

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC136451.2 - phase: 0 /pseudo
(355 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
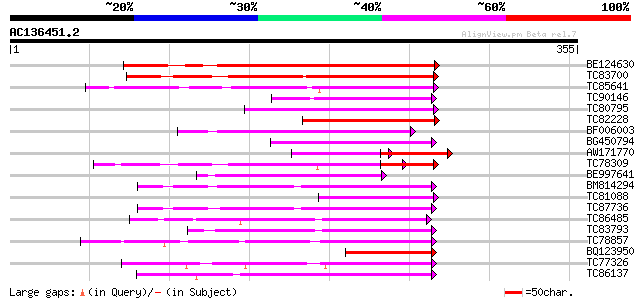
Score E
Sequences producing significant alignments: (bits) Value
BE124630 weakly similar to GP|14209594|db cytochrome P450-like p... 191 3e-49
TC83700 similar to GP|9587211|gb|AAF89209.1| cytochrome P450 {Vi... 129 2e-30
TC85641 weakly similar to GP|8777295|dbj|BAA96885.1 cytochrome P... 89 3e-18
TC90146 weakly similar to SP|Q42569|C901_ARATH Cytochrome P450 9... 88 4e-18
TC80795 weakly similar to GP|23495769|dbj|BAC19980. 5-alpha-taxa... 83 1e-16
TC82228 similar to GP|10177005|dbj|BAB10255. cytochrome P450 {Ar... 82 3e-16
BF006003 similar to GP|11934677|gb| ent-kaurenoic acid oxidase {... 80 1e-15
BG450794 similar to SP|O23051|C883 Cytochrome P450 88A3 (EC 1.14... 77 8e-15
AW171770 similar to GP|20259299|gb| putative cytochrome P450 pro... 47 3e-13
TC78309 similar to GP|20259299|gb|AAM14385.1 putative cytochrome... 72 3e-13
BE997641 weakly similar to PIR|T02739|T02 probable cytochrome P4... 69 4e-12
BM814294 weakly similar to GP|6002279|emb cytochrome P450 monoox... 68 5e-12
TC81088 weakly similar to GP|23495770|dbj|BAC19981. 5-alpha-taxa... 66 2e-11
TC87736 weakly similar to GP|6002279|emb|CAB56741.1 cytochrome P... 65 5e-11
TC86485 similar to GP|23315211|gb|AAN20392.1 Sequence 405 from p... 63 2e-10
TC83793 weakly similar to GP|6002279|emb|CAB56741.1 cytochrome P... 60 1e-09
TC78857 weakly similar to SP|O81974|C7D8_SOYBN Cytochrome P450 7... 57 1e-08
BQ123950 similar to SP|O23051|C883 Cytochrome P450 88A3 (EC 1.14... 55 4e-08
TC77326 similar to GP|12004682|gb|AAG44132.1 cytochrome P450 {Pi... 54 1e-07
TC86137 weakly similar to PIR|A96673|A96673 probable cytochrome ... 53 2e-07
>BE124630 weakly similar to GP|14209594|db cytochrome P450-like protein
{Oryza sativa (japonica cultivar-group)}, partial (33%)
Length = 608
Score = 191 bits (486), Expect = 3e-49
Identities = 101/199 (50%), Positives = 136/199 (67%), Gaps = 1/199 (0%)
Frame = +1
Query: 72 LYKSLKAKERMMKMVGRIVE-ERMKLIMDNNAEDDEKVGVVNDVVDALLRDKGELSQSSS 130
LY+SL+AK++M+K V + V+ +R K I + V DVVD LL D +
Sbjct: 16 LYQSLQAKKKMVK*VRKTVQAKRNKGIFE----------VPRDVVDVLLND--------T 141
Query: 131 SSNLMVEMISQNIIEFMIPGEETLPTAMTLALKFLTDYPLALSKLMEENMELKKQKTNCP 190
S L ++I+ NII+ MIPGE+++P MTLA K+L++ P AL +L EN++LKK K
Sbjct: 142 SEKLTDDLIADNIIDMMIPGEDSVPVLMTLATKYLSECPPALQQLTVENIKLKKLKDQLG 321
Query: 191 DGYTWTDYMSLPFTQNVISETLRMANIVNGIWRKAVKDVEIKGYLIPKDWGVMASLTSVH 250
