

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC136449.11 - phase: 0 /pseudo
(982 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
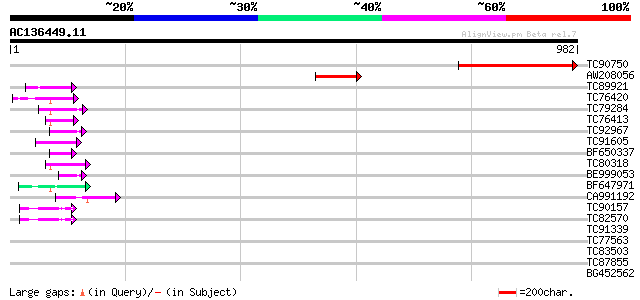
Score E
Sequences producing significant alignments: (bits) Value
TC90750 homologue to PIR|T05426|T05426 hypothetical protein F28A... 414 e-115
AW208056 homologue to PIR|T05426|T0 hypothetical protein F28A23.... 135 8e-32
TC89921 similar to GP|21594296|gb|AAM65991.1 unknown {Arabidopsi... 65 2e-10
TC76420 similar to PIR|T45715|T45715 hypothetical protein F1P2.1... 65 2e-10
TC79284 similar to GP|10178077|dbj|BAB11496. gene_id:K19B1.7~unk... 62 8e-10
TC76413 similar to GP|3461820|gb|AAC32914.1| unknown protein {Ar... 57 3e-08
TC92967 similar to GP|9972376|gb|AAG10626.1| Unknown protein {Ar... 56 8e-08
TC91605 similar to GP|15809927|gb|AAL06891.1 At3g09760/F8A24.19 ... 55 2e-07
BF650337 similar to GP|8885610|dbj| contains similarity to unkno... 49 9e-06
TC80318 similar to GP|21593635|gb|AAM65602.1 unknown {Arabidopsi... 48 2e-05
BE999053 similar to GP|9972376|gb|A Unknown protein {Arabidopsis... 46 6e-05
BF647971 similar to GP|21537278|gb unknown {Arabidopsis thaliana... 46 8e-05
CA991192 weakly similar to GP|21594296|gb unknown {Arabidopsis t... 43 5e-04
TC90157 similar to PIR|T51853|T51853 RING-H2 finger protein RHF1... 42 9e-04
TC82570 similar to PIR|T51853|T51853 RING-H2 finger protein RHF1... 42 9e-04
TC91339 homologue to PIR|B84609|B84609 hypothetical protein At2g... 40 0.004
TC77563 similar to GP|15450687|gb|AAK96615.1 AT5g59550/f2o15_210... 39 0.010
TC83503 weakly similar to PIR|T51851|T51851 RING-H2 finger prote... 39 0.013
TC87855 similar to GP|15081801|gb|AAK82555.1 AT5g41350/MYC6_6 {A... 39 0.013
BG452562 similar to GP|9293902|dbj contains similarity to RING z... 37 0.049
>TC90750 homologue to PIR|T05426|T05426 hypothetical protein F28A23.140 -
Arabidopsis thaliana, partial (15%)
Length = 880
Score = 414 bits (1063), Expect = e-115
Identities = 202/205 (98%), Positives = 202/205 (98%)
Frame = +3
Query: 778 LFELLVIVPMRVPVDESPVFLLYQDWALGLIFLKIWTRLVMLDHMMPLMDESWRVKFERV 837
LFELLVIVPMRVPVDESPVFLLYQDWALGLIFLKIWTRLVMLDHMMPLMDESWRVKFERV
Sbjct: 3 LFELLVIVPMRVPVDESPVFLLYQDWALGLIFLKIWTRLVMLDHMMPLMDESWRVKFERV 182
Query: 838 RDDGFSRLQGLWVLREIVLPIIMKLLTALCVPYVLARGVFPALGYPLVVNSAVYRFAWLG 897
RDDGFSRLQGLWVLREIVLPIIMKLLTALCVPYVLARGVFPALGYPLVVNSAVYRFAWLG
