

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC135801.5 - phase: 0
(736 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
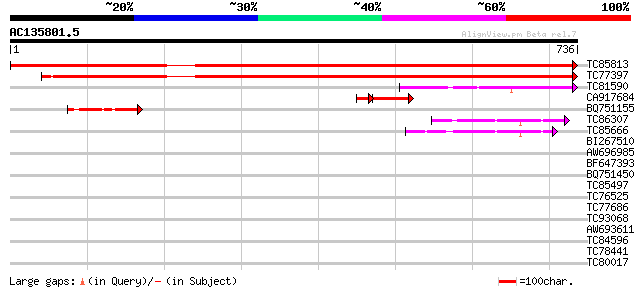
Score E
Sequences producing significant alignments: (bits) Value
TC85813 homologue to GP|6503253|gb|AAC09468.2| putative NADPH-cy... 1398 0.0
TC77397 homologue to PIR|S37159|S37159 NADPH--ferrihemoprotein r... 943 0.0
TC81590 similar to GP|6041790|gb|AAF02110.1| putative NADPH-ferr... 154 9e-38
CA917684 homologue to GP|6503253|gb|A putative NADPH-cytochrome ... 104 1e-30
BQ751155 similar to GP|10041541|em unnamed protein product {Beau... 69 6e-12
TC86307 homologue to PIR|T06773|T06773 ferredoxin--NADP+ reducta... 66 4e-11
TC85666 homologue to SP|P10933|FENR_PEA Ferredoxin--NADP reducta... 63 5e-10
BI267510 weakly similar to PIR|C96782|C96 unknown protein F22H5.... 40 0.004
AW696985 weakly similar to PIR|C96782|C967 unknown protein F22H5... 33 0.39
BF647393 similar to GP|6041790|gb|A putative NADPH-ferrihemoprot... 33 0.39
BQ751450 similar to GP|21954534|db 6-phosphogluconate dehydrogen... 33 0.51
TC85497 similar to PIR|T05174|T05174 hypothetical protein T6K22.... 32 0.88
TC76525 similar to GP|15450864|gb|AAK96703.1 Unknown protein {Ar... 31 1.5
TC77686 similar to GP|16649123|gb|AAL24413.1 unknown protein {Ar... 30 3.3
TC93068 similar to GP|3170523|gb|AAC18088.1| coatomer alpha subu... 30 3.3
AW693611 weakly similar to GP|20197699|gb| putative LRR receptor... 29 5.7
TC84596 weakly similar to GP|17064866|gb|AAL32587.1 Unknown prot... 29 5.7
TC78441 similar to GP|16226543|gb|AAL16196.1 AT3g15000/K15M2_14 ... 28 9.7
TC80017 similar to GP|10177301|dbj|BAB10562. gene_id:MDC12.17~un... 28 9.7
>TC85813 homologue to GP|6503253|gb|AAC09468.2| putative NADPH-cytochrome
P450 reductase {Pisum sativum}, complete
Length = 2581
Score = 1398 bits (3618), Expect = 0.0
Identities = 701/736 (95%), Positives = 701/736 (95%)
Frame = +3
Query: 1 MQDSSSMKFSPLDLMSALIKGKIDPSNGTVPASLILENREFVMILTTSIAVLIGCVVVLI 60
MQDSSSMKFSPLDLMSALIKGKIDPSNGTVPASLILENREFVMILTTSIAVLIGCVVVLI
