

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC136286.2 - phase: 0
(251 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
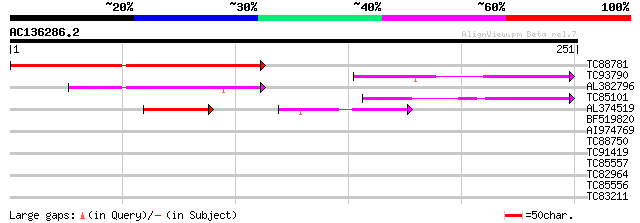
TC88781 similar to GP|9369366|gb|AAF87115.1| F10A5.18 {Arabidops... 134 4e-32
TC93790 weakly similar to GP|9369366|gb|AAF87115.1| F10A5.18 {Ar... 70 1e-12
AL382796 weakly similar to GP|20161084|dbj putative glyoxal oxid... 60 6e-10
TC85101 similar to GP|15809876|gb|AAL06866.1 AT3g53950/F5K20_250... 58 4e-09
AL374519 similar to GP|15809876|gb| AT3g53950/F5K20_250 {Arabido... 50 7e-09
BF519820 weakly similar to GP|15809876|gb AT3g53950/F5K20_250 {A... 30 1.1
AI974769 29 2.0
TC88750 similar to GP|16604523|gb|AAL24267.1 At2g31810/F20M17.15... 28 3.3
TC91419 similar to GP|21594030|gb|AAM65948.1 unknown {Arabidopsi... 27 5.7
TC85557 homologue to GP|1389640|dbj|BAA12982.1 PNDKN1 {Pisum sat... 27 7.5
TC82964 similar to GP|22655058|gb|AAM98120.1 predicted protein {... 27 7.5
TC85556 homologue to GP|1389640|dbj|BAA12982.1 PNDKN1 {Pisum sat... 27 7.5
TC83211 similar to GP|5803275|dbj|BAA83585.1 EST AU056133(S20320... 27 9.7
>TC88781 similar to GP|9369366|gb|AAF87115.1| F10A5.18 {Arabidopsis thaliana},
partial (67%)
Length = 1391
Score = 134 bits (336), Expect = 4e-32
Identities = 66/113 (58%), Positives = 82/113 (72%)
Frame = +1
Query: 1 MEVMPVPRVMPDMLLLPTGNVIILNGAANGTAGWENAANPVLYPVLYKPGLDNPFMKFEL 60
ME MP RVM DM+LLP G+V+++NGAA G+AGWE+ NPVL P LYKP +F+L
Sbjct: 1033 METMPGGRVMSDMILLPNGDVLLINGAAVGSAGWESGRNPVLNPFLYKPN-SLVGSRFQL 1209
Query: 61 LAPASTPRMYHSSAVLLPDGRILVGGSNPHRLYDFQAKYPTELSLDAYYPDYL 113
P+ PRMYHS+AVLL DGR+LV GSNPH Y+F +PTEL L+A+ P YL
Sbjct: 1210 QKPSDIPRMYHSTAVLLRDGRVLVAGSNPHVNYNFTRLFPTELRLEAFSPSYL 1368
>TC93790 weakly similar to GP|9369366|gb|AAF87115.1| F10A5.18 {Arabidopsis
thaliana}, partial (11%)
Length = 574
Score = 69.7 bits (169), Expect = 1e-12
Identities = 40/99 (40%), Positives = 54/99 (54%), Gaps = 1/99 (1%)
Frame = +1
Query: 153 NRIRVSMVAPSFTTHSFAMNQRLLFL-EVTALEEVVNSMQDQNFGEFGFGSSLGPGKIAN 211
N + V+M APSF THSF+MNQRLL L E+++ + VV +
Sbjct: 13 NMVSVTMYAPSFNTHSFSMNQRLLVLDELSSSKNVVGE--------------------ST 132
Query: 212 SVYKATVRGPPSLNVAPPGYYMLFVIHVGIPSVATWVHV 250
S Y+ V P S +APPGYY+LFV+H +PS WV +
Sbjct: 133 STYQVEVAIPNSPILAPPGYYLLFVVHQEVPSQGVWVQL 249
>AL382796 weakly similar to GP|20161084|dbj putative glyoxal oxidase {Oryza
sativa (japonica cultivar-group)}, partial (4%)
Length = 465
Score = 60.5 bits (145), Expect = 6e-10
Identities = 34/89 (38%), Positives = 46/89 (51%), Gaps = 2/89 (2%)
Frame = +3
Query: 27 AANGTAGWENAANPVLYPVLYKPGLDNPFMKFELLAPASTPRMYHSSAVLLPDGRILVGG 86
A N T A P P+LY P + ++ R+YHSSA L+PDG + V G
