 7.5
Identities = 33/132 (25%), Positives = 50/132 (37%), Gaps = 2/132 (1%)
Frame = +2
Query: 56 LSTASLMMGVGAVTKDVKTMILTGIAGLVAGACSMAIGEFVSVYSQYDIEFAQMKRQGNI 115
LS S+ + D+ + TG+ V A M I V + D+E
Sbjct: 476 LSIISVSLLAAEKLSDISPLFFTGVLEAVVAALFMNI-YIVGLNQLSDVEI--------- 625
Query: 116 SQKDKLPNPYYAAFASAIAFAVGAFVPLLGAAFVKDYKVRLGVVVGVVSLALFG--FGLL 173
DK+ PY + +FA GA + + + V G+ L + G FGL
Sbjct: 626 ---DKINKPYLPLASGEYSFATGAIIGVSSSF----------*VFGLPGL*VHGHCFGLF 766
Query: 174 SAVLGKAPLVKS 185
S+VL L+ S
Sbjct: 767 SSVLC*GRLIPS 802
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.321 0.135 0.380
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 5,125,521
Number of Sequences: 36976
Number of extensions: 50685
Number of successful extensions: 268
Number of sequences better than 10.0: 12
Number of HSP's better than 10.0 without gapping: 265
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 265
length of query: 212
length of database: 9,014,727
effective HSP length: 92
effective length of query: 120
effective length of database: 5,612,935
effective search space: 673552200
effective search space used: 673552200
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 56 (26.2 bits)
Medicago: description of AC136472.16