W DY+SLPFTQ VI+ETLRM NI+NG+ RKA+KDVEIKGYLIP+ W V A+ SVH
Sbjct: 322 KPLCWNDYLSLPFTQKVITETLRMGNIINGVMRKALKDVEIKGYLIPQGWCVFANFRSVH 501
Query: 251 MDSKNYENPYKFDPWRWEI 269
+D KNYE PY+F+PWRW++
Sbjct: 502 LDEKNYECPYQFNPWRWQV 558
>TC83700 similar to GP|9587211|gb|AAF89209.1| cytochrome P450 {Vigna
radiata}, partial (42%)
Length = 658
Score = 129 bits (324), Expect = 2e-30
Identities = 69/195 (35%), Positives = 119/195 (60%)
Frame = +3
Query: 74 KSLKAKERMMKMVGRIVEERMKLIMDNNAEDDEKVGVVNDVVDALLRDKGELSQSSSSSN 133
+++KA+ ++ + + +V +R K E++ + ND++ ALL + S
Sbjct: 54 RAIKARTKVAEALTLVVRQRRK-------ENEIGIQKKNDMLGALLASGEQFSN------ 194
Query: 134 LMVEMISQNIIEFMIPGEETLPTAMTLALKFLTDYPLALSKLMEENMELKKQKTNCPDGY 193
E I ++ ++ G ET T MTLA+KFLT+ PLAL++L EE+ +++ ++
Sbjct: 195 ---EEIVDFMLALLVAGYETTSTIMTLAIKFLTETPLALAQLKEEHEQIRA-RSEPEAAL 362
Query: 194 TWTDYMSLPFTQNVISETLRMANIVNGIWRKAVKDVEIKGYLIPKDWGVMASLTSVHMDS 253
WTDY S+ FTQ V++ETLR+ANI+ I+R+ D++IKGY IPK V+AS +VH++
Sbjct: 363 EWTDYKSMTFTQCVVNETLRVANIIGAIFRRTTTDIDIKGYTIPKGMKVIASFRAVHLNP 542
Query: 254 KNYENPYKFDPWRWE 268
+++++ F+PWRW+
Sbjct: 543 EHFKDARTFNPWRWQ 587
>TC85641 weakly similar to GP|8777295|dbj|BAA96885.1 cytochrome P450-like
{Arabidopsis thaliana}, partial (71%)
Length = 1941
Score = 88.6 bits (218), Expect = 3e-18
Identities = 60/223 (26%), Positives = 108/223 (47%), Gaps = 2/223 (0%)
Frame = +1
Query: 48 ETLDFIASGYSSCPVTFMEKRKSILYKSLKAKERMMKMVGRIVEERMKLIMDNNAEDDEK 107
+ IA+G S P+ + + K++KA + K + +I+++R + + A +
Sbjct: 715 DPFQLIAAGIISLPI---DLPGTPFNKAIKASNFIRKELIKIIKQRRVDLAEGTASPTQ- 882
Query: 108 VGVVNDVVDALLRDKGELSQSSSSSNLMVEMISQNIIEFMIPGEETLPTAMTLALKFLTD 167
D++ +L E +S + N I+ I+ +I G +T A T +K+L +
Sbjct: 883 -----DILSHMLLTSDENGKSMNELN-----IADKILGLLIGGHDTASVACTFLVKYLGE 1032
Query: 168 YPLALSKLMEENMELKKQKTNCPDG--YTWTDYMSLPFTQNVISETLRMANIVNGIWRKA 225
P K+ +E ME+ K K P G W D + ++ NV E +R++ + G +R+A
Sbjct: 1033LPHIYDKVYQEQMEIAKSK---PAGELLNWDDLKKMKYSWNVACEVMRLSPPLQGGFREA 1203
Query: 226 VKDVEIKGYLIPKDWGVMASLTSVHMDSKNYENPYKFDPWRWE 268
+ D G+ IPK W + S S H +++ + P KFDP R+E
Sbjct: 1204ITDFMFNGFSIPKGWKLYWSANSTHKNAECFPMPEKFDPTRFE 1332
Score = 40.4 bits (93), Expect = 0.001
Identities = 23/77 (29%), Positives = 45/77 (57%), Gaps = 9/77 (11%)
Frame = +1
Query: 8 YVSIIVLL-----MSLWFVKKEKNNIKLRNGIKIPKGNSGWPLLGETLDFIASGYSSCPV 62
Y+S+++L +SL+F+ K ++ + +P G G+P++GE+L+F+++G+ P
Sbjct: 127 YLSLLLLFVTFISLSLFFIF-----YKQKSPLNLPPGKMGYPIIGESLEFLSTGWKGHPE 291
Query: 63 TF----MEKRKSILYKS 75
F M K S L+K+
Sbjct: 292 KFIFDRMRKYSSELFKT 342
>TC90146 weakly similar to SP|Q42569|C901_ARATH Cytochrome P450 90A1 (EC
