

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC135566.1 + phase: 0 /pseudo
(437 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
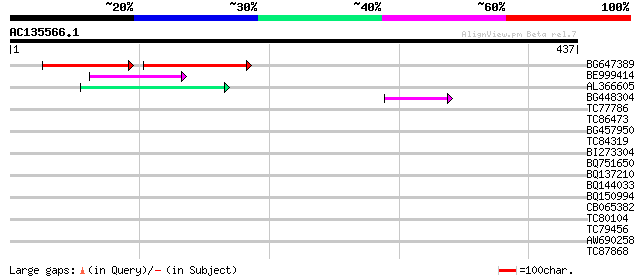
Score E
Sequences producing significant alignments: (bits) Value
BG647389 108 1e-41
BE999414 48 7e-06
AL366605 47 2e-05
BG448304 46 2e-05
TC77786 35 0.074
TC86473 similar to PIR|E84828|E84828 probable WD-40 repeat prote... 33 0.22
BG457950 similar to GP|4996286|dbj lipoyltransferase {Arabidopsi... 32 0.48
TC84319 GP|10440402|dbj|BAB15734. FLJ00034 protein {Homo sapiens... 32 0.63
BI273304 weakly similar to GP|10177311|dbj gene_id:MDC12.27~unkn... 31 1.1
BQ751650 weakly similar to PIR|A32567|A32 stress-induced protein... 30 2.4
BQ137210 similar to SP|P09789|GRP1_ Glycine-rich cell wall struc... 29 3.1
BQ144033 similar to GP|6523547|emb| hydroxyproline-rich glycopro... 29 4.1
BQ150994 similar to GP|22122173|dbj preproneuropeptide B {Homo s... 29 4.1
CB065382 similar to PIR|T09671|T096 RPE15 protein - alfalfa (fra... 28 7.0
TC80104 similar to GP|20197699|gb|AAD20910.3 putative LRR recept... 28 7.0
TC79456 similar to PIR|T51518|T51518 ubiquitin-fusion degradatio... 28 7.0
AW690258 GP|22900886|gb amt2-like protein {Medicago truncatula},... 28 9.1
TC87868 similar to PIR|T05112|T05112 splicing factor 9G8-like SR... 28 9.1
>BG647389
Length = 718
Score = 108 bits (271), Expect(2) = 1e-41
Identities = 52/83 (62%), Positives = 64/83 (76%)
Frame = +1
Query: 104 DAALRWYMSLPCFSIMGYQDMIKKFTQQFSGTRHRKVLSTSLFNVRQGPSESLREYLARF 163
+A +WYM+LP FSI GYQ+M K QFS ++H KV +T+LFNVRQ P+ESL EYL F
Sbjct: 454 EATQQWYMNLPRFSITGYQNMTHKLVHQFSASKHCKVSTTNLFNVRQDPNESLPEYLV*F 633
Query: 164 NDSTIKVSNPNQEVFVGAFQNGL 186
N++TIKV NP QE+F GAFQNGL
Sbjct: 634 NNATIKVVNPKQELFAGAFQNGL 702
Score = 79.3 bits (194), Expect(2) = 1e-41
Identities = 36/70 (51%), Positives = 46/70 (65%)
Frame = +2
Query: 26 QQQEKTDSHHGDLDEPEPQPFSAKIWNAPVPDNFKPPHLPTFDGRGDPSEHVTVFNTRMS 85
QQ E+ + H D+DE +PQP S IWNAPV +N K L +FD + DP EH T FNT+M+
Sbjct: 224 QQHEQDELHREDVDELDPQPLSMDIWNAPVLENLKSLSLLSFDDKSDPVEHATTFNTQMA 403
Query: 86 VYGVADSLKC 95
V G SL+C
Sbjct: 404 VIGALKSLRC 433
>BE999414
Length = 613
Score = 48.1 bits (113), Expect = 7e-06
Identities = 23/75 (30%), Positives = 39/75 (51%)
Frame = +3
Query: 62 PHLPTFDGRGDPSEHVTVFNTRMSVYGVADSLKCKLLAGTFADAALRWYMSLPCFSIMGY 121
P + ++DG DP EH+ ++ V ++KCKL T A+ W+ +L SI +
Sbjct: 3 PQMDSYDGT*DPDEHMENIEVVLTYRSVRGAVKCKLFVTTLRRGAMTWFKNLRRNSIGSW 182
Query: 122 QDMIKKFTQQFSGTR 136