Sbjct: 183 RDDGFSRLQGLWVLREIVLPIIMKLLTALCVPYVLARGVFPALGYPLVVNSAVYRFAWLG 362
Query: 898 CLSFSFVCFCAKRFHVWFTNLHNSIRDDRYLIGRRLHNFGEHVEKANEAATSTGVQDAIL 957
CLSFSFVCFCAKRFHVWFTNLHNSIRDDRYLIGRR HNFGEHVEKANEAATSTGVQDAIL
Sbjct: 363 CLSFSFVCFCAKRFHVWFTNLHNSIRDDRYLIGRR*HNFGEHVEKANEAATSTGVQDAIL 542
Query: 958 LGPNINQQDRDADVGLRLRHINQQA 982
LGPNINQQDRDA VGL LRHINQQA
Sbjct: 543 LGPNINQQDRDAXVGLXLRHINQQA 617
>AW208056 homologue to PIR|T05426|T0 hypothetical protein F28A23.140 -
Arabidopsis thaliana, partial (12%)
Length = 411
Score = 135 bits (340), Expect = 8e-32
Identities = 69/80 (86%), Positives = 72/80 (89%)
Frame = +2
Query: 530 GIVYMLQISIFVSLLRGVLRNGVLYFLRDPADPNYNPSRDLIDNPVHKHARRVLLSVAVY 589
GIVYMLQISIFVSLLRGVLRNGVLYFLRDPADPNYNP RDLID+PVHKHARRVLLSVAVY
Sbjct: 8 GIVYMLQISIFVSLLRGVLRNGVLYFLRDPADPNYNPFRDLIDDPVHKHARRVLLSVAVY 187
Query: 590 GSLMVMLVILPR*LGIADGP 609
GSL+VMLV LP L + P
Sbjct: 188 GSLIVMLVFLPVKLAMRMAP 247
>TC89921 similar to GP|21594296|gb|AAM65991.1 unknown {Arabidopsis
thaliana}, partial (45%)
Length = 1649
Score = 64.7 bits (156), Expect = 2e-10
Identities = 30/88 (34%), Positives = 46/88 (52%)
Frame = +1
Query: 28 SSSSSPRGSKGKEIDAEAVATASTAPPSAKYDDDDEDEEDVCRICRNPGDADNPLRYPCA 87
S S + R + KE + T AP D++ +E VCRIC + D N + C
Sbjct: 628 SVSFNTRSEQDKEDTNDDQITP--APVEVTEDEEIPEEAAVCRICLDECDEGNTFKMECY 801
Query: 88 CSGSIKFVHQDCLLQWLNHSNARQCEVC 115
C G ++ VH++CL++WLN +CE+C
Sbjct: 802 CKGDLRLVHEECLIKWLNTKGTNKCEIC 885
>TC76420 similar to PIR|T45715|T45715 hypothetical protein F1P2.100 -
Arabidopsis thaliana, partial (67%)
Length = 1176
Score = 64.7 bits (156), Expect = 2e-10
Identities = 39/120 (32%), Positives = 57/120 (47%), Gaps = 5/120 (4%)
Frame = +1
Query: 5 NEPPPSLDGTPIAATTPSSSSPSSSSSSPRGSKGKEIDAEAVATASTAPPSAKYDDDDED 64
+ P +D P A PS SPS S ++ A+A ++ S + DD ++
Sbjct: 88 DHPVVPVDQQP--AHLPSDPSPSPSPAT----------ADAGSSGSGSAVDRDGDDCGDE 231
Query: 65 EEDV-----CRICRNPGDADNPLRYPCACSGSIKFVHQDCLLQWLNHSNARQCEVCKHPF 119
EE + CRIC+ N L PCACSGS+K+ H+ C+ W N CE+C P+
Sbjct: 232 EEPLIQMVECRICQEEDSVSN-LETPCACSGSLKYAHRKCVQHWCNEKGDITCEICHQPY 408
>TC79284 similar to GP|10178077|dbj|BAB11496. gene_id:K19B1.7~unknown
protein {Arabidopsis thaliana}, partial (56%)
Length = 1009
Score = 62.4 bits (150), Expect = 8e-10
Identities = 35/94 (37%), Positives = 44/94 (46%), Gaps = 9/94 (9%)
Frame = +3
