

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC135799.3 + phase: 0
(242 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
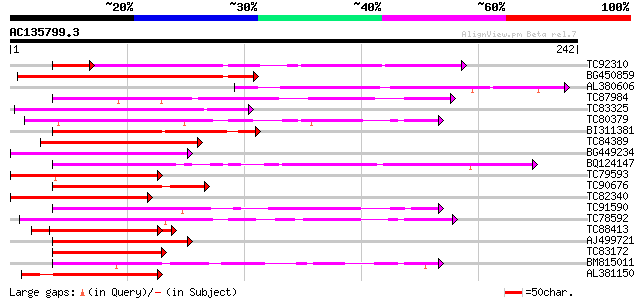
Score E
Sequences producing significant alignments: (bits) Value
TC92310 weakly similar to PIR|T47778|T47778 hypothetical protein... 94 1e-21
BG450859 weakly similar to GP|21553400|gb| unknown {Arabidopsis ... 85 3e-17
AL380606 80 5e-16
TC87984 weakly similar to GP|10177368|dbj|BAB10659. contains sim... 75 3e-14
TC83325 similar to PIR|D84455|D84455 hypothetical protein At2g04... 73 1e-13
TC80379 similar to GP|9802755|gb|AAF99824.1| Unknown protein {Ar... 71 4e-13
BI311381 weakly similar to PIR|T46128|T461 hypothetical protein ... 70 7e-13
TC84389 weakly similar to GP|10177368|dbj|BAB10659. contains sim... 69 2e-12
BG449234 weakly similar to GP|8979942|gb|A Contains a F-box PF|0... 61 3e-10
BQ124147 similar to GP|10177836|dbj emb|CAB62440.1~gene_id:MCD7.... 60 6e-10
TC79593 similar to GP|10177368|dbj|BAB10659. contains similarity... 60 6e-10
TC90676 similar to GP|6503308|gb|AAF14684.1| Contains PF|00646 F... 58 3e-09
TC82340 similar to GP|10178240|dbj|BAB11672. gene_id:MDJ22.8~unk... 57 5e-09
TC91590 weakly similar to GP|10178235|dbj|BAB11667. gene_id:MDJ2... 57 6e-09
TC78592 similar to GP|9802755|gb|AAF99824.1| Unknown protein {Ar... 56 1e-08
TC88413 weakly similar to GP|10176786|dbj|BAB09900. gene_id:MIK1... 56 1e-08
AJ499721 weakly similar to PIR|T48291|T482 hypothetical protein ... 56 1e-08
TC83172 weakly similar to GP|20042907|gb|AAM08735.1 Hypothetical... 55 2e-08
BM815011 similar to PIR|T49178|T491 hypothetical protein T20N10.... 54 7e-08
AL381150 weakly similar to GP|14028988|gb| Unknown protein {Oryz... 54 7e-08
>TC92310 weakly similar to PIR|T47778|T47778 hypothetical protein F17J16.10
- Arabidopsis thaliana, partial (8%)
Length = 1019
Score = 94.0 bits (232), Expect(2) = 1e-21
Identities = 61/161 (37%), Positives = 92/161 (56%)
Frame = +2
Query: 35 LVSTKEAVATSILSKRWNNLWLSLPNIDFNNIKIDSIESNSKFIDSVYSVLVSPDTAIGG 94
+ +TK+AV TS+LSKRW NLW +P +DF++IK+ ES F +V++ +T
Sbjct: 86 MCATKDAVTTSVLSKRWTNLWCFVPFLDFSDIKLADHESFLWFNQFFCTVMLFRETY--A 259
Query: 95 SHFINRFCLDVQFRNGNPLYGYDNPHLLYKRSCHNVVKWVNLVVQRRLKYLRLNLRLGYD 154
