

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC136451.10 - phase: 0 /pseudo
(820 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
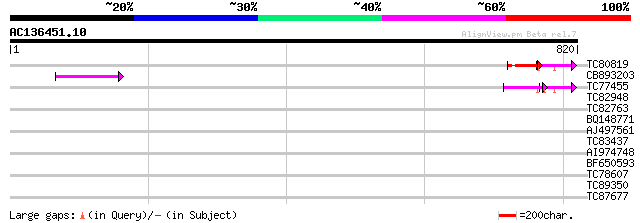
Score E
Sequences producing significant alignments: (bits) Value
TC80819 weakly similar to GP|10140689|gb|AAG13524.1 putative non... 62 9e-10
CB893203 55 1e-07
TC77455 similar to GP|22335695|dbj|BAC10549. nine-cis-epoxycarot... 50 3e-06
TC82948 39 0.011
TC82763 38 0.014
BQ148771 37 0.023
AJ497561 weakly similar to GP|10140689|gb putative non-LTR retro... 33 0.44
TC83437 weakly similar to PIR|D86384|D86384 unknown protein [imp... 27 0.59
AI974748 32 0.76
BF650593 32 0.76
TC78607 similar to GP|20466746|gb|AAM20690.1 putative protein {A... 30 3.7
TC89350 similar to GP|21213868|emb|CAD12767. LHY protein {Phaseo... 30 4.9
TC87677 29 6.4
>TC80819 weakly similar to GP|10140689|gb|AAG13524.1 putative non-LTR
retroelement reverse transcriptase {Oryza sativa
(japonica cultivar-group)}, partial (2%)
Length = 1262
Score = 62.0 bits (149), Expect = 9e-10
Identities = 34/57 (59%), Positives = 38/57 (66%), Gaps = 7/57 (12%)
Frame = +1
Query: 721 GLGVRRIREFNSALLGKWSWMVLVEKESLWYRVLVARYGEEGG-------RLLDGGR 770
GLGV FN +LLGKW W +LV+KE LW+RVL ARYGEEGG LL GGR
Sbjct: 31 GLGVGA---FNLSLLGKWCWRLLVDKEGLWHRVLKARYGEEGGGFVRVVDSLLCGGR 192
Score = 54.3 bits (129), Expect = 2e-07
Identities = 22/64 (34%), Positives = 34/64 (52%), Gaps = 6/64 (9%)
Frame = +3
Query: 762 GGRLLDGGRTASAWWRAIATLHSEA------WFGTNVKRSVGNGENIYFWSDVWVGGVSF 815
GG L +GGR +S WW+ I + WF N++ VG+G + +FW D W G V
Sbjct: 144 GGWLCEGGRQSSMWWKTICKVREGVGEGVGNWFEENIRMVVGDGRDAFFWYDTWAGDVPL 323
Query: 816 KERF 819
+ ++
Sbjct: 324 RLKY 335
>CB893203
Length = 800
Score = 54.7 bits (130), Expect = 1e-07
Identities = 42/98 (42%), Positives = 54/98 (54%)
Frame = -2
Query: 67 *KVVLDLA*LFSAGLVTRVAGSLPVNIVC**GKLGA*TVTIVEMLGKFSWL*KLCSRSVA 126
*K+V +A* S ++R SL N+ * GKLGA + VEMLG S L +C +
Sbjct: 793 *KLVY*MA*YDSKSSLSRSIRSLSYNVND*RGKLGASLASYVEMLG*SSGLSSICQG*ME 614
Query: 127 FLSGGGMGRICLERKIHVD*PSFERLAS*TLSKFTCKN 164
F+SG MGR+C KI RLAS*+ S+F C N
Sbjct: 613 FISGVKMGRLCA*GKIKNGQKQP*RLAS*SYSEFRCSN 500
>TC77455 similar to GP|22335695|dbj|BAC10549. nine-cis-epoxycarotenoid
dioxygenase1 {Pisum sativum}, partial (43%)
Length = 1865
Score = 50.1 bits (118), Expect = 3e-06
Identities = 32/76 (42%), Positives = 39/76 (51%), Gaps = 12/76 (15%)
Frame = -3
Query: 714 CLSKEVGGLGVRRIREFNSALLGKWSWMVLVEKESLWYRVLVARYGEEG----------G 763
CL + GGLGVR IR N +LL KW W +L ++ SLW VL YG G