Sbjct: 102 MQDSSSMKFSPLDLMSALIKGKIDPSNGTVPASLILENREFVMILTTSIAVLIGCVVVLI 281
Query: 61 WRRSNSQKPKPIEVPKRVIEKLPELEIDDGTKKVTVFFGTQTGTAEGFAKAIAEEAKARY 120
WRRSNSQKPKPIEVPKRVIEKLPELEIDDGTKKVTVFFGTQTGTAEGFAKAIAEEAKARY
Sbjct: 282 WRRSNSQKPKPIEVPKRVIEKLPELEIDDGTKKVTVFFGTQTGTAEGFAKAIAEEAKARY 461
Query: 121 EKAKFKVVDMDDYAADDDEYEEKLKKETMALFFLATYGDGEPTDNAARFYKWFEEFEGEE 180
EKAKFKVVDMDDYAADDDEYEEKLKKETMALFFLATYGDGEPTDNAARFYKWFEEFEGEE
Sbjct: 462 EKAKFKVVDMDDYAADDDEYEEKLKKETMALFFLATYGDGEPTDNAARFYKWFEEFEGEE 641
Query: 181 DSFKNLQYGVFGLGNRQYEHFNKVCTQALMHSCGFLSFDYSDGVASFRAFSDIYHSTQVA 240
DSFKNLQYGVFGLGNRQYEHFNKV A
Sbjct: 642 DSFKNLQYGVFGLGNRQYEHFNKV-----------------------------------A 716
Query: 241 KIVDDKLLEKGGNRLVPVGLGDDDQCIEDDFTAWKEELWPALDQLLRDEDDATVATPYTA 300
KIVDDKLLEKGGNRLVPVGLGDDDQCIEDDFTAWKEELWPALDQLLRDEDDATVATPYTA
Sbjct: 717 KIVDDKLLEKGGNRLVPVGLGDDDQCIEDDFTAWKEELWPALDQLLRDEDDATVATPYTA 896
Query: 301 SVLEYRVVIRDQLDATVDEKKQLNGNGHAVVDAHHPVRANVAVRKELHTPASDRSCTHLE 360
SVLEYRVVIRDQLDATVDEKKQLNGNGHAVVDAHHPVRANVAVRKELHTPASDRSCTHLE
Sbjct: 897 SVLEYRVVIRDQLDATVDEKKQLNGNGHAVVDAHHPVRANVAVRKELHTPASDRSCTHLE 1076
Query: 361 FDISGTGVVYETGDHVGVYCENLSDTVEEAERILGLSPDTYFSVHTDDEDGKPLGGSSLP 420
FDISGTGVVYETGDHVGVYCENLSDTVEEAERILGLSPDTYFSVHTDDEDGKPLGGSSLP
Sbjct: 1077FDISGTGVVYETGDHVGVYCENLSDTVEEAERILGLSPDTYFSVHTDDEDGKPLGGSSLP 1256
Query: 421 PPFPPCTLRTALAKYADVLSSPKKSALLALAAHASDPSEADRLRHLASPAGKDEYAEWVI 480
PPFPPCTLRTALAKYADVLSSPKKSALLALAAHASDPSEADRLRHLASPAGKDEYAEWVI
Sbjct: 1257PPFPPCTLRTALAKYADVLSSPKKSALLALAAHASDPSEADRLRHLASPAGKDEYAEWVI 1436
Query: 481 ASQRSLLEVMAEFSSAKPPIGVFFASVAPRLQPRYYSISSSPRVAPSRIHVTCALVHDKM 540
ASQRSLLEVMAEFSSAKPPIGVFFASVAPRLQPRYYSISSSPRVAPSRIHVTCALVHDKM
Sbjct: 1437ASQRSLLEVMAEFSSAKPPIGVFFASVAPRLQPRYYSISSSPRVAPSRIHVTCALVHDKM 1616
Query: 541 PTGRIHQGVCSTWMKNSAPLEKSQDCSWAPIFVRQSNFRLPADNKVPIIMIGPGTGLAPF 600
PTGRIHQGVCSTWMKNSAPLEKSQDCSWAPIFVRQSNFRLPADNKVPIIMIGPGTGLAPF
Sbjct: 1617PTGRIHQGVCSTWMKNSAPLEKSQDCSWAPIFVRQSNFRLPADNKVPIIMIGPGTGLAPF 1796
Query: 601 RGFLQERLALKEEGAELGPSVLFFGCRNRQVDYIYEDELNHFVHGGALSELIVAFSREGP 660
RGFLQERLALKEEGAELGPSVLFFGCRNRQVDYIYEDELNHFVHGGALSELIVAFSREGP