S+ I+ F LD+Q+ HL Y S + W+ L+ +R +KYL L+L
Sbjct: 260 SNSIDSFSLDIQY-----------AHLNY-FSILRLNNWLYLLGKRNVKYLNLHLNFLNA 403
Query: 155 LHLDVDDYDNSYLPKLPITIFTCRTLVSLDLHHFSVKGVCY 195
L +DV + + PKLP TIFTC+TLV L++ F+ KG +
Sbjct: 404 LLVDV-QHRKTLTPKLPSTIFTCKTLVVLNISWFAFKGFSF 523
Score = 26.2 bits (56), Expect(2) = 1e-21
Identities = 10/18 (55%), Positives = 15/18 (82%)
Frame = +1
Query: 19 DKISRLPDDVLHYILSLV 36
D+IS+L D ++H+ILS V
Sbjct: 37 DRISKLDDKIIHHILSYV 90
>BG450859 weakly similar to GP|21553400|gb| unknown {Arabidopsis thaliana},
partial (11%)
Length = 549
Score = 84.7 bits (208), Expect = 3e-17
Identities = 44/103 (42%), Positives = 66/103 (63%)
Frame = +2
Query: 4 QKRRRRSSKASLAVADKISRLPDDVLHYILSLVSTKEAVATSILSKRWNNLWLSLPNIDF 63
++ R S + S A D+I LPD++L +ILS V TK+A TSILSKRW +LW +P +DF
Sbjct: 77 RRNRMSSPENSTATVDRIGTLPDEILIHILSFVPTKQAFTTSILSKRWIHLWRYVPILDF 256
Query: 64 NNIKIDSIESNSKFIDSVYSVLVSPDTAIGGSHFINRFCLDVQ 106
++ +S +F + ++SV+ S +A G+H IN F LD+Q
Sbjct: 257 TETNLEDRDSVIRFEEFIFSVIRSRHSA--GNHSINTFILDIQ 379
>AL380606
Length = 437
Score = 80.5 bits (197), Expect = 5e-16
Identities = 59/149 (39%), Positives = 74/149 (49%), Gaps = 6/149 (4%)
Frame = +1
Query: 97 FINRFCLDVQFRNGNPLYGYDNPHLLYKRSCHNVVKWVNLVVQRRLKYLRLNLRLGYDLH 156
FI+ F L++Q Y + HL Y + N+ KWVN VVQR LKYL L LR+ H
Sbjct: 1 FIDSFHLNIQ---------YSDSHLAYNKEFPNLTKWVNSVVQRGLKYLHLRLRVPLPDH 153
Query: 157 LDVDDYDNSYLPKLPITIFTCRTLVSLDLHHFSVKGVCYR---FLLKALSTSNSLRLDTF 213
Y PKLPI+IFTC+TLVSL+L F V G + F +L T SLRL F
Sbjct: 154 FS----GYPYFPKLPISIFTCKTLVSLNLSWFRVDGFSFTSVGFEFPSLKTL-SLRLIKF 318
Query: 214 KLYRSIYQVRF---IMNTMHVIYFLFLSE 239
R + I+ +HV F E
Sbjct: 319 SEVRDFMLLLAGCPILEDLHVALVFFYYE 405
>TC87984 weakly similar to GP|10177368|dbj|BAB10659. contains similarity to
heat shock transcription factor HSF30~gene_id:K16L22.13,
partial (5%)
Length = 696
Score = 74.7 bits (182), Expect = 3e-14
Identities = 45/176 (25%), Positives = 89/176 (50%), Gaps = 4/176 (2%)
Frame = +3
Query: 19 DKISRLPDDVLHYILSLVSTKEAVATS-ILSKRWNNLWLSLPNIDF---NNIKIDSIESN 74
D+I++LP+ +LH+ILS + TK V T+ ++S++W NLW +L +DF ++ +I +++
Sbjct: 105 DRINQLPNSLLHHILSFLPTKTCVQTTPLVSRKWRNLWKNLEALDFCDSSSPQIYEFDND 284
Query: 75 SKFIDSVYSVLVSPDTAIGGSHFINRFCLDVQFRNGNPLYGYDNPHLLYKRSCHNVVKWV 134