Query: 50 STAPPSAKYDDDDEDEEDV---------CRICRNPGDADNPLRYPCACSGSIKFVHQDCL 100
S A PS DD+ ED CRIC+ DN L PCACSGS+K+ H+ C+
Sbjct: 183 SPAGPSGSSTADDDVGEDEPLLQMEMAECRICQEEDGIDN-LENPCACSGSLKYAHRKCV 359
Query: 101 LQWLNHSNARQCEVCKHPFSFSPVYAENAPARLP 134
+W N CE+C S+ P Y P P
Sbjct: 360 QRWCNEKGDTTCEICHK--SYEPDYTAPPPRPHP 455
>TC76413 similar to GP|3461820|gb|AAC32914.1| unknown protein {Arabidopsis
thaliana}, partial (54%)
Length = 1310
Score = 57.4 bits (137), Expect = 3e-08
Identities = 25/62 (40%), Positives = 34/62 (54%), Gaps = 4/62 (6%)
Frame = +1
Query: 62 DEDEEDV----CRICRNPGDADNPLRYPCACSGSIKFVHQDCLLQWLNHSNARQCEVCKH 117
DE+E + CRIC+ N L PCACSGS+K+ H+ C+ W N CE+C
Sbjct: 847 DEEEPLIQMVECRICQEEDSVSN-LETPCACSGSLKYAHRKCVQHWCNEKGDITCEICHQ 1023
Query: 118 PF 119
P+
Sbjct: 1024PY 1029
>TC92967 similar to GP|9972376|gb|AAG10626.1| Unknown protein {Arabidopsis
thaliana}, partial (32%)
Length = 701
Score = 55.8 bits (133), Expect = 8e-08
Identities = 23/65 (35%), Positives = 34/65 (51%), Gaps = 1/65 (1%)
Frame = +1
Query: 69 CRICRNPG-DADNPLRYPCACSGSIKFVHQDCLLQWLNHSNARQCEVCKHPFSFSPVYAE 127
CRIC ++ L PC+CSG++KF H+DC+ +W N CE+C + P Y
Sbjct: 136 CRICHEEEFESSKALEAPCSCSGTVKFAHRDCIQRWCNEKGNTTCEICLQ--QYEPGYTA 309
Query: 128 NAPAR 132
P +
Sbjct: 310 PPPKK 324
>TC91605 similar to GP|15809927|gb|AAL06891.1 At3g09760/F8A24.19
{Arabidopsis thaliana}, partial (21%)
Length = 948
Score = 54.7 bits (130), Expect = 2e-07
Identities = 27/83 (32%), Positives = 41/83 (48%), Gaps = 4/83 (4%)
Frame = +3
Query: 46 VATASTAPPSAKYDDDDED---EEDVCRICRNP-GDADNPLRYPCACSGSIKFVHQDCLL 101
V+ AS + + +D ED EE VCRIC G+ + + C+C G + H++C +
Sbjct: 567 VSVASPKGENVETEDGGEDIPEEEAVCRICMAELGEGADNFKLECSCKGELSLAHKECAV 746
Query: 102 QWLNHSNARQCEVCKHPFSFSPV 124
+W R C+VCK PV
Sbjct: 747 KWFGIKGNRTCDVCKQEVQNLPV 815
>BF650337 similar to GP|8885610|dbj| contains similarity to unknown
protein~gb|AAC32914.1~gene_id:F16F17.8 {Arabidopsis
thaliana}, partial (20%)
Length = 472
Score = 48.9 bits (115), Expect = 9e-06
Identities = 18/48 (37%), Positives = 29/48 (59%)
Frame = +3
Query: 69 CRICRNPGDADNPLRYPCACSGSIKFVHQDCLLQWLNHSNARQCEVCK 116
CRIC + D D + PC+C G++K+ H+ C+ +W N CE+C+
Sbjct: 318 CRICHDE-DEDLNMDTPCSCCGTLKYAHRICVQRWCNEKGDTTCEICQ 458
>TC80318 similar to GP|21593635|gb|AAM65602.1 unknown {Arabidopsis
thaliana}, partial (80%)
Length = 1172
Score = 47.8 bits (112), Expect = 2e-05