Sbjct: 30 ATNKTPRLHTAGKPTRNPLLYNPH-EKKGSRWSKFTEDPIIRVYHSSATLIPDGTVFVAG 206
Query: 87 SNPHRLY--DFQAKYPTELSLDAYYPDYL 113
SNP+R Y Q ++PTE + +YP YL
Sbjct: 207 SNPNRYYCGMDQCEHPTEHRAEKFYPPYL 293
>TC85101 similar to GP|15809876|gb|AAL06866.1 AT3g53950/F5K20_250
{Arabidopsis thaliana}, partial (12%)
Length = 519
Score = 57.8 bits (138), Expect = 4e-09
Identities = 33/94 (35%), Positives = 48/94 (50%)
Frame = +2
Query: 157 VSMVAPSFTTHSFAMNQRLLFLEVTALEEVVNSMQDQNFGEFGFGSSLGPGKIANSVYKA 216
V++ + F THSF+ QRL+ L G S++ G + ++
Sbjct: 5 VNLGSAPFATHSFSQGQRLIKL--------------------GVASAMVDG---DQRWRI 115
Query: 217 TVRGPPSLNVAPPGYYMLFVIHVGIPSVATWVHV 250
PPS VAPPGYYMLF ++ G+PSVA W+H+
Sbjct: 116 RCTAPPSGMVAPPGYYMLFAVNQGVPSVARWIHM 217
>AL374519 similar to GP|15809876|gb| AT3g53950/F5K20_250 {Arabidopsis
thaliana}, partial (17%)
Length = 532
Score = 49.7 bits (117), Expect(2) = 7e-09
Identities = 20/31 (64%), Positives = 26/31 (83%)
Frame = +1
Query: 60 LLAPASTPRMYHSSAVLLPDGRILVGGSNPH 90
+L P + PR+YHS+A LLPDGR+L+ GSNPH
Sbjct: 4 VLNPGTVPRLYHSTANLLPDGRVLLAGSNPH 96
Score = 26.9 bits (58), Expect(2) = 7e-09
Identities = 20/60 (33%), Positives = 32/60 (53%), Gaps = 1/60 (1%)
Frame = +3
Query: 120 LRPVIVAV-EVVNSTLSYESLFSVSFLLREVKDVNRIRVSMVAPSFTTHSFAMNQRLLFL 178
+RP I+ V E V + ++ + SV + V I V++ + F THSF+ QRL+ L
Sbjct: 204 IRPKILEVPETVLYGVGFDVVVSVPLPV-----VGIIEVNLGSAPFATHSFSQGQRLIKL 368
>BF519820 weakly similar to GP|15809876|gb AT3g53950/F5K20_250 {Arabidopsis
thaliana}, partial (34%)
Length = 574
Score = 29.6 bits (65), Expect = 1.1
Identities = 21/83 (25%), Positives = 36/83 (43%), Gaps = 2/83 (2%)
Frame = +1
Query: 15 LLPTGNVIILNGAANGTAGWENAANPVLYPVLYKPGLDNPFMKFELLAPASTPRM--YHS 72
+LP G++II+ G + T V + P +N + F L +M +
Sbjct: 172 ILPDGSIIIIGGRGSNT-------------VEFYPKRENGVVLFPFLQETEDTQMDNLYP 312
Query: 73 SAVLLPDGRILVGGSNPHRLYDF 95
LLP+G++ V + +YDF
Sbjct: 313 YVHLLPNGKLFVFANTKSVMYDF 381
>AI974769
Length = 361
Score = 28.9 bits (63), Expect = 2.0
Identities = 19/67 (28%), Positives = 36/67 (53%), Gaps = 13/67 (19%)
Frame = +1
Query: 39 NPVLYPVLYKPGLDNPFMKFELLAPASTPRMYHSSAVLLP-------------DGRILVG 85
+P+L P ++K L+ F K E+L + P++ +SS++ + +GRI++G
Sbjct: 115 SPILDPTIFKNFLNFKFPKIEILCNFN-PQLRNSSSLTIFYFLIFYF*FFEKFEGRIVIG 291
Query: 86 GSNPHRL 92
G N + L
Sbjct: 292 GGNLNNL 312
>TC88750 similar to GP|16604523|gb|AAL24267.1 At2g31810/F20M17.15
{Arabidopsis thaliana}, partial (36%)
Length = 803
Score = 28.1 bits (61), Expect = 3.3
Identities = 25/98 (25%), Positives = 45/98 (45%), Gaps = 7/98 (7%)
Frame = +3
Query: 120 LRPVIVAVEVVNSTLSYESLFSVSFLLREVKDV-------NRIRVSMVAPSFTTHSFAMN 172
LR V+ ++ + + L E L + RE+ V NR + + +F ++