Sbjct: 1797RGFLQERLALKEEGAELGPSVLFFGCRNRQVDYIYEDELNHFVHGGALSELIVAFSREGP 1976
Query: 661 TKEYVQHKMIEKASDIWNMISQGAYIYVCGDAKGMAKDVHRTLHTILQEQGSLDNSKTES 720
TKEYVQHKMIEKASDIWNMISQGAYIYVCGDAKGMAKDVHRTLHTILQEQGSLDNSKTES
Sbjct: 1977TKEYVQHKMIEKASDIWNMISQGAYIYVCGDAKGMAKDVHRTLHTILQEQGSLDNSKTES 2156
Query: 721 MVKNLQMTGRYLRDVW 736
MVKNLQMTGRYLRDVW
Sbjct: 2157MVKNLQMTGRYLRDVW 2204
>TC77397 homologue to PIR|S37159|S37159 NADPH--ferrihemoprotein reductase
(EC 1.6.2.4) - spring vetch, complete
Length = 2519
Score = 943 bits (2437), Expect = 0.0
Identities = 467/700 (66%), Positives = 556/700 (78%), Gaps = 5/700 (0%)
Frame = +2
Query: 42 VMILTTSIAVLIGCVVVLIWRRSN--SQKPKPIEVPKRVI-EKLPELEIDDGTKKVTVFF 98
V+I+TTS AV+IG ++V +W++S+ S++ KP+ VPK ++ E+ + +I DG KVTVFF
Sbjct: 167 VLIVTTSAAVIIG-LLVFLWKKSSDRSKELKPVIVPKSLVKEEDDDADIADGKTKVTVFF 343
Query: 99 GTQTGTAEGFAKAIAEEAKARYEKAKFKVVDMDDYAADDDEYEEKLKKETMALFFLATYG 158
GTQTGTAEGFAKA+AEE KARYEKA KVVDMDDYAADDD+YEEKLKKET+A F LATYG
Sbjct: 344 GTQTGTAEGFAKALAEEIKARYEKAFVKVVDMDDYAADDDQYEEKLKKETLAFFMLATYG 523
Query: 159 DGEPTDNAARFYKWFEEFEGEEDSF-KNLQYGVFGLGNRQYEHFNKVCTQALMHSCGFLS 217
DGEPTDNAARFYKWF E + E ++ + L YGVFGLGNRQYEHFNK+
Sbjct: 524 DGEPTDNAARFYKWFTEGKDERGTWLQQLTYGVFGLGNRQYEHFNKI------------- 664
Query: 218 FDYSDGVASFRAFSDIYHSTQVAKIVDDKLLEKGGNRLVPVGLGDDDQCIEDDFTAWKEE 277
K+VDD L E+G RLVP+G+GDDDQ IEDDF AWKE
Sbjct: 665 ----------------------GKVVDDDLSEQGAKRLVPLGMGDDDQSIEDDFNAWKES 778
Query: 278 LWPALDQLLRDEDDA-TVATPYTASVLEYRVVIRDQLDATVDEKKQLNGNGHAVVDAHHP 336
LWP LDQLLRDEDD TV+TPYTA++ EYRVV D E NG AV D HHP
Sbjct: 779 LWPELDQLLRDEDDVNTVSTPYTAAISEYRVVFHDPTVTPSYENHFNAANGGAVFDIHHP 958
Query: 337 VRANVAVRKELHTPASDRSCTHLEFDISGTGVVYETGDHVGVYCENLSDTVEEAERILGL 396
RANVAVR+ELH P SDRSC HLEFD+SGTGV YETGDHVGVY +N +TV+EA ++LG
Sbjct: 959 CRANVAVRRELHKPQSDRSCIHLEFDVSGTGVTYETGDHVGVYADNCDETVKEAGKLLGQ 1138
Query: 397 SPDTYFSVHTDDEDGKPLGGSSLPPPFPPCTLRTALAKYADVLSSPKKSALLALAAHASD 456
D FS+HTD+EDG LGGS LPP PCT+RTALA+YAD+L+ P+K+AL+ALAAHAS+
Sbjct: 1139DLDLLFSLHTDNEDGTSLGGSLLPPFPGPCTVRTALARYADLLNPPRKAALIALAAHASE 1318
Query: 457 PSEADRLRHLASPAGKDEYAEWVIASQRSLLEVMAEFSSAKPPIGVFFASVAPRLQPRYY 516