D+ +FT F+ +R
Sbjct: 183 GDLCHEFTTHFTVSR 227
>AL366605
Length = 422
Score = 46.6 bits (109), Expect = 2e-05
Identities = 29/115 (25%), Positives = 45/115 (38%)
Frame = -3
Query: 55 VPDNFKPPHLPTFDGRGDPSEHVTVFNTRMSVYGVADSLKCKLLAGTFADAALRWYMSLP 114
VP FK P ++G P H+ + +M Y DSL + + A WY SL
Sbjct: 345 VPKKFKIPDFDRYNGLTCPQNHIIKYVRKMGNYKDNDSLMIHCFQDSLMEDAAEWYTSLS 166
Query: 115 CFSIMGYQDMIKKFTQQFSGTRHRKVLSTSLFNVRQGPSESLREYLARFNDSTIK 169
I + ++ F + K L ++ Q ES REY R+ + +
Sbjct: 165 KNDIHTFDELAAAFKSHYGFNTRLKPNREFLRSLSQKKEESFREYAQRWRGAAAR 1
>BG448304
Length = 637
Score = 46.2 bits (108), Expect = 2e-05
Identities = 21/52 (40%), Positives = 27/52 (51%)
Frame = +2
Query: 290 PPPFSRARVMGQDKDAWCKYHLVQGHNTDDCVHLKREIEKLLLSGKLRGYAK 341
PP G DK +C++H H TDDC+HLK IE L+ G+L K
Sbjct: 62 PPKTPTQENKGTDKTKYCRFHKCHRHLTDDCIHLKDTIEILIQRGRLNQLTK 217
>TC77786
Length = 1094
Score = 34.7 bits (78), Expect = 0.074
Identities = 20/54 (37%), Positives = 28/54 (51%), Gaps = 1/54 (1%)
Frame = +3
Query: 302 DKDAWCKYHLVQGHNTDDCVHLKREIEKLLLSGKL-RGYAKERRHNERQEDKPN 354
DK C YH +GH+T++C L+ IE L+ KL R K R N ++ N
Sbjct: 810 DKAK*CCYHRSRGHDTEECFQLREAIEDLIKKVKLGRFIDKGHRKNSPRKSP*N 971
>TC86473 similar to PIR|E84828|E84828 probable WD-40 repeat protein [imported]
- Arabidopsis thaliana, partial (68%)
Length = 2654
Score = 33.1 bits (74), Expect = 0.22
Identities = 13/45 (28%), Positives = 27/45 (59%), Gaps = 2/45 (4%)
Frame = +2
Query: 242 YVPPNRDRGTFKKPYERNQNRYVPEHFTPLNTRP--ERILKEVFE 284
Y+P + +++ YE ++ +++P+ FT + + P E +KE FE
Sbjct: 989 YIPTQEEINSYQLMYEEDRPKFIPQRFTSMRSIPAYENAMKESFE 1123
>BG457950 similar to GP|4996286|dbj lipoyltransferase {Arabidopsis thaliana},
partial (77%)
Length = 686
Score = 32.0 bits (71), Expect = 0.48
Identities = 17/57 (29%), Positives = 28/57 (48%), Gaps = 1/57 (1%)
Frame = -3
Query: 252 FKKPYERNQNRYVP-EHFTPLNTRPERILKEVFESKIIPPPPFSRARVMGQDKDAWC 307
F P + + + P + FTP I+ +F + + P P SR+ ++G K AWC
Sbjct: 570 FLSPTQTPVSHFFPGQAFTPYRPASSIIVDSIFSTNFLAPSPVSRSDMIGY-KMAWC 403
>TC84319 GP|10440402|dbj|BAB15734. FLJ00034 protein {Homo sapiens}, partial
(0%)
Length = 692
Score = 31.6 bits (70), Expect = 0.63
Identities = 19/54 (35%), Positives = 25/54 (46%), Gaps = 6/54 (11%)
Frame = -1
Query: 22 IQQVQQQEKTDSHH------GDLDEPEPQPFSAKIWNAPVPDNFKPPHLPTFDG 69
+QQ QQQ+ H GDL E Q + +AP+P + PP L F G
Sbjct: 341 VQQPQQQQHHQRPHPSALRPGDLTAAESQDLRQQFVSAPIPADQGPPDLLHFQG 180
>BI273304 weakly similar to GP|10177311|dbj gene_id:MDC12.27~unknown protein