1.14.-.-). [Mouse-ear cress] {Arabidopsis thaliana},
partial (7%)
Length = 777
Score = 88.2 bits (217), Expect = 4e-18
Identities = 45/103 (43%), Positives = 60/103 (57%)
Frame = +2
Query: 165 LTDYPLALSKLMEENMELKKQKTNCPDGYTWTDYMSLPFTQNVISETLRMANIVNGIWRK 224
L YP+ L KL EENM +K D T D +L +T V+ E +RMANI +RK
Sbjct: 2 LAKYPIVLKKLREENMAFRKGSPE--DFITPKDVSNLKYTNKVVEEVIRMANIAACAFRK 175
Query: 225 AVKDVEIKGYLIPKDWGVMASLTSVHMDSKNYENPYKFDPWRW 267
+V+ KGY IPK W V+ L +H DSKN+++P F+P RW
Sbjct: 176 VDTEVDYKGYKIPKGWNVILLLRYLHTDSKNFKDPMNFNPDRW 304
>TC80795 weakly similar to GP|23495769|dbj|BAC19980.
5-alpha-taxadienol-10-beta-hydroxylase-like protein
{Oryza sativa (japonica cultivar-group)}, partial (12%)
Length = 796
Score = 83.2 bits (204), Expect = 1e-16
Identities = 43/121 (35%), Positives = 68/121 (55%)
Frame = +3
Query: 148 IPGEETLPTAMTLALKFLTDYPLALSKLMEENMELKKQKTNCPDGYTWTDYMSLPFTQNV 207
+ +T T M+L + L+ +K++EE ME+ KQ+ +G TW + + +T V
Sbjct: 3 VASHDTSATLMSLMIWKLSRDQEVYNKVLEEQMEILKQREANEEGLTWGEIQKMKYTWRV 182
Query: 208 ISETLRMANIVNGIWRKAVKDVEIKGYLIPKDWGVMASLTSVHMDSKNYENPYKFDPWRW 267
E +RM + G +RKA+KD +GY IPK W V + + H + +ENPYKF+P R+
Sbjct: 183 AQELMRMIPPLFGAFRKALKDTCYQGYDIPKGWQVYWASSGTHENKDIFENPYKFEPSRF 362
Query: 268 E 268
E
Sbjct: 363 E 365
>TC82228 similar to GP|10177005|dbj|BAB10255. cytochrome P450 {Arabidopsis
thaliana}, partial (30%)
Length = 1003
Score = 82.0 bits (201), Expect = 3e-16
Identities = 37/86 (43%), Positives = 56/86 (65%)
Frame = +3
Query: 184 KQKTNCPDGYTWTDYMSLPFTQNVISETLRMANIVNGIWRKAVKDVEIKGYLIPKDWGVM 243
KQ+ G W D ++P T VI ETLR+A+I++ +R+A +DVE +GYLIPK W V+
Sbjct: 33 KQENGEEIGLNWEDTKNMPITSRVIQETLRVASILSFTFREATEDVEYQGYLIPKGWKVL 212
Query: 244 ASLTSVHMDSKNYENPYKFDPWRWEI 269
++H +N++ P KFDP R+E+
Sbjct: 213 PLFRNIHHSPENFKEPEKFDPSRFEV 290
>BF006003 similar to GP|11934677|gb| ent-kaurenoic acid oxidase {Cucurbita
maxima}, partial (14%)
Length = 441
Score = 80.1 bits (196), Expect = 1e-15
Identities = 50/149 (33%), Positives = 74/149 (49%)
Frame = +2
Query: 106 EKVGVVNDVVDALLRDKGELSQSSSSSNLMVEMISQNIIEFMIPGEETLPTAMTLALKFL 165
+ VG D++D LL K E + L + IS +I + G E+ T + ++ L
Sbjct: 5 QHVGEKKDLMDILLEIKDE-----NGRKLEDQDISDLLIGLLFAGHESTATGIMWSVAHL 169
Query: 166 TDYPLALSKLMEENMELKKQKTNCPDGYTWTDYMSLPFTQNVISETLRMANIVNGIWRKA 225
T +P L K EE E+ K + + + + + VI E LR ANI I+R+A
Sbjct: 170 TQHPHILQKAKEEQEEILKIRPASQKRLSLNEVKQMIYLSYVIDEMLRFANIAFSIFREA 349
Query: 226 VKDVEIKGYLIPKDWGVMASLTSVHMDSK 254
DV I GYLIPK W V+ ++HMDS+
Sbjct: 350 TSDVNINGYLIPKGWRVLIWARAIHMDSE 436
>BG450794 similar to SP|O23051|C883 Cytochrome P450 88A3 (EC 1.14.-.-).