PSEA+RL+ L+SP GKDEY++WV+ S R+LLEVMA+F SAKPP+GVFFA++APRLQPRYY
Sbjct: 1319PSEAERLKFLSSPQGKDEYSKWVVGSHRTLLEVMADFPSAKPPLGVFFAAIAPRLQPRYY 1498
Query: 517 SISSSPRVAPSRIHVTCALVHDKMPTGRIHQGVCSTWMKNSAPLEKSQDCSWAPIFVRQS 576
SISSSPR AP R+HVTCALV PTGRIH+GVCSTWMKN+ P E+S+DCSWAPIF+R S
Sbjct: 1499SISSSPRFAPQRVHVTCALVEGPTPTGRIHKGVCSTWMKNAIPSEESRDCSWAPIFIRPS 1678
Query: 577 NFRLPADNKVPIIMIGPGTGLAPFRGFLQERLALKEEGAELGPSVLFFGCRNRQVDYIYE 636
NF+LPAD +PIIM+GPGTGLAPFRGFLQER ALKE+G +LGP++LFFGCRNRQ+D+IYE
Sbjct: 1679NFKLPADPSIPIIMVGPGTGLAPFRGFLQERFALKEDGVQLGPALLFFGCRNRQMDFIYE 1858
Query: 637 DELNHFVHGGALSELIVAFSREGPTKEYVQHKMIEKASDIWNMISQGAYIYVCGDAKGMA 696
+ELN+FV G+LSELIVAFSREGP KEYVQHKM++KAS W++ISQG Y+YVCGDAKGMA
Sbjct: 1859EELNNFVEQGSLSELIVAFSREGPEKEYVQHKMMDKASYFWSLISQGGYLYVCGDAKGMA 2038
Query: 697 KDVHRTLHTILQEQGSLDNSKTESMVKNLQMTGRYLRDVW 736
+DVHRTLHTI+Q+Q + D+SK E+ VK LQM GRYLRDVW
Sbjct: 2039RDVHRTLHTIVQQQENADSSKAEATVKKLQMDGRYLRDVW 2158
>TC81590 similar to GP|6041790|gb|AAF02110.1| putative
NADPH-ferrihemoprotein reductase {Arabidopsis thaliana},
partial (33%)
Length = 1095
Score = 154 bits (390), Expect = 9e-38
Identities = 84/236 (35%), Positives = 129/236 (54%), Gaps = 6/236 (2%)
Frame = +3
Query: 507 VAPRLQPRYYSISSSPRVAPSRIHVTCALVHDKMPTGRIHQGVCSTWMKNSAPLEKSQDC 566
+ P L+ R +SISSS V P+++H+T ++V P R +G+CS+W+ P
Sbjct: 12 LVPMLKTRAFSISSSQSVHPNQVHLTVSVVSWTTPYKRKKKGLCSSWLATLDPRNAVGIP 191
Query: 567 SWAPIFVRQSNFRLPADNKVPIIMIGPGTGLAPFRGFLQERLALKEEGAELGPSVLFFGC 626
+W Q A +P+I++GPGTG APFRGF++ER AL+ + P + FFGC
Sbjct: 192 AWF-----QKGSLPTASPSLPLILVGPGTGCAPFRGFIEER-ALQSKTISTAPIMFFFGC 353
Query: 627 RNRQVDYIYEDE-LNHFVHGGALSE-----LIVAFSREGPTKEYVQHKMIEKASDIWNMI 680
N D++Y+D LNH + G LSE VAFSR+ P K YVQHKM E + +WN++
Sbjct: 354 WNEDGDFLYKDLWLNHSQNNGVLSEANGGGFYVAFSRDQPEKVYVQHKMKEHSGRVWNLL 533
Query: 681 SQGAYIYVCGDAKGMAKDVHRTLHTILQEQGSLDNSKTESMVKNLQMTGRYLRDVW 736
+ GA +Y+ G + M DV I+ ++ + ++ L+ G+Y + W
Sbjct: 534 ADGAAVYIAGSSTKMPTDVTSAFEEIVSKENEVSKEDAVRWIRALERCGKYHIEAW 701
>CA917684 homologue to GP|6503253|gb|A putative NADPH-cytochrome P450
reductase {Pisum sativum}, partial (10%)
