

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
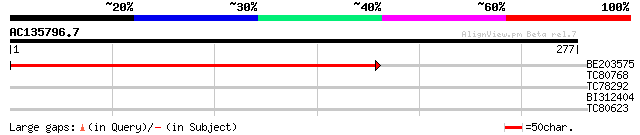
Query= AC135796.7 - phase: 0
(277 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
BE203575 weakly similar to GP|12324017|gb hypothetical protein; ... 368 e-102
TC80768 similar to GP|15081692|gb|AAK82501.1 At1g50380/F14I3_27 ... 28 3.0
TC78292 weakly similar to PIR|T09617|T09617 isoliquiritigenin 2'... 28 5.0
BI312404 27 8.6
TC80623 27 8.6
>BE203575 weakly similar to GP|12324017|gb hypothetical protein; 1267-3026
{Arabidopsis thaliana}, partial (50%)
Length = 632
Score = 368 bits (944), Expect = e-102
Identities = 181/181 (100%), Positives = 181/181 (100%)
Frame = +1
Query: 1 MKAKLIVFPIRGRNWCFTRSIDHTLPASSSTADFSQSPSTLKQLWKSINTGDKPFNAKAE 60
MKAKLIVFPIRGRNWCFTRSIDHTLPASSSTADFSQSPSTLKQLWKSINTGDKPFNAKAE
Sbjct: 88 MKAKLIVFPIRGRNWCFTRSIDHTLPASSSTADFSQSPSTLKQLWKSINTGDKPFNAKAE 267
Query: 61 LFTDYVANKMNNGWVTLENAPDGSFKKKIHGLGLWLLSRVKPSEIFLKSISKDVTGVEVV 120
LFTDYVANKMNNGWVTLENAPDGSFKKKIHGLGLWLLSRVKPSEIFLKSISKDVTGVEVV
Sbjct: 268 LFTDYVANKMNNGWVTLENAPDGSFKKKIHGLGLWLLSRVKPSEIFLKSISKDVTGVEVV 447
Query: 121 YPSSMNARLVRRRLRHIAMRGTIIHRKFFYGSVSLIPLSSAFSILPLPNVPFFWILFRSY 180
YPSSMNARLVRRRLRHIAMRGTIIHRKFFYGSVSLIPLSSAFSILPLPNVPFFWILFRSY
Sbjct: 448 YPSSMNARLVRRRLRHIAMRGTIIHRKFFYGSVSLIPLSSAFSILPLPNVPFFWILFRSY 627
Query: 181 S 181
S
Sbjct: 628 S 630
>TC80768 similar to GP|15081692|gb|AAK82501.1 At1g50380/F14I3_27
{Arabidopsis thaliana}, partial (35%)
Length = 831
Score = 28.5 bits (62), Expect = 3.0
Identities = 29/103 (28%), Positives = 49/103 (47%), Gaps = 4/103 (3%)
Frame = +3
Query: 156 IPLSSAFSILPLPNVPFFWILFRSYSHWRALQGSEKLFQLVSDGSQSSNTYSGKKETEHE 215
IPL+ S+L L ++PF S+ H R + S Q +S SQS K+ EH+
Sbjct: 123 IPLT--VSVLTL-SLPFSTTAAFSHHHRRPMTTSPPQSQSLSPHSQSPLPPPVAKKVEHK 293
Query: 216 DSENESLGLDEPHWVLTPSK---ELENIVRQEDG-NDGLSRGT 254
+ +D +W+ S+ ++ + +RQE+ D + GT
Sbjct: 294 MELFNDVRIDNYYWLRDDSRSDPDVLSYLRQENAYTDSVMSGT 422
>TC78292 weakly similar to PIR|T09617|T09617 isoliquiritigenin
2'-O-methyltransferase - alfalfa, partial (86%)
Length = 1327
Score = 27.7 bits (60), Expect = 5.0
Identities = 17/60 (28%), Positives = 34/60 (56%), Gaps = 6/60 (10%)
Frame = +1
Query: 62 FTDYVANK-MNNGWVTLENA---PDGSFKKKIHGLGLWLLSRVKP--SEIFLKSISKDVT 115
FT ++++ ++ W+ + A P+G KK+HG+ ++ + P +EIF KS++ T
Sbjct: 466 FTSFLSHPALSMVWLNFKEAIIDPEGDLFKKVHGITMFEYFKKDPQINEIFNKSMTDTCT 645
>BI312404
Length = 759
Score = 26.9 bits (58), Expect = 8.6
Identities = 11/21 (52%), Positives = 16/21 (75%)
Frame = +3
Query: 128 RLVRRRLRHIAMRGTIIHRKF 148
++VR RLR I +RG +HR+F
Sbjct: 150 KVVRTRLRFIMLRGNWMHREF 212
>TC80623
Length = 1125
Score = 26.9 bits (58), Expect = 8.6
Identities = 18/43 (41%), Positives = 22/43 (50%)
Frame = +1
Query: 217 SENESLGLDEPHWVLTPSKELENIVRQEDGNDGLSRGTIEEIC 259
S+N SL L+ L S LEN +E+ DGL T EIC
Sbjct: 394 SQNVSLPLNCEQQSLGASVTLEN---KENSQDGLGNATRSEIC 513
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.317 0.134 0.400
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 9,127,478
Number of Sequences: 36976
Number of extensions: 128058
Number of successful extensions: 731
Number of sequences better than 10.0: 10
Number of HSP's better than 10.0 without gapping: 728
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 730
length of query: 277
length of database: 9,014,727
effective HSP length: 95
effective length of query: 182
effective length of database: 5,502,007
effective search space: 1001365274
effective search space used: 1001365274
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 58 (26.9 bits)
Medicago: description of AC135796.7