[Mouse-ear cress] {Arabidopsis thaliana}, partial (33%)
Length = 691
Score = 77.4 bits (189), Expect = 8e-15
Identities = 40/104 (38%), Positives = 56/104 (53%)
Frame = +1
Query: 164 FLTDYPLALSKLMEENMELKKQKTNCPDGYTWTDYMSLPFTQNVISETLRMANIVNGIWR 223
FL +P L K EE E+ K++ G T + + F VI ET+R+ ++R
Sbjct: 1 FLQKHPEYLQKAKEEQEEIIKRRPPTQKGLTLKEIRGMDFLYKVIDETMRVITFSLVVFR 180
Query: 224 KAVKDVEIKGYLIPKDWGVMASLTSVHMDSKNYENPYKFDPWRW 267
+A DV I GY IPK W V+ SVH+D + Y NP +F+P RW
Sbjct: 181 EAKSDVMINGYTIPKGWKVLTWFRSVHLDPEIYPNPKEFNPNRW 312
>AW171770 similar to GP|20259299|gb| putative cytochrome P450 protein
{Arabidopsis thaliana}, partial (16%)
Length = 539
Score = 47.4 bits (111), Expect(2) = 3e-13
Identities = 20/45 (44%), Positives = 28/45 (61%)
Frame = +2
Query: 233 GYLIPKDWGVMASLTSVHMDSKNYENPYKFDPWRWEIIYLHEAII 277
GYLIPK W V+ ++H N+++P KFDP R+E+I E I
Sbjct: 383 GYLIPKGWKVLPLFRNIHHSPNNFKDPEKFDPSRFEVIIEQELFI 517
Score = 45.1 bits (105), Expect(2) = 3e-13
Identities = 22/64 (34%), Positives = 37/64 (57%)
Frame = +1
Query: 177 EENMELKKQKTNCPDGYTWTDYMSLPFTQNVISETLRMANIVNGIWRKAVKDVEIKGYLI 236
+E++ K++ G W D + T VI ETLR+A+I++ +R+AV+DVE +G
Sbjct: 112 QESILKSKEENGEEKGLNWEDTKKMVITSRVIQETLRVASILSFTFREAVEDVEYQGQYF 291
Query: 237 PKDW 240
K +
Sbjct: 292 SKSY 303
>TC78309 similar to GP|20259299|gb|AAM14385.1 putative cytochrome P450
protein {Arabidopsis thaliana}, partial (89%)
Length = 2063
Score = 72.0 bits (175), Expect = 3e-13
Identities = 51/199 (25%), Positives = 98/199 (48%), Gaps = 2/199 (1%)
Frame = +1
Query: 53 IASGYSSCPVTFMEKRKSILYKSLKAKERMMKMVGRIVEERMKLIMDNNAEDDEKVGVVN 112
+ GY+S P+ ++ +K++KA++ + +++ +I+ R EK
Sbjct: 664 LEKGYNSMPINLPG---TLFHKAMKARKELAQILAQIISSRR-----------EKKEEYK 801
Query: 113 DVVDALLRDKGELSQSSSSSNLMVEMISQNIIEFMIPGEETLPTAMTLALKFLTDYPLAL 172
D++ + + +K LS E I+ N+I + +T + +T +K+L + AL
Sbjct: 802 DLLGSFMDEKSGLSD---------EQIANNVIGVIFATRDTTASVLTWIVKYLGENISAL 954
Query: 173 SKLMEENMELKKQKTNCPD--GYTWTDYMSLPFTQNVISETLRMANIVNGIWRKAVKDVE 230
++EE + K K + G W D + T VI ETLR+A+I++ +R+AV+DVE
Sbjct: 955 ESVIEEQESILKSKEENGEEKGLNWEDTKKMVITSRVIQETLRVASILSFTFREAVEDVE 1134
Query: 231 IKGYLIPKDWGVMASLTSV 249
+G K + S+T++
Sbjct: 1135YQGQYYTK---TVISITNI 1182