Length = 705
Score = 104 bits (259), Expect(2) = 1e-30
Identities = 51/53 (96%), Positives = 53/53 (99%)
Frame = +3
Query: 472 KDEYAEWVIASQRSLLEVMAEFSSAKPPIGVFFASVAPRLQPRYYSISSSPRV 524
+DEYAEWVIASQRSLLEVMAEFSSAKPPIGVFFASVAPRLQPRYYSISSSPR+
Sbjct: 150 QDEYAEWVIASQRSLLEVMAEFSSAKPPIGVFFASVAPRLQPRYYSISSSPRL 308
Score = 47.8 bits (112), Expect(2) = 1e-30
Identities = 22/22 (100%), Positives = 22/22 (100%)
Frame = +2
Query: 451 AAHASDPSEADRLRHLASPAGK 472
AAHASDPSEADRLRHLASPAGK
Sbjct: 2 AAHASDPSEADRLRHLASPAGK 67
>BQ751155 similar to GP|10041541|em unnamed protein product {Beauveria
bassiana}, partial (20%)
Length = 812
Score = 68.9 bits (167), Expect = 6e-12
Identities = 37/97 (38%), Positives = 60/97 (61%)
Frame = +1
Query: 76 KRVIEKLPELEIDDGTKKVTVFFGTQTGTAEGFAKAIAEEAKARYEKAKFKVVDMDDYAA 135
+ ++EK+ E K VF+G+QTGTAE +A +A+E K+RY + V D++DY
Sbjct: 520 RNILEKMEE-----SGKNCIVFYGSQTGTAEDYASRLAKEGKSRY-GLETMVADLEDY-- 675
Query: 136 DDDEYEEKLKKETMALFFLATYGDGEPTDNAARFYKW 172
D + + + + + +F LATYG+GEPT +A FY++
Sbjct: 676 -DYDNLDAIPSDKVVMFVLATYGEGEPTRHAVDFYEF 783
>TC86307 homologue to PIR|T06773|T06773 ferredoxin--NADP+ reductase (EC
1.18.1.2) - garden pea (fragment), complete
Length = 1648
Score = 66.2 bits (160), Expect = 4e-11
Identities = 56/186 (30%), Positives = 86/186 (46%), Gaps = 7/186 (3%)
Frame = +3
Query: 548 GVCSTWMKNSAPLEKSQDCSWAPIFVRQSNFRLPADN-KVPIIMIGPGTGLAPFRGFLQE 606
GVCS ++ +S P +K + + + LP D+ IMI GTG+AP+RG+L+
Sbjct: 828 GVCSNFLCDSKPGDKIKITGPSGKIML-----LPEDDPNATHIMIATGTGVAPYRGYLRR 992
Query: 607 RLALKEEGAELGPSV-LFFGCRNRQVDYIYEDELNHFVHGGALS-ELIVAFSREGPT--- 661
+ G LF G N +Y+DE ++ + A SRE
Sbjct: 993 MFMESVPTFKFGGLAWLFLGVANSD-SLLYDDEFTKYLKDYPENFRYDRALSREEKNRNG 1169
Query: 662 -KEYVQHKMIEKASDIWNMISQGAYIYVCGDAKGMAKDVHRTLHTILQEQGSLDNSKTES 720
K YVQ K+ E + +I+ ++ GA+IY CG KGM + TL + + +G K
Sbjct: 1170 GKMYVQDKIEEYSDEIFKLLDNGAHIYFCG-LKGMMPGIQETLKRVAENRGESWEEKLSQ 1346
Query: 721 MVKNLQ 726
+ KN Q
Sbjct: 1347 LKKNKQ 1364
>TC85666 homologue to SP|P10933|FENR_PEA Ferredoxin--NADP reductase leaf
isozyme chloroplast precursor (EC 1.18.1.2) (FNR).