Sbjct: 408 LRQVVEQLQKLVNVLKVEDLSREPQVERELMLVKVHADPQNRAEIKWLVDTFRAKIVDIS 587
Query: 173 QRLLFLEVTALEEVVNSMQDQNFGEFGFGSSLGPGKIA 210
+ + +EVT + ++Q +NFG+FG GKIA
Sbjct: 588 EHSVTIEVTGDPGKMVAVQ-RNFGKFGIKEIARTGKIA 698
>TC91419 similar to GP|21594030|gb|AAM65948.1 unknown {Arabidopsis
thaliana}, partial (27%)
Length = 817
Score = 27.3 bits (59), Expect = 5.7
Identities = 11/22 (50%), Positives = 14/22 (63%)
Frame = +1
Query: 221 PPSLNVAPPGYYMLFVIHVGIP 242
PP +AP YY+ FV +GIP
Sbjct: 475 PPEWGLAPSSYYVPFVPIIGIP 540
>TC85557 homologue to GP|1389640|dbj|BAA12982.1 PNDKN1 {Pisum sativum},
partial (75%)
Length = 1129
Score = 26.9 bits (58), Expect = 7.5
Identities = 11/32 (34%), Positives = 18/32 (55%)
Frame = +2
Query: 30 GTAGWENAANPVLYPVLYKPGLDNPFMKFELL 61
G A W+++ + +Y YKP +D F F+ L
Sbjct: 806 GPAAWQSSLHSWIYE*RYKPEVDGEFSHFDPL 901
>TC82964 similar to GP|22655058|gb|AAM98120.1 predicted protein {Arabidopsis
thaliana}, partial (73%)
Length = 1244
Score = 26.9 bits (58), Expect = 7.5
Identities = 16/42 (38%), Positives = 21/42 (49%)
Frame = -1
Query: 199 GFGSSLGPGKIANSVYKATVRGPPSLNVAPPGYYMLFVIHVG 240
GF G GKI N V TV+ PP++ + P Y +I G
Sbjct: 350 GFPFEEGTGKI-NPVLYDTVQSPPTIELTTPDYVAEALIQRG 228
>TC85556 homologue to GP|1389640|dbj|BAA12982.1 PNDKN1 {Pisum sativum},
complete
Length = 785
Score = 26.9 bits (58), Expect = 7.5
Identities = 11/32 (34%), Positives = 18/32 (55%)
Frame = +2
Query: 30 GTAGWENAANPVLYPVLYKPGLDNPFMKFELL 61
G A W+++ + +Y YKP +D F F+ L
Sbjct: 464 GPAAWQSSLHSWIYE*RYKPEVDGEFSHFDPL 559
>TC83211 similar to GP|5803275|dbj|BAA83585.1 EST AU056133(S20320)
corresponds to a region of the predicted gene.~Similar
to Caenorhabditis, partial (22%)
Length = 690
Score = 26.6 bits (57), Expect = 9.7
Identities = 15/41 (36%), Positives = 20/41 (48%)
Frame = -1
Query: 108 YYPDYLRPELDTLRPVIVAVEVVNSTLSYESLFSVSFLLRE 148
Y D +R + L P + + VNS SYE + SF L E
Sbjct: 204 YLHD*VRKCITPLTPTVTQLSNVNSRRSYEYEITTSFQLHE 82
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.321 0.138 0.412
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 7,139,564
Number of Sequences: 36976
Number of extensions: 94237
Number of successful extensions: 473
Number of sequences better than 10.0: 26
Number of HSP's better than 10.0 without gapping: 465
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 469
length of query: 251
length of database: 9,014,727
effective HSP length: 94
effective length of query: 157
effective length of database: 5,538,983
effective search space: 869620331
effective search space used: 869620331
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 57 (26.6 bits)
Medicago: description of AC136286.2