+F+ ++SV V+ + S + +FCL +P Y H++ W+
Sbjct: 285 EQFL--IFSVFVNTVLTLRKSRVVRKFCLSCYHVQLDPFYN------------HSIDTWI 422
Query: 135 NLVVQRRLKYLRLNLRLGYDLHLDVDDYDNSYLPKLPITIFTCRTLVSLDLHHFSV 190
N + L+ L L + ++P+++F+C LVSL + + +
Sbjct: 423 NATIGPNLEEFHLTLLTAAGFN------------RVPLSLFSCPNLVSLSFNDYII 554
>TC83325 similar to PIR|D84455|D84455 hypothetical protein At2g04230
[imported] - Arabidopsis thaliana, partial (4%)
Length = 755
Score = 72.8 bits (177), Expect = 1e-13
Identities = 40/102 (39%), Positives = 61/102 (59%)
Frame = +2
Query: 3 RQKRRRRSSKASLAVADKISRLPDDVLHYILSLVSTKEAVATSILSKRWNNLWLSLPNID 62
+ K+RR+++ D+IS LPD +L +ILS TK AV T+ILS RWNNLW LP +
Sbjct: 104 QMKKRRKNNHNHTENKDRISDLPDCILFHILSS*ETKHAVQTTILSTRWNNLWKRLPTLR 283
Query: 63 FNNIKIDSIESNSKFIDSVYSVLVSPDTAIGGSHFINRFCLD 104
N+ +++S +KF+ V S L TA+ +F + C++
Sbjct: 284 LNSSHFKTLKSFTKFVSRVLS-LRDDSTALHTLYFQRKGCVE 406
>TC80379 similar to GP|9802755|gb|AAF99824.1| Unknown protein {Arabidopsis
thaliana}, partial (5%)
Length = 672
Score = 70.9 bits (172), Expect = 4e-13
Identities = 65/191 (34%), Positives = 91/191 (47%), Gaps = 12/191 (6%)
Frame = +2
Query: 7 RRRSSKASLAVAD----KISRLPDDVLHYILSLVSTKEAVATSILSKRWNNLWLSLPNID 62
R+RS L + D +IS LP V+ ILSL+ KEAV TS+LS W N W +LPN+
Sbjct: 146 RKRSKSTHLTIKDAELDRISCLPGHVIDQILSLMPIKEAVRTSVLSSSWRNKWYTLPNLV 325
Query: 63 FNNIKIDSIES------NSKFIDSVYSVLVSPDTAIGGSHFINRFCLDVQFRNGNPLYGY 116
FN I S N+KF+ V VL+ S IN+F L
Sbjct: 326 FNKHCISVAASKYTSVINNKFLRIVDHVLLQ------HSGPINKFMLS------------ 451
Query: 117 DNPHLLYKRSC--HNVVKWVNLVVQRRLKYLRLNLRLGYDLHLDVDDYDNSYLPKLPITI 174
DN + L R C +V +WV + +R +K L L+ L D Y K+P+++
Sbjct: 452 DNENCLI-RVCDVSDVHRWVLHLSRRSIKELLLHFWL-----------DKRY--KIPLSL 589
Query: 175 FTCRTLVSLDL 185
F+C++L L L
Sbjct: 590 FSCQSLRHLSL 622
>BI311381 weakly similar to PIR|T46128|T461 hypothetical protein T2J13.140 -
Arabidopsis thaliana, partial (12%)
Length = 803
Score = 70.1 bits (170), Expect = 7e-13
Identities = 43/89 (48%), Positives = 55/89 (61%)
Frame = +1
Query: 19 DKISRLPDDVLHYILSLVSTKEAVATSILSKRWNNLWLSLPNIDFNNIKIDSIESNSKFI 78
D IS LPD +L +ILS + T A ATS+LSKRWN+LW S+ + F ++ SN FI
Sbjct: 103 DIISTLPDAILCHILSFLETNYADATSVLSKRWNHLWRSVHTLRFIT-QVTDHNSNHDFI 279