Sbjct: 969 CLPRCKGGLGVRDIRLVNVSLLAKWWWRLLQDQSSLWKEVLEDIYGPRVSSRTMMVGRIG 790
Query: 764 RLL--DGGRTASAWWR 777
RL+ DGG+ W R
Sbjct: 789 RLMLQDGGKI**VWRR 742
Score = 44.7 bits (104), Expect = 1e-04
Identities = 21/60 (35%), Positives = 29/60 (48%), Gaps = 7/60 (11%)
Frame = -2
Query: 767 DGGRT----ASAWWRAIATLHSEA---WFGTNVKRSVGNGENIYFWSDVWVGGVSFKERF 819
DGG AS WW+ + +L WF + R VG+G + +FW D W V +E F
Sbjct: 808 DGGENWPAYASRWWKDLMSLEEVGRVRWFPRELIRKVGDGRSSFFWKDAWDSSVPLRESF 629
>TC82948
Length = 705
Score = 38.5 bits (88), Expect = 0.011
Identities = 18/48 (37%), Positives = 26/48 (53%)
Frame = +3
Query: 704 KITWVDWNYVCLSKEVGGLGVRRIREFNSALLGKWSWMVLVEKESLWY 751
K+ V W VC + G LG+R + + N AL K W +++ KE WY
Sbjct: 255 KVVKVSWEKVCRPIKEGSLGIRSLSKLNEALNLKLCWDMMISKEQ-WY 395
>TC82763
Length = 1376
Score = 38.1 bits (87), Expect = 0.014
Identities = 24/74 (32%), Positives = 40/74 (53%), Gaps = 1/74 (1%)
Frame = +1
Query: 710 WNYVCLSKEVGGLGVRRIREFNSALLGKWSWMVLVEKESLWYRVLVARYGEEGGRLLDG- 768
WN++C GG+ +R +R FN A+ SW + E S + VL +YG L+G
Sbjct: 244 WNWICRPIR-GGIRIRNLRFFNIAVR**-SWTLKTETNSFHFMVLFHKYGS----WLEGV 405
Query: 769 GRTASAWWRAIATL 782
++S WWR ++++
Sbjct: 406 NCSSSLWWRYLSSI 447
>BQ148771
Length = 680
Score = 37.4 bits (85), Expect = 0.023
Identities = 25/94 (26%), Positives = 40/94 (41%), Gaps = 2/94 (2%)
Frame = -3
Query: 694 FFLGGNDDHKKITWVDWNYVCLSKEVGGLGVRRIREFNSALLGKWSWMVLVEKESLWYRV 753
F G + ++ V W + K + GLG+RR+ N A + K W + SL V
Sbjct: 426 FVWGDTEVSRRYHAVGWETMSKPKTIYGLGLRRLDVMNKACIMKLGWSIYSGSNSLCTEV 247
Query: 754 LVARY--GEEGGRLLDGGRTASAWWRAIATLHSE 785
+ +Y E + T S+ W+A+ L E
Sbjct: 246 MRGKYQRSESLEEIFLEKPTDSSLWKALVKLWPE 145
>AJ497561 weakly similar to GP|10140689|gb putative non-LTR retroelement
reverse transcriptase {Oryza sativa (japonica
cultivar-group)}, partial (2%)
Length = 621
Score = 33.1 bits (74), Expect = 0.44
Identities = 17/44 (38%), Positives = 29/44 (65%)
Frame = +3
Query: 709 DWNYVCLSKEVGGLGVRRIREFNSALLGKWSWMVLVEKESLWYR 752
++N + K VGGLG++ +EFN ++LGK W +L + L+Y+
Sbjct: 300 EFNNLARHKSVGGLGLQN-QEFNISMLGKQRWKLL--SQPLYYQ 422
>TC83437 weakly similar to PIR|D86384|D86384 unknown protein [imported] -
Arabidopsis thaliana, partial (6%)
Length = 951
Score = 26.9 bits (58), Expect(2) = 0.59
Identities = 18/38 (47%), Positives = 20/38 (52%)
Frame = +3
Query: 419 RWYSGGK*GCG*CQEM*KGTSSFQSGFRKSL*FHRLDL 456
RWYS K*GCG* E + + GF K L RL L
Sbjct: 243 RWYSYSK*GCG*G*ETEERSFVV*GGF*KGLSLCRLGL 356
Score = 24.3 bits (51), Expect(2) = 0.59
Identities = 14/31 (45%), Positives = 18/31 (57%)
Frame = +1