[Garden pea], partial (96%)
Length = 1389
Score = 62.8 bits (151), Expect = 5e-10
Identities = 59/209 (28%), Positives = 96/209 (45%), Gaps = 11/209 (5%)
Frame = +3
Query: 514 RYYSISSSPR--VAPSRIHVTCA--LVHDKMPTGRIHQGVCSTWMKNSAPLEKSQDCSWA 569
R YSI+SS S+ C LV+ G + +GVCS ++ + P + Q
Sbjct: 507 RLYSIASSALGDFGDSKTVSLCVKRLVYTN-DAGEVVKGVCSNFLCDLRPGSEVQITG-- 677
Query: 570 PIFVRQSNFRLPADNKVPIIMIGPGTGLAPFRGFLQERLALKEEGAEL-GPSVLFFGCRN 628
P+ +P D +IM+G GTG+APFR FL + K E + G + LF G
Sbjct: 678 PV---GKEMLMPKDPNATVIMLGTGTGIAPFRSFLWKMFFEKHEDYKFNGLAWLFLGVPT 848
Query: 629 RQVDYIYEDELNHFVHGGALS-ELIVAFSREGPT----KEYVQHKMIEKASDIWNMISQ- 682
+Y++E + L A SRE K Y+Q +M + A ++W ++ +
Sbjct: 849 SS-SLLYKEEFEKMKEKAPENFRLDFAVSREQVNDKGEKMYIQTRMAQYAEELWELLKKD 1025
Query: 683 GAYIYVCGDAKGMAKDVHRTLHTILQEQG 711
++Y+CG KGM K + + ++ + G
Sbjct: 1026NTFVYMCG-LKGMEKGIDDIMVSLAAKDG 1109
>BI267510 weakly similar to PIR|C96782|C96 unknown protein F22H5.13
[imported] - Arabidopsis thaliana, partial (20%)
Length = 634
Score = 39.7 bits (91), Expect = 0.004
Identities = 39/167 (23%), Positives = 67/167 (39%)
Frame = +1
Query: 96 VFFGTQTGTAEGFAKAIAEEAKARYEKAKFKVVDMDDYAADDDEYEEKLKKETMALFFLA 155
+ F +QT T++ A + + F ++D +Y +D L KET+ L +
Sbjct: 226 ILFISQTNTSKTLANHLHRFLTSN--NLAFDIIDGRNYEPED------LPKETLLLIVAS 381
Query: 156 TYGDGEPTDNAARFYKWFEEFEGEEDSFKNLQYGVFGLGNRQYEHFNKVCTQALMHSCGF 215
T+ DG+P F W E ED F +G+ ++ C F
Sbjct: 382 TWEDGKPPPEYKFFADWLAE--SSED---------FRVGS------------LILSKCRF 492
Query: 216 LSFDYSDGVASFRAFSDIYHSTQVAKIVDDKLLEKGGNRLVPVGLGD 262
F G + D +++ V K + ++ GGN ++P+G GD
Sbjct: 493 AVFGVGSG-----GYGDNFNA--VGKEIVKRMKALGGNEMIPLGEGD 612
>AW696985 weakly similar to PIR|C96782|C967 unknown protein F22H5.13
[imported] - Arabidopsis thaliana, partial (14%)
Length = 736
Score = 33.1 bits (74), Expect = 0.39
Identities = 22/83 (26%), Positives = 36/83 (42%)
Frame = +3
Query: 96 VFFGTQTGTAEGFAKAIAEEAKARYEKAKFKVVDMDDYAADDDEYEEKLKKETMALFFLA 155
+ F +QT T++ A + + F ++D +Y +D L KET+ L +
Sbjct: 171 ILFISQTNTSKTLANHLHRFLTSN--NLAFDIIDGRNYEPED------LPKETLLLIVAS 326
Query: 156 TYGDGEPTDNAARFYKWFEEFEG 178
T+ DG+P F W E G
Sbjct: 327 TWEDGKPPPEYKFFADWLAESSG 395
>BF647393 similar to GP|6041790|gb|A putative NADPH-ferrihemoprotein
reductase {Arabidopsis thaliana}, partial (7%)
Length = 522
Score = 33.1 bits (74), Expect = 0.39
Identities = 23/90 (25%), Positives = 42/90 (46%)
Frame = +3
Query: 76 KRVIEKLPELEIDDGTKKVTVFFGTQTGTAEGFAKAIAEEAKARYEKAKFKVVDMDDYAA 135
KRV + E++ K+ + + TQTG A A+ +A EA+ R ++ + Y
Sbjct: 273 KRV*RIMKEIQQQQHHHKLLILYATQTGNALDAAERLAREAERR--ACPINLLSLHQYEP 446