Score = 43.1 bits (100), Expect = 2e-04
Identities = 17/36 (47%), Positives = 24/36 (66%)
Frame = +3
Query: 233 GYLIPKDWGVMASLTSVHMDSKNYENPYKFDPWRWE 268
GYLIPK W V+ ++H N+++P KFDP R+E
Sbjct: 1251 GYLIPKGWKVLPLFRNIHHSPNNFKDPEKFDPSRFE 1358
Score = 34.7 bits (78), Expect = 0.058
Identities = 21/71 (29%), Positives = 36/71 (50%), Gaps = 3/71 (4%)
Frame = +1
Query: 8 YVSIIVLLMSLWFVKKEKNNIKLRNGIKIPKGNSGWPLLGETLDFIASGYSSCPVTFME- 66
+ SI+ +++ F+K + + R+ + +P G+ GWP +GET YS P F
Sbjct: 94 FTSILFIVLFRIFIK---SFVSKRHDLPLPPGSMGWPYIGETFQL----YSQDPNVFFAS 252
Query: 67 --KRKSILYKS 75
KR ++KS
Sbjct: 253 KIKRYGSMFKS 285
>BE997641 weakly similar to PIR|T02739|T02 probable cytochrome P450 At2g29090
[imported] - Arabidopsis thaliana, partial (21%)
Length = 399
Score = 68.6 bits (166), Expect = 4e-12
Identities = 40/119 (33%), Positives = 64/119 (53%)
Frame = +3
Query: 118 LLRDKGELSQSSSSSNLMVEMISQNIIEFMIPGEETLPTAMTLALKFLTDYPLALSKLME 177
LLR KG+ + L ++ NII + +T +A+T LK+L D L + +
Sbjct: 54 LLRGKGD----EKMNQLTDSQVADNIIGGIFAAHDTTTSALTWVLKYLHDNVNILETVTK 221
Query: 178 ENMELKKQKTNCPDGYTWTDYMSLPFTQNVISETLRMANIVNGIWRKAVKDVEIKGYLI 236
E E++ + G W D +PFT VI TLR A+I++ +R+AV+DV+++GY I
Sbjct: 222 EQEEIRSIASRESRGLAWEDTRQMPFTSRVIQYTLRSASILSFTFREAVRDVKLQGYSI 398
>BM814294 weakly similar to GP|6002279|emb cytochrome P450 monooxygenase
{Cicer arietinum}, partial (37%)
Length = 793
Score = 68.2 bits (165), Expect = 5e-12
Identities = 51/188 (27%), Positives = 92/188 (48%), Gaps = 1/188 (0%)
Frame = +1
Query: 81 RMMKMVGRIVEERMKLIMDNNAEDDEKVGVVNDVVDALLRDKGELSQSSSSSNLMVEMIS 140
++ + I+++R+KL +++ ND++D+LL E + L E I
Sbjct: 61 KLFNVFDNIIDQRLKL------REEDGFFTNNDMLDSLLDIPEE-----NRKELDREKIE 207
Query: 141 QNIIEFMIPGEETLPTAMTLALKFLTDYPLALSKLMEENMELKKQKTNCPDGYTWTDYMS 200
+ + ++ G +T + A+ L P +SK+ +E + + +D
Sbjct: 208 HLLHDLLVGGTDTTTYTLEWAMAELLHNPNIMSKVKKE----LEDTIGIGNPLEESDITR 375
Query: 201 LPFTQNVISETLRMANIVNGIW-RKAVKDVEIKGYLIPKDWGVMASLTSVHMDSKNYENP 259
LP+ Q VI ETLR+ I + RKA +DVE+ GY IPKD + ++ ++ D + ++NP
Sbjct: 376 LPYLQAVIKETLRLHPIAPLLLPRKAKEDVEVNGYTIPKDAQIFVNVWAIGRDPEVWDNP 555
Query: 260 YKFDPWRW 267
Y F P R+
Sbjct: 556 YLFSPERF 579
>TC81088 weakly similar to GP|23495770|dbj|BAC19981.