Query: 79 DSVYSVLVSPDTAIGGSHFINRFCLDVQF 107
D VYSVL+S D A+ I F L+V F
Sbjct: 280 DFVYSVLLSRDPALP----IKTFHLEVTF 354
>TC84389 weakly similar to GP|10177368|dbj|BAB10659. contains similarity to
heat shock transcription factor HSF30~gene_id:K16L22.13,
partial (8%)
Length = 692
Score = 68.6 bits (166), Expect = 2e-12
Identities = 34/69 (49%), Positives = 47/69 (67%)
Frame = +3
Query: 14 SLAVADKISRLPDDVLHYILSLVSTKEAVATSILSKRWNNLWLSLPNIDFNNIKIDSIES 73
S++ D+IS LPD ++ +ILS V TK A TSILSKRWN LWLS+ + F + ++ES
Sbjct: 252 SISTEDRISALPDPIIWHILSFVPTKTAAITSILSKRWNPLWLSVLILHFEDETFQNMES 431
Query: 74 NSKFIDSVY 82
S F+ SV+
Sbjct: 432 FSHFMSSVF 458
>BG449234 weakly similar to GP|8979942|gb|A Contains a F-box PF|00646 domain.
{Arabidopsis thaliana}, partial (6%)
Length = 655
Score = 61.2 bits (147), Expect = 3e-10
Identities = 31/78 (39%), Positives = 47/78 (59%)
Frame = +3
Query: 1 MQRQKRRRRSSKASLAVADKISRLPDDVLHYILSLVSTKEAVATSILSKRWNNLWLSLPN 60
M +++R+R K + D+IS L D +L +ILS ++T+EAV T ILSKRW NLW +LP
Sbjct: 102 MGTRRQRKRQPKDDSQIQDRISVLADCLLIHILSFLNTREAVQTCILSKRWINLWKTLPT 281
Query: 61 IDFNNIKIDSIESNSKFI 78
+ + + E F+
Sbjct: 282 LTLDCYQFSDCEIYENFL 335
>BQ124147 similar to GP|10177836|dbj emb|CAB62440.1~gene_id:MCD7.13~similar
to unknown protein {Arabidopsis thaliana}, partial (11%)
Length = 677
Score = 60.5 bits (145), Expect = 6e-10
Identities = 57/211 (27%), Positives = 97/211 (45%), Gaps = 4/211 (1%)
Frame = +1
Query: 19 DKISRLPDDVLHYILSLVSTKEAVATSILSKRWNNLWLSLPNIDFNNIKIDSIESNSKFI 78
D+IS LPDD+L +ILS + TK+A TS+LSKRW NLW +P ++F K ++ SK++
Sbjct: 25 DRISILPDDILVHILSPLPTKQAFVTSVLSKRWKNLWCLVPALEFVGTKTVTV---SKYV 195
Query: 79 DSVYSVLVSPDTAIGGSHFINRFCLDVQFRNGNPLYGYDNPHLLYKRSCHNVVKWVNLVV 138
VS A G+H+++ N +Y + P+ ++ R ++K L +
Sbjct: 196 -------VSRQAA--GNHYMH---------NWERIYP-EFPNTIFTRETLVILKLSELFM 318
Query: 139 QRRLKYLRLNLRLGYDLHLDVDDYDNSYLPKLPITIFTCRTLVSLDLHHFSVKGVCY--- 195
+ L LHL D + L + C L L L+ S + +
Sbjct: 319 GSGFCNYLIRLPSLKTLHL--KDIEFQQYGDLDCLLEGCPVLEDLQLYDISYVSLLFSNA 492
Query: 196 -RFLLKALSTSNSLRLDTFKLYRSIYQVRFI 225
R L L+ ++ ++ D +++ V F+
Sbjct: 493 CRKTLTKLNRADIIQCDCGVRMKALSNVEFL 585
>TC79593 similar to GP|10177368|dbj|BAB10659. contains similarity to heat
shock transcription factor HSF30~gene_id:K16L22.13,