Identities = 28/89 (31%), Positives = 43/89 (47%), Gaps = 12/89 (13%)
Frame = +1
Query: 63 EDEEDV-------CRIC-RNPGDADNPLRYPCACSGSIKFVHQDCLLQWLNHSNA---RQ 111
+D+ED+ CRIC + D ++ L PC C G+ +FVH+ CL W +
Sbjct: 205 KDQEDIEAGSLPCCRICLESDSDPEDELISPCMCKGTQQFVHRSCLDHWRSVKEGFAFSH 384
Query: 112 CEVCKHPFSFS-PVYAENAPARLPFQEFV 139
C CK F Y +N+ ++ F+ FV
Sbjct: 385 CTTCKAQFHLRVETYEDNSWRKIKFRLFV 471
>BE999053 similar to GP|9972376|gb|A Unknown protein {Arabidopsis thaliana},
partial (19%)
Length = 173
Score = 46.2 bits (108), Expect = 6e-05
Identities = 19/48 (39%), Positives = 24/48 (49%)
Frame = +3
Query: 85 PCACSGSIKFVHQDCLLQWLNHSNARQCEVCKHPFSFSPVYAENAPAR 132
PCACSG+ KF H+DC+ W N CE C + P Y P +
Sbjct: 24 PCACSGTDKFAHRDCIQTWRNEKGNTTCETCLQ--QYEPGYTTPPPKK 161
>BF647971 similar to GP|21537278|gb unknown {Arabidopsis thaliana}, partial
(47%)
Length = 656
Score = 45.8 bits (107), Expect = 8e-05
Identities = 42/139 (30%), Positives = 54/139 (38%), Gaps = 15/139 (10%)
Frame = +1
Query: 16 IAATTPSSSSPSSSSSSPRGSKGKEIDAEAVATASTAPPSAKYDDDDEDEEDV------- 68
+A+ T + S SSSS + K P A +DDD D E
Sbjct: 157 LASNTDNQDSSSSSSETQPILKNH--------LCEIIPIPANDEDDDYDNESSHLLVVNA 312
Query: 69 ----CRICRNPGDADNPLRYPCACSGSIKFVHQDCLLQWLNHSNA---RQCEVCKHPFSF 121
CRIC + G D L PC C G+ K+VH+ CL W + C C+ F
Sbjct: 313 DHPQCRICLDLGGED--LIAPCHCKGTQKYVHRSCLDNWRSTKEGFAFSHCTECRALFII 486
Query: 122 SP-VYAENAPARLPFQEFV 139
P V + RL FQ V
Sbjct: 487 RPNVPKDRWWLRLKFQFLV 543
>CA991192 weakly similar to GP|21594296|gb unknown {Arabidopsis thaliana},
partial (28%)
Length = 782
Score = 43.1 bits (100), Expect = 5e-04
Identities = 31/130 (23%), Positives = 57/130 (43%), Gaps = 17/130 (13%)
Frame = +2
Query: 80 NPLRYPCACSGSIKFVHQDCLLQWLNHSNARQCEVCKHPFSFSPVYAENAPARL------ 133
N + C+C G + VH++CL++W + ++C+VC +N PA L
Sbjct: 23 NIFKMECSCKGDQRLVHEECLIKWFSTKRNKKCDVCL-------AEVQNLPANLVHECRS 181
Query: 134 ---------PFQEFVVGMAMKACHVLQFFVRLSF--VLSVWLLIIPFITFWIWRLAFVRS 182
+Q FVV + + F V L + + + ++I ++F + LA V +
Sbjct: 182 VQPRNIRLSAWQNFVVLVLISTLWYFHFIVDLLYRDLKTRGIIIAAAVSFTLSLLASVFA 361
Query: 183 FGEAQRLFFY 192
F A R + +
Sbjct: 362 FFLAIREYMW 391
>TC90157 similar to PIR|T51853|T51853 RING-H2 finger protein RHF1a
[imported] - Arabidopsis thaliana (fragment), partial
(44%)
Length = 1486
Score = 42.4 bits (98), Expect = 9e-04
Identities = 31/98 (31%), Positives = 41/98 (41%)
Frame = +1
Query: 18 ATTPSSSSPSSSSSSPRGSKGKEIDAEAVATASTAPPSAKYDDDDEDEEDVCRICRNPGD 77