5-alpha-taxadienol-10-beta-hydroxylase-like protein
{Oryza sativa (japonica cultivar-group)}, partial (28%)
Length = 724
Score = 66.2 bits (160), Expect = 2e-11
Identities = 31/75 (41%), Positives = 42/75 (55%)
Frame = +3
Query: 194 TWTDYMSLPFTQNVISETLRMANIVNGIWRKAVKDVEIKGYLIPKDWGVMASLTSVHMDS 253
TW D + FT V ETLR + + G +RK D+E GY+IPK W + + HMD+
Sbjct: 48 TWEDLSKMKFTWRVAMETLRRFSPIFGGFRKTATDIEYGGYIIPKGWQIFWVTSMTHMDN 227
Query: 254 KNYENPYKFDPWRWE 268
+ P KFDP R+E
Sbjct: 228 NIFPEPSKFDPSRFE 272
>TC87736 weakly similar to GP|6002279|emb|CAB56741.1 cytochrome P450
monooxygenase {Cicer arietinum}, partial (55%)
Length = 1367
Score = 64.7 bits (156), Expect = 5e-11
Identities = 48/188 (25%), Positives = 92/188 (48%), Gaps = 1/188 (0%)
Frame = +3
Query: 81 RMMKMVGRIVEERMKLIMDNNAEDDEKVGVVNDVVDALLRDKGELSQSSSSSNLMVEMIS 140
++ + I+++++KL + + ND++D+LL ++ + L E I
Sbjct: 333 KLFGIFDNIIDQKLKL------REGDGFVTNNDMLDSLLAEENK-------KELDREKIQ 473
Query: 141 QNIIEFMIPGEETLPTAMTLALKFLTDYPLALSKLMEENMELKKQKTNCPDGYTWTDYMS 200
+ + ++ G +T + A+ L P +SK+ +E ++ + +D
Sbjct: 474 HLLHDLLVGGTDTTTYTLEWAMAELLHNPNVMSKVKKE----LEETIGIGNPIEESDVTR 641
Query: 201 LPFTQNVISETLRMANIVNGIW-RKAVKDVEIKGYLIPKDWGVMASLTSVHMDSKNYENP 259
LP+ Q +I ETLR+ I + RKA +DVE+ GYLIPK + ++ ++ D K ++NP
Sbjct: 642 LPYLQAIIKETLRLHPIAPLLLPRKAKEDVEVNGYLIPKGAQIFVNVWAIGRDPKVWDNP 821
Query: 260 YKFDPWRW 267
F P R+
Sbjct: 822 NLFSPERF 845
>TC86485 similar to GP|23315211|gb|AAN20392.1 Sequence 405 from patent US
6410718, partial (90%)
Length = 1927
Score = 63.2 bits (152), Expect = 2e-10
Identities = 43/193 (22%), Positives = 93/193 (47%), Gaps = 4/193 (2%)
Frame = +3
Query: 76 LKAKERMMKMVGRIVEERMKLIMDNNAEDDEKVGVVNDVVDALLRDKGELSQSSSSSNLM 135
+KA++ + + +I++E + + DE+ +V++++ A ++ +L+ S N
Sbjct: 861 VKARKELDSFIDKIIDEH---VQKKKSVVDEETDMVDELL-AFYSEEAKLNNESDDLNNS 1028
Query: 136 VEMISQNI----IEFMIPGEETLPTAMTLALKFLTDYPLALSKLMEENMELKKQKTNCPD 191
+++ NI ++ M G ET+ +A+ A+ L P L K+ +E E+ +
Sbjct: 1029 IKLTKDNIKAIIMDVMFGGTETVASAIEWAMAELMKSPEDLKKVQQELAEVVGLSRQVEE 1208
Query: 192 GYTWTDYMSLPFTQNVISETLRMANIVNGIWRKAVKDVEIKGYLIPKDWGVMASLTSVHM 251