partial (6%)
Length = 713
Score = 60.5 bits (145), Expect = 6e-10
Identities = 31/66 (46%), Positives = 45/66 (67%), Gaps = 1/66 (1%)
Frame = +1
Query: 1 MQRQKRRRRSSKASLAVA-DKISRLPDDVLHYILSLVSTKEAVATSILSKRWNNLWLSLP 59
M R+ R+RR + S+ D+IS LPD ++H+ILS + +K+A AT++LSKRW LWL
Sbjct: 487 MSRRLRQRRQLQLSIPEEEDRISALPDSLVHHILSFLPSKDATATTVLSKRWKPLWLLEL 666
Query: 60 NIDFNN 65
I FN+
Sbjct: 667 IIHFNH 684
>TC90676 similar to GP|6503308|gb|AAF14684.1| Contains PF|00646 F-box
domain. ESTs gb|AA586135 gb|Z26675 gb|AI993239 and
gb|AA585907 come from, partial (5%)
Length = 1119
Score = 58.2 bits (139), Expect = 3e-09
Identities = 31/67 (46%), Positives = 43/67 (63%)
Frame = +3
Query: 19 DKISRLPDDVLHYILSLVSTKEAVATSILSKRWNNLWLSLPNIDFNNIKIDSIESNSKFI 78
D IS LP+ +L+YILS +ST+EAV TSIL+ +W LW LP DF + S + NSK
Sbjct: 204 DMISDLPEGILYYILSRLSTEEAVRTSILATKWRYLWTQLPVFDFRD---SSPKRNSKSP 374
Query: 79 DSVYSVL 85
D + ++
Sbjct: 375 DCLVDLV 395
>TC82340 similar to GP|10178240|dbj|BAB11672. gene_id:MDJ22.8~unknown
protein {Arabidopsis thaliana}, partial (9%)
Length = 641
Score = 57.4 bits (137), Expect = 5e-09
Identities = 28/61 (45%), Positives = 38/61 (61%)
Frame = +3
Query: 1 MQRQKRRRRSSKASLAVADKISRLPDDVLHYILSLVSTKEAVATSILSKRWNNLWLSLPN 60
M R K +RRS + D++S LPD +L YILS + T+ AV T ILS RW +LW +P
Sbjct: 270 MFRDKMKRRSRSENEESEDRLSDLPDCILIYILSFLKTENAVQTCILSTRWKHLWKRIPK 449
Query: 61 I 61
+
Sbjct: 450 L 452
>TC91590 weakly similar to GP|10178235|dbj|BAB11667. gene_id:MDJ22.3~unknown
protein {Arabidopsis thaliana}, partial (9%)
Length = 712
Score = 57.0 bits (136), Expect = 6e-09
Identities = 41/170 (24%), Positives = 79/170 (46%), Gaps = 3/170 (1%)
Frame = +2
Query: 19 DKISRLPDDVLHYILSLVSTKEAVATSILSKRWNNLWLSLPNIDFNNIKIDSIE---SNS 75
D IS L + +L ILS + +AV+TS+LS+RW ++W + N+DF++ + S +
Sbjct: 164 DMISTLHESILSQILSFIPIVDAVSTSVLSRRWVDVWKCITNLDFDDSLLGSRKKRMQKE 343
Query: 76 KFIDSVYSVLVSPDTAIGGSHFINRFCLDVQFRNGNPLYGYDNPHLLYKRSCHNVVKWVN 135
+F++ V VL+ HF N + + + ++ + +W++
Sbjct: 344 QFVNFVEKVLI---------HFTN-----------SSIQSFSLSLTSHQYDASKLSEWIS 463
Query: 136 LVVQRRLKYLRLNLRLGYDLHLDVDDYDNSYLPKLPITIFTCRTLVSLDL 185
+++RR++ L + D +LP ++F C +LV L L
Sbjct: 464 FILERRVQKLHIQYA------------DKVFLPS--DSLFRCNSLVDLTL 571