AT S SSPSSSSS+P + PS+ DD+ ED C IC
Sbjct: 76 ATFTSFSSPSSSSSTP---------------PPFSAPSSSSAFDDDSFEDSCSICLESFS 210
Query: 78 ADNPLRYPCACSGSIKFVHQDCLLQWLNHSNARQCEVC 115
+P C C H C+L+W S ++C +C
Sbjct: 211 VHDPSTVTC-CKHE---YHLHCILEWSQRS--KECPIC 306
>TC82570 similar to PIR|T51853|T51853 RING-H2 finger protein RHF1a
[imported] - Arabidopsis thaliana (fragment), partial
(23%)
Length = 738
Score = 42.4 bits (98), Expect = 9e-04
Identities = 31/98 (31%), Positives = 41/98 (41%)
Frame = +2
Query: 18 ATTPSSSSPSSSSSSPRGSKGKEIDAEAVATASTAPPSAKYDDDDEDEEDVCRICRNPGD 77
AT S SSPSSSSS+P + PS+ DD+ ED C IC
Sbjct: 104 ATFTSFSSPSSSSSTP---------------PPFSAPSSSSAFDDDSFEDSCSICLESFS 238
Query: 78 ADNPLRYPCACSGSIKFVHQDCLLQWLNHSNARQCEVC 115
+P C C H C+L+W S ++C +C
Sbjct: 239 VHDPSTVTC-CKHE---YHLHCILEWSQRS--KECPIC 334
>TC91339 homologue to PIR|B84609|B84609 hypothetical protein At2g22120
[imported] - Arabidopsis thaliana, partial (47%)
Length = 681
Score = 40.0 bits (92), Expect = 0.004
Identities = 30/102 (29%), Positives = 40/102 (38%), Gaps = 10/102 (9%)
Frame = +1
Query: 48 TASTAPPSAKYDDDDED------EEDVCRICRNPGDADNPLRYPCACSGSIKFVHQDCLL 101
T+ PP D D E+ CRIC D C C G+ K+VH++CL
Sbjct: 154 TSPLVPPLPPVDPSSIDLEAGPSEQIQCRICLETDGRD--FIASCKCKGTSKYVHRECLD 327
Query: 102 QWLNHSNA---RQCEVCKHPFSFS-PVYAENAPARLPFQEFV 139
W + C CK P+ V A+ L F+ FV
Sbjct: 328 HWRSVKEGFAFAHCTTCKAPYHLRVHVAADRKWRTLKFRFFV 453
>TC77563 similar to GP|15450687|gb|AAK96615.1 AT5g59550/f2o15_210
{Arabidopsis thaliana}, partial (39%)
Length = 1468
Score = 38.9 bits (89), Expect = 0.010
Identities = 22/81 (27%), Positives = 37/81 (45%), Gaps = 1/81 (1%)
Frame = +2
Query: 38 GKEIDAEAVATASTAPPSAKYDDDDEDEEDVCRICRNPGDADNPLR-YPCACSGSIKFVH 96
G+ + A A + P+ + D + + E C +C+ + + R PC H
Sbjct: 524 GRSENPPASKAAIESIPTVEITDSEMESEIHCAVCKEQFELGSEARKMPCN-----HLYH 688
Query: 97 QDCLLQWLNHSNARQCEVCKH 117
DC+L WL+ N+ C VC+H
Sbjct: 689 SDCILPWLSMRNS--CPVCRH 745
>TC83503 weakly similar to PIR|T51851|T51851 RING-H2 finger protein RHB1a
[imported] - Arabidopsis thaliana, partial (44%)
Length = 835
Score = 38.5 bits (88), Expect = 0.013
Identities = 34/132 (25%), Positives = 53/132 (39%), Gaps = 16/132 (12%)
Frame = +1
Query: 6 EPPPSLDGTPIAATTPSSSSPSSSSSSPRGSKGK---------EIDAEAVA-TASTAPPS 55
+PPP+ + P S+ S + GS + E D +A A +A +P
Sbjct: 352 QPPPAPLPYDVVLGVPVSTDSESGKDTVSGSSFETLVTCEDIEESDLKAQAKSAPISPTK 531