Query: 358 YFHCSYS*GREPATSE*ISPDIFGG*YVQNS 388
+FH +S*G +P+ * S FGG V NS
Sbjct: 58 HFHSPHS*G*QPSAP**FSSYFFGGKSV*NS 150
>AI974748
Length = 261
Score = 32.3 bits (72), Expect = 0.76
Identities = 11/15 (73%), Positives = 14/15 (93%)
Frame = +1
Query: 796 VGNGENIYFWSDVWV 810
VGNGE+IYFW+D W+
Sbjct: 1 VGNGESIYFWTDRWL 45
>BF650593
Length = 486
Score = 32.3 bits (72), Expect = 0.76
Identities = 13/31 (41%), Positives = 17/31 (53%)
Frame = +1
Query: 749 LWYRVLVARYGEEGGRLLDGGRTASAWWRAI 779
LW+R LV +YG G + R S WW+ I
Sbjct: 88 LWFRALVNKYGLNRGSITIENRGVSLWWKDI 180
>TC78607 similar to GP|20466746|gb|AAM20690.1 putative protein {Arabidopsis
thaliana}, partial (55%)
Length = 1734
Score = 30.0 bits (66), Expect = 3.7
Identities = 12/29 (41%), Positives = 17/29 (58%), Gaps = 4/29 (13%)
Frame = +2
Query: 785 EAWF----GTNVKRSVGNGENIYFWSDVW 809
+AWF G +V V NG ++Y+W D W
Sbjct: 47 KAWFR*PHGASVS*KVSNGCSVYWWQDTW 133
>TC89350 similar to GP|21213868|emb|CAD12767. LHY protein {Phaseolus
vulgaris}, partial (13%)
Length = 752
Score = 29.6 bits (65), Expect = 4.9
Identities = 25/89 (28%), Positives = 41/89 (45%), Gaps = 2/89 (2%)
Frame = -1
Query: 485 GFGQWVPNRRVPFEEGFEARRPAFPFP-LSLGCRRFQYFDEVNVG*SVLPWVQGGFRCYY 543
G W P+ ++ G + RP FP+P L++ C+ FDE V PW++ CY+
Sbjct: 449 GVSSWDPSLQL----GCKPFRPCFPWPLLNIFCKV*TLFDES*VYLVQDPWLEIPPHCYF 282
Query: 544 -GCLTLTICG*YFNNS*KILGQCSGDACG 571
+ + G ++ K + GD CG
Sbjct: 281 VQMVGFAVPG----HNYKCISCSQGDLCG 207
>TC87677
Length = 682
Score = 29.3 bits (64), Expect = 6.4
Identities = 10/20 (50%), Positives = 15/20 (75%)
Frame = +3
Query: 740 WMVLVEKESLWYRVLVARYG 759
WM+++E +W+ VL ARYG
Sbjct: 264 WMIILESLIMWWDVLYARYG 323
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.361 0.164 0.647
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 29,793,753
Number of Sequences: 36976
Number of extensions: 509135
Number of successful extensions: 4698
Number of sequences better than 10.0: 26
Number of HSP's better than 10.0 without gapping: 1298
Number of HSP's successfully gapped in prelim test: 163
Number of HSP's that attempted gapping in prelim test: 3332
Number of HSP's gapped (non-prelim): 1616
length of query: 820
length of database: 9,014,727
effective HSP length: 104
effective length of query: 716
effective length of database: 5,169,223
effective search space: 3701163668
effective search space used: 3701163668
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 14 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 37 (21.9 bits)
S2: 62 (28.5 bits)
Medicago: description of AC136451.10