>TC78592 similar to GP|9802755|gb|AAF99824.1| Unknown protein {Arabidopsis
thaliana}, partial (8%)
Length = 1909
Score = 56.2 bits (134), Expect = 1e-08
Identities = 51/193 (26%), Positives = 93/193 (47%), Gaps = 6/193 (3%)
Frame = +2
Query: 5 KRRRRSSKASLAVADKISRLPDDVLHYILSLVSTKEAVATSILSKRWNNLWLSLPNIDFN 64
+ R+S K + +D+IS LPD ++ + LS +S KEA TS+LS +W W + P++ F+
Sbjct: 419 EENRQSLKMTDKESDRISCLPDHIIDHFLSRLSIKEAGRTSVLSNKWRKKWSTQPDLVFD 598
Query: 65 N------IKIDSIESNSKFIDSVYSVLVSPDTAIGGSHFINRFCLDVQFRNGNPLYGYDN 118
+ D +KF+ +V VL+ S IN+F + +G D
Sbjct: 599 KQCVSAAVSEDPSVIENKFVRTVDHVLLL------HSGPINKFKISD--------FGCDF 736
Query: 119 PHLLYKRSCHNVVKWVNLVVQRRLKYLRLNLRLGYDLHLDVDDYDNSYLPKLPITIFTCR 178
++ S +V +W+ + +R +K L L++ ++ Y +P +F+C+
Sbjct: 737 IGMI---SVADVDRWILHLTRRSIKNLELDIW-----------FEQRY--NIPWCLFSCQ 868
Query: 179 TLVSLDLHHFSVK 191
+L L+L S+K
Sbjct: 869 SLHHLNLFCSSLK 907
>TC88413 weakly similar to GP|10176786|dbj|BAB09900.
gene_id:MIK19.29~pir||T01344~similar to unknown protein
{Arabidopsis thaliana}, partial (10%)
Length = 1288
Score = 56.2 bits (134), Expect = 1e-08
Identities = 26/62 (41%), Positives = 42/62 (66%)
Frame = +1
Query: 10 SSKASLAVADKISRLPDDVLHYILSLVSTKEAVATSILSKRWNNLWLSLPNIDFNNIKID 69
S++ +A D++S LPD++L ILS + ++E +TS+LSKRW +WL +PN D +I D
Sbjct: 91 SNQTMMAEEDRMSGLPDELLCQILSFLPSEEIPSTSLLSKRWRRVWLGMPNADRISILPD 270
Query: 70 SI 71
+
Sbjct: 271 EL 276
Score = 43.9 bits (102), Expect = 6e-05
Identities = 21/48 (43%), Positives = 34/48 (70%)
Frame = +1
Query: 18 ADKISRLPDDVLHYILSLVSTKEAVATSILSKRWNNLWLSLPNIDFNN 65
AD+IS LPD++L ILSL+ K+ + TS+LS+RW +L + I+ ++
Sbjct: 244 ADRISILPDELLCRILSLLPAKQIMVTSLLSRRWRSLRPRMTEINIDD 387
>AJ499721 weakly similar to PIR|T48291|T482 hypothetical protein F9G14.10 -
Arabidopsis thaliana, partial (8%)
Length = 502
Score = 56.2 bits (134), Expect = 1e-08
Identities = 22/60 (36%), Positives = 39/60 (64%)
Frame = +1
Query: 19 DKISRLPDDVLHYILSLVSTKEAVATSILSKRWNNLWLSLPNIDFNNIKIDSIESNSKFI 78
D++ +PD ++H+ILS + T++A+ T +LSKRW +W S+P + FN+ + KF+
Sbjct: 298 DRLGDMPDCLIHHILSFMETRDAIRTCVLSKRWRYIWKSIPCLYFNSKSFTRLVDFKKFV 477
>TC83172 weakly similar to GP|20042907|gb|AAM08735.1 Hypothetical protein
{Oryza sativa (japonica cultivar-group)}, partial (31%)