+D+ L + + + ETLR+ + + + ++ + GY IPK VM + ++
Sbjct: 1209 ----SDFEKLTYLKCALKETLRLHPPIPLLLHETAEEATVNGYFIPKQARVMINAWAIGR 1376
Query: 252 DSKNYENPYKFDP 264
D+ +E P F P
Sbjct: 1377 DANCWEEPQSFKP 1415
>TC83793 weakly similar to GP|6002279|emb|CAB56741.1 cytochrome P450
monooxygenase {Cicer arietinum}, partial (48%)
Length = 923
Score = 60.5 bits (145), Expect = 1e-09
Identities = 42/157 (26%), Positives = 81/157 (50%), Gaps = 1/157 (0%)
Frame = +2
Query: 112 NDVVDALLRDKGELSQSSSSSNLMVEMISQNIIEFMIPGEETLPTAMTLALKFLTDYPLA 171
+D++DA+L D+ ++ + + I + ++ + G +T+ + + A+ L P
Sbjct: 170 SDMLDAMLDDE------KNAGEMYKDKIERLSVDLFVAGTDTVTSTLEWAMAELLHNPNI 331
Query: 172 LSKLMEENMELKKQKTNCPDGYTWTDYMSLPFTQNVISETLRMANIVNGIW-RKAVKDVE 230
+SK E ++ + + + +D LP+ Q +I ET R+ V + RKA D+E
Sbjct: 332 MSKAKSELNQIIGKGNSVEE----SDIGKLPYLQAIIKETFRLHPAVPLLLPRKAEIDLE 499
Query: 231 IKGYLIPKDWGVMASLTSVHMDSKNYENPYKFDPWRW 267
I GY +PK V+ ++ ++ DS +ENP +F P R+
Sbjct: 500 INGYKVPKGAQVLINVWAIGRDSNLWENPNEFLPERF 610
>TC78857 weakly similar to SP|O81974|C7D8_SOYBN Cytochrome P450 71D8 (EC
1.14.-.-) (P450 CP7). [Soybean] {Glycine max}, partial
(38%)
Length = 1724
Score = 56.6 bits (135), Expect = 1e-08
Identities = 56/225 (24%), Positives = 95/225 (41%), Gaps = 2/225 (0%)
Frame = +3
Query: 45 LLGETLDFIASGYSSCPVTFMEKRKSILYKSLKAKERMMKMVGRIVEERMK--LIMDNNA 102
LL ET S + S + FM L ++ + K + E+ +K L +N
Sbjct: 702 LLNETQAIFLSFFVSDYIPFMGWVDK-LTGAIARVDNTFKALDEFFEQVLKEHLNPNNRK 878
Query: 103 EDDEKVGVVNDVVDALLRDKGELSQSSSSSNLMVEMISQNIIEFMIPGEETLPTAMTLAL 162
+DDE+ D+VD LL K + S +N ++ + N++ + +T +
Sbjct: 879 KDDEE----KDIVDVLLELKNQGRLSIDLTNNHIKAVVMNLL---VAATDTSAATSVWVM 1037
Query: 163 KFLTDYPLALSKLMEENMELKKQKTNCPDGYTWTDYMSLPFTQNVISETLRMANIVNGIW 222
L P A+ K EE +KK+ + D + + VI ETLR +
Sbjct: 1038TGLIKNPRAMKKAQEEIRNIKKEFID------EDDIQKFVYFKAVIKETLRFYSPAPLAP 1199
Query: 223 RKAVKDVEIKGYLIPKDWGVMASLTSVHMDSKNYENPYKFDPWRW 267
R+ K + GY I V S+ S+H D + +++P +F P R+
Sbjct: 1200RETSKSFTLNGYKIEPKTSVFVSIWSIHRDPETWKDPDEFYPERF 1334
>BQ123950 similar to SP|O23051|C883 Cytochrome P450 88A3 (EC 1.14.-.-).