Query: 136 DDDEYEEKLKKETMALFFLATYGDGEPTDN 165
+ L +E +F ++T G G+ D+
Sbjct: 447 N------LLPQEEAVVFVVSTTGXGDSPDS 518
>BQ751450 similar to GP|21954534|db 6-phosphogluconate dehydrogenase
{Aspergillus oryzae}, partial (36%)
Length = 661
Score = 32.7 bits (73), Expect = 0.51
Identities = 38/154 (24%), Positives = 60/154 (38%), Gaps = 1/154 (0%)
Frame = -1
Query: 416 GSSLPPPFPPCTLRTALAKYADVLSSPKKSALLALAAHASDPSEADRLRHLASPAGKDEY 475
GSS PP P C + A + + +SP +A + +S + + RL SPAG+
Sbjct: 529 GSSTSPPLPSCGVVAASSAPSSSRTSPTLTAPTPTSRTSSSTTSSTRLSTRPSPAGE--- 359
Query: 476 AEWVIASQRSLLEVMAEFSSAKPPIGVFFASVAPRLQPRYYSISSSPRVAPSRIHVTCAL 535
S L +++ + PP A+ APR P S P+ + +
Sbjct: 358 -----TSLPRPLSLVSRLPPSLPPCPGLTAT-APRTSPPTSCRPSVTTSVPTPSASSPSH 197
Query: 536 VHDKMPTGR-IHQGVCSTWMKNSAPLEKSQDCSW 568
+K P G+ IH SA + + SW
Sbjct: 196 ASEKYPEGQDIHVNWTGRGGNVSASTYNA*NISW 95
>TC85497 similar to PIR|T05174|T05174 hypothetical protein T6K22.50 -
Arabidopsis thaliana, partial (14%)
Length = 1121
Score = 32.0 bits (71), Expect = 0.88
Identities = 24/95 (25%), Positives = 37/95 (38%), Gaps = 2/95 (2%)
Frame = +2
Query: 152 FFL--ATYGDGEPTDNAARFYKWFEEFEGEEDSFKNLQYGVFGLGNRQYEHFNKVCTQAL 209
FFL A + P Y+W E E+ K +YGV + + Y FN Q +
Sbjct: 8 FFLDKAPFNSEAPVTVTMSAYRWKSFEENEDRPSKPRRYGVTEMRSPHYTLFNHNVLQDI 187
Query: 210 MHSCGFLSFDYSDGVASFRAFSDIYHSTQVAKIVD 244
S G DY DG+ + + +++D
Sbjct: 188 FESMG----DYVDGLKFSGGSDSLMPKAFIKQVID 280
>TC76525 similar to GP|15450864|gb|AAK96703.1 Unknown protein {Arabidopsis
thaliana}, partial (75%)
Length = 2266
Score = 31.2 bits (69), Expect = 1.5
Identities = 21/73 (28%), Positives = 32/73 (43%), Gaps = 1/73 (1%)
Frame = +1
Query: 631 VDYIYEDELNHFVHGGA-LSELIVAFSREGPTKEYVQHKMIEKASDIWNMISQGAYIYVC 689
VD + L+ +H L+++ GPT + + +S W GA +Y
Sbjct: 1360 VDVVCLKNLDSLLHAMPNLNQIHALIEHYGPTVYFHPDEKYMPSSVSW-FFKNGAILYTA 1536
Query: 690 GDAKGMAKDVHRT 702
G+AKG A D H T
Sbjct: 1537 GNAKGKAIDYHGT 1575
>TC77686 similar to GP|16649123|gb|AAL24413.1 unknown protein {Arabidopsis
thaliana}, partial (49%)
Length = 1007
Score = 30.0 bits (66), Expect = 3.3
Identities = 15/55 (27%), Positives = 29/55 (52%), Gaps = 5/55 (9%)
Frame = -1
Query: 311 DQLDATVDEKKQLNGNGHAVVDAHHPVRANVAVR-----KELHTPASDRSCTHLE 360
D D V+E+ ++ G VV A ++ ++ ++ ++LH P + SC HL+
Sbjct: 179 DDDDVLVEEEGKIEKEGRVVVVAEDELKDSIIIKIDSQSQKLHKPKYNTSCLHLK 15
>TC93068 similar to GP|3170523|gb|AAC18088.1| coatomer alpha subunit
{Aspergillus nidulans}, partial (8%)
Length = 700
Score = 30.0 bits (66), Expect = 3.3
Identities = 19/76 (25%), Positives = 29/76 (38%)
Frame = -2