Query: 56 AKYDDDDE------DEEDVCRICRNPGDADNPLRYPCACSGSIKFVHQDCLLQWLNHSNA 109
A+ +E +EED C IC D +NP S H C+L+W+ S++
Sbjct: 532 AELWKSNELHALVIEEEDGCPICLEEYDVENPK----TLSKCEHHFHLACILEWMERSDS 699
Query: 110 RQCEVCKHPFSF 121
C +C F
Sbjct: 700 --CPICDQEMIF 729
>TC87855 similar to GP|15081801|gb|AAK82555.1 AT5g41350/MYC6_6 {Arabidopsis
thaliana}, partial (48%)
Length = 1194
Score = 38.5 bits (88), Expect = 0.013
Identities = 31/104 (29%), Positives = 41/104 (38%)
Frame = +1
Query: 19 TTPSSSSPSSSSSSPRGSKGKEIDAEAVATASTAPPSAKYDDDDEDEEDVCRICRNPGDA 78
T+P S P S KG+ + A S S + + EED C IC DA
Sbjct: 610 TSPKSEEPKESEC-----KGQTDIEQDTAKDSEIELSKLGEPINLVEEDTCPICLEEYDA 774
Query: 79 DNPLRYPCACSGSIKFVHQDCLLQWLNHSNARQCEVCKHPFSFS 122
+NP + C H C+L+W+ S C VC FS
Sbjct: 775 ENP-KLTTQCGHDF---HLACILEWMERSET--CPVCDQVTVFS 888
>BG452562 similar to GP|9293902|dbj contains similarity to RING zinc finger
protein~gene_id:MVE11.15 {Arabidopsis thaliana}, partial
(59%)
Length = 664
Score = 36.6 bits (83), Expect = 0.049
Identities = 36/124 (29%), Positives = 52/124 (41%), Gaps = 1/124 (0%)
Frame = +2
Query: 18 ATTPSSSSPSSSSSSPRGSKGKEIDAEAVATASTAPPSAKYDDDDEDEEDVCRICRNPGD 77
A SSS SSS+SSP+ K I +A+ T T S + D E C IC +
Sbjct: 302 ALNNGSSSSSSSNSSPQ-LANKGIKKKALKTFPTVSYSTELKLPTLDTE--CVICLSEFT 472
Query: 78 ADNPLRYPCACSGSIKFVHQDCLLQWL-NHSNARQCEVCKHPFSFSPVYAENAPARLPFQ 136
+R C+ H C+ +WL +HS+ +C C V +E P LP
Sbjct: 473 KGEKVRILPKCNHGF---HVRCIDKWLKSHSSCPKCRQCLLETCRKIVGSEAPPXMLPVP 643
Query: 137 EFVV 140
++
Sbjct: 644 XTII 655
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.328 0.141 0.441
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 32,447,105
Number of Sequences: 36976
Number of extensions: 543937
Number of successful extensions: 6450
Number of sequences better than 10.0: 96
Number of HSP's better than 10.0 without gapping: 4133
Number of HSP's successfully gapped in prelim test: 202
Number of HSP's that attempted gapping in prelim test: 1081
Number of HSP's gapped (non-prelim): 5144
length of query: 982
length of database: 9,014,727
effective HSP length: 105
effective length of query: 877
effective length of database: 5,132,247
effective search space: 4500980619
effective search space used: 4500980619
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.8 bits)
S2: 63 (28.9 bits)
Medicago: description of AC136449.11