Length = 1097
Score = 55.1 bits (131), Expect = 2e-08
Identities = 26/49 (53%), Positives = 34/49 (69%)
Frame = +2
Query: 19 DKISRLPDDVLHYILSLVSTKEAVATSILSKRWNNLWLSLPNIDFNNIK 67
D++S LPD V+ +ILS + T +AV T ILSKRWNNLW LP ++ K
Sbjct: 206 DRLSDLPDCVILHILSFLDTIDAVQTCILSKRWNNLWKYLPTLNIKFFK 352
>BM815011 similar to PIR|T49178|T491 hypothetical protein T20N10.300 -
Arabidopsis thaliana, partial (4%)
Length = 628
Score = 53.5 bits (127), Expect = 7e-08
Identities = 53/169 (31%), Positives = 76/169 (44%), Gaps = 2/169 (1%)
Frame = +2
Query: 19 DKISRLPDDVLHYILSLVSTKEAVAT-SILSKRWNNLWLSLPNIDFNNIKIDSIESNSKF 77
D++S LPD VL +I+S + T +VAT LS RW +LW L F++ D S +F
Sbjct: 5 DRLSSLPDSVLCHIMSFLPTITSVATIDRLSSRWRHLWKDLQVFRFSS---DDSYSFKRF 175
Query: 78 IDSVYSVLVSPDTAIGGSHFINRFCLDVQFRNGNPLYGYDNPHLLYKRSCHNVVKWVNLV 137
V +VL A+ S I +F +FR Y+ C + WV+
Sbjct: 176 AFFVNAVL-----ALRRSRHIRKFDFTFEFRE-------------YEVEC--IEMWVHAA 295
Query: 138 VQRRLKYLRLNLRLGYDLHLDVDDYDNSYLPKLPITIFT-CRTLVSLDL 185
+ RL+ LD+D Y+ LP++ FT C LVSL L
Sbjct: 296 IGPRLE------------ELDLDIYEADI--NLPLSFFTSCNNLVSLRL 400
>AL381150 weakly similar to GP|14028988|gb| Unknown protein {Oryza sativa},
partial (10%)
Length = 469
Score = 53.5 bits (127), Expect = 7e-08
Identities = 25/60 (41%), Positives = 40/60 (66%)
Frame = +1
Query: 6 RRRRSSKASLAVADKISRLPDDVLHYILSLVSTKEAVATSILSKRWNNLWLSLPNIDFNN 65
+R+R K+ D+IS LPD VL +I+ ++ K++V T +LSKRW +LW SL N+ ++
Sbjct: 76 KRKRQKKS-----DRISELPDHVLLHIIEFMNIKQSVQTCVLSKRWKSLWKSLANLTLHH 240
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.326 0.141 0.423
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 8,840,302
Number of Sequences: 36976
Number of extensions: 137612
Number of successful extensions: 1191
Number of sequences better than 10.0: 109
Number of HSP's better than 10.0 without gapping: 1167
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1173
length of query: 242
length of database: 9,014,727
effective HSP length: 93
effective length of query: 149
effective length of database: 5,575,959
effective search space: 830817891
effective search space used: 830817891
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.7 bits)
S2: 57 (26.6 bits)
Medicago: description of AC135799.3