Query: 494 SSAKPPIGVFFASVAPRLQPRYYSISSSPRVAPSRIHVTCALVHDKMPTGRIHQGVCSTW 553
++A +G + PR P Y + +S + A +R C + P R C TW
Sbjct: 576 NAAPSSLGGTLHHLYPRTNPHYSLLPASTQKASARCRA-CQPRTPRTPCKRCPCTWCGTW 400
Query: 554 MKNSAPLEKSQDCSWA 569
+ L C WA
Sbjct: 399 SRQMRTLRTQCRCKWA 352
>AW693611 weakly similar to GP|20197699|gb| putative LRR receptor protein
kinase {Arabidopsis thaliana}, partial (8%)
Length = 421
Score = 29.3 bits (64), Expect = 5.7
Identities = 18/66 (27%), Positives = 31/66 (46%)
Frame = +1
Query: 429 RTALAKYADVLSSPKKSALLALAAHASDPSEADRLRHLASPAGKDEYAEWVIASQRSLLE 488
R++L Y DVL ++ +++ +DP++ L L + G WV + RSL
Sbjct: 148 RSSLKIYGDVLFGFLLFCIIQISSSLTDPTDVAALNSLHTSLGSPLLPGWVSSRWRSLRR 327
Query: 489 VMAEFS 494
+A S
Sbjct: 328 RVARHS 345
>TC84596 weakly similar to GP|17064866|gb|AAL32587.1 Unknown protein
{Arabidopsis thaliana}, partial (40%)
Length = 863
Score = 29.3 bits (64), Expect = 5.7
Identities = 19/56 (33%), Positives = 26/56 (45%)
Frame = +2
Query: 413 PLGGSSLPPPFPPCTLRTALAKYADVLSSPKKSALLALAAHASDPSEADRLRHLAS 468
PLG PPP PP T ++ K S P K + +A +D S L +L+S
Sbjct: 35 PLGKPLPPPPLPPLT-SSSTNKNVSATSPPTKKTHHSTSAPMADASSLRPLPYLSS 199
>TC78441 similar to GP|16226543|gb|AAL16196.1 AT3g15000/K15M2_14
{Arabidopsis thaliana}, partial (43%)
Length = 1476
Score = 28.5 bits (62), Expect = 9.7
Identities = 15/46 (32%), Positives = 26/46 (55%), Gaps = 2/46 (4%)
Frame = +3
Query: 377 GVYC-ENLSDTVEEAERILGLSPDTYFSVHTDDEDGKP-LGGSSLP 420
G C E LS ++E ++ + PD+Y +V D G+P + G ++P
Sbjct: 465 GALCSEELSYKLKELPKVRWVLPDSYLNVREKDYGGEPFINGQAVP 602
>TC80017 similar to GP|10177301|dbj|BAB10562. gene_id:MDC12.17~unknown
protein {Arabidopsis thaliana}, partial (45%)
Length = 879
Score = 28.5 bits (62), Expect = 9.7
Identities = 14/26 (53%), Positives = 14/26 (53%)
Frame = +2
Query: 407 DDEDGKPLGGSSLPPPFPPCTLRTAL 432
DDED S LPPP PP T T L
Sbjct: 95 DDEDDNNHSSSLLPPPLPPHTEITRL 172
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.318 0.135 0.400
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 21,030,149
Number of Sequences: 36976
Number of extensions: 293256
Number of successful extensions: 1518
Number of sequences better than 10.0: 38
Number of HSP's better than 10.0 without gapping: 1484
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1509
length of query: 736
length of database: 9,014,727
effective HSP length: 103
effective length of query: 633
effective length of database: 5,206,199
effective search space: 3295523967
effective search space used: 3295523967
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 62 (28.5 bits)
Medicago: description of AC135801.5