[Mouse-ear cress] {Arabidopsis thaliana}, partial (23%)
Length = 791
Score = 55.1 bits (131), Expect = 4e-08
Identities = 25/57 (43%), Positives = 35/57 (60%)
Frame = +2
Query: 211 TLRMANIVNGIWRKAVKDVEIKGYLIPKDWGVMASLTSVHMDSKNYENPYKFDPWRW 267
T+R+ ++R+A DV I GY IPK W V+ SVH+D + Y NP +F+P RW
Sbjct: 17 TIRVITFSLVVFREAKSDVMINGYTIPKGWKVLTWFRSVHLDPEIYPNPKEFNPNRW 187
>TC77326 similar to GP|12004682|gb|AAG44132.1 cytochrome P450 {Pisum
sativum}, complete
Length = 1864
Score = 53.5 bits (127), Expect = 1e-07
Identities = 52/208 (25%), Positives = 95/208 (45%), Gaps = 11/208 (5%)
Frame = +1
Query: 71 ILYKSLKAKERMMKMVGRIVEERMKLIMDNNAEDDEKVG--VVNDVVDALLRDKGELSQS 128
I + L+ + MK + + + M+ +++ + E + V V D+VD LL Q
Sbjct: 742 IHFLDLQGYVKRMKTLSKKFDRFMEHVLEEHIERRKNVKDYVAKDMVDVLL-------QL 900
Query: 129 SSSSNLMVEMISQNIIEF----MIPGEETLPTAMTLALKFLTDYPLALSKLMEE-NMELK 183
+ NL V++ + F + G E+ + A+ L P K EE + +
Sbjct: 901 AEDPNLEVKLERHGVKAFTQDLIAGGTESSAVTVEWAISELVRKPEIFKKATEELDRVIG 1080
Query: 184 KQKTNCPDGYTWT---DYMSLPFTQNVISETLRMANIVNGIW-RKAVKDVEIKGYLIPKD 239
K + W D +LP+ + ET+R+ + + R+A +D ++ GY IPK
Sbjct: 1081KDR--------WVEEKDIANLPYVYAIAKETMRLHPVAPFLVPREAREDCKVDGYDIPKG 1236
Query: 240 WGVMASLTSVHMDSKNYENPYKFDPWRW 267
V+ + ++ DS+ +ENPY+F P R+
Sbjct: 1237TIVLVNTWTIARDSEVWENPYEFMPERF 1320
>TC86137 weakly similar to PIR|A96673|A96673 probable cytochrome P450
F13O11.25 [imported] - Arabidopsis thaliana, partial
(69%)
Length = 1695
Score = 53.1 bits (126), Expect = 2e-07
Identities = 52/193 (26%), Positives = 86/193 (43%), Gaps = 5/193 (2%)
Frame = +3
Query: 80 ERMMKMVGRIVEERMKLIMDNNAEDDEKVGVVNDVV---DALLRDKGELSQSSSSSNLMV 136
E +K++ + + LI + K+ +N VV D LL + + S N MV
Sbjct: 777 EEFLKLLKDQEDVLLPLIRARKQVKESKLNNINTVVSYADTLLELEWPEEKRKLSENEMV 956
Query: 137 EMISQNIIEFMIPGEETLPTAMTLALKFLTDYPLALSKLMEENMELKKQKTNCP-DGYTW 195
+ S EF+ G +T T++ + + YP +L+EE E+ N +
Sbjct: 957 NLCS----EFLNGGTDTTSTSLQWIMANVVKYPEVQGRLVEEIREVMGGDENGEKEEVKE 1124
Query: 196 TDYMSLPFTQNVISETLRMANIVNGIWRKAVK-DVEIKGYLIPKDWGVMASLTSVHMDSK 254
D L + + V+ E LR AVK DV + GYL+PK+ V L + +D +
Sbjct: 1125EDLQKLRYLKCVVLEGLRRHPSGKFPLPHAVKEDVVLDGYLVPKNGTVNFLLAEMALDRR 1304
Query: 255 NYENPYKFDPWRW 267
+E+P +F P R+
Sbjct: 1305VWEDPLEFKPERF 1343
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.328 0.142 0.433
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 10,978,332
Number of Sequences: 36976
Number of extensions: 148413
Number of successful extensions: 1026
Number of sequences better than 10.0: 97
Number of HSP's better than 10.0 without gapping: 1001
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1012
length of query: 355
length of database: 9,014,727
effective HSP length: 97
effective length of query: 258
effective length of database: 5,428,055
effective search space: 1400438190
effective search space used: 1400438190
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.7 bits)
S2: 59 (27.3 bits)
Medicago: description of AC136451.2