{Arabidopsis thaliana}, partial (11%)
Length = 678
Score = 30.8 bits (68), Expect = 1.1
Identities = 21/92 (22%), Positives = 39/92 (41%)
Frame = +3
Query: 170 VSNPNQEVFVGAFQNGLWAGQFNESIAQKPADSMEEIIARAECYVKGEESNAEKKAREAK 229
++NP E+ + + AG F E R + + + ++KAREA
Sbjct: 78 MTNPEDELINRFKKEVIMAGVFQE--------------CRRRRFFETPQDKIKRKAREAS 215
Query: 230 ERGNSGGERRNHYVPPNRDRGTFKKPYERNQN 261
+R +R Y+PP RD+ + P ++ +
Sbjct: 216 KRNR----KRRPYIPPARDKYEYDYPNKKKDD 299
>BQ751650 weakly similar to PIR|A32567|A32 stress-induced protein STI1 -
yeast (Saccharomyces cerevisiae), partial (32%)
Length = 723
Score = 29.6 bits (65), Expect = 2.4
Identities = 22/60 (36%), Positives = 27/60 (44%), Gaps = 5/60 (8%)
Frame = +1
Query: 243 VPPNRDRGTFKKPYERNQNR---YVPEHFTPLNTRPE--RILKEVFESKIIPPPPFSRAR 297
VPP R R T K+P +NR P TP P R + V+ + PP P RAR
Sbjct: 304 VPPTRSRATSKRPLRTTRNRSRSIAPPTSTPSFAPPSATRSSRRVWPTS-TPPRPRRRAR 480
>BQ137210 similar to SP|P09789|GRP1_ Glycine-rich cell wall structural
protein 1 precursor. [Petunia] {Petunia hybrida},
partial (13%)
Length = 990
Score = 29.3 bits (64), Expect = 3.1
Identities = 18/65 (27%), Positives = 30/65 (45%)
Frame = +1
Query: 222 EKKAREAKERGNSGGERRNHYVPPNRDRGTFKKPYERNQNRYVPEHFTPLNTRPERILKE 281
++K R RG GGE R + PP + + T K + NQ H +T+P +
Sbjct: 16 KRKGRRRGGRGG-GGEGRRSHPPPPQTKNTNKTQEQDNQKTQPRPHPGHRSTQPRVVYAG 192
Query: 282 VFESK 286
V+ ++
Sbjct: 193 VYNTQ 207
>BQ144033 similar to GP|6523547|emb| hydroxyproline-rich glycoprotein DZ-HRGP
{Volvox carteri f. nagariensis}, partial (8%)
Length = 1433
Score = 28.9 bits (63), Expect = 4.1
Identities = 16/48 (33%), Positives = 24/48 (49%)
Frame = +1
Query: 215 KGEESNAEKKAREAKERGNSGGERRNHYVPPNRDRGTFKKPYERNQNR 262
KG ES E++ RE +ERG G +RR+ ++ Y+R R
Sbjct: 763 KGSES--ERRGREYEERGRRGAKRRSQRQARREREEEVREQYQRYARR 900
>BQ150994 similar to GP|22122173|dbj preproneuropeptide B {Homo sapiens},
partial (15%)
Length = 1036
Score = 28.9 bits (63), Expect = 4.1
Identities = 15/40 (37%), Positives = 23/40 (57%), Gaps = 2/40 (5%)
Frame = +1
Query: 225 AREAKERG--NSGGERRNHYVPPNRDRGTFKKPYERNQNR 262
A+ K RG N+G +R+N PN+++ T KK R + R
Sbjct: 448 AQRDKRRGAQNAGAQRKNDEPSPNKNQTTEKKDQTREKQR 567
>CB065382 similar to PIR|T09671|T096 RPE15 protein - alfalfa (fragment),
partial (9%)
Length = 624
Score = 28.1 bits (61), Expect = 7.0
Identities = 17/49 (34%), Positives = 24/49 (48%), Gaps = 1/49 (2%)
Frame = -2
Query: 289 PPPPFSRARVMGQDKDAWCKYHL-VQGHNTDDCVHLKREIEKLLLSGKL 336
P PP R+ D C +H GH+ + C LK ++KL+ GKL
Sbjct: 383 PLPPRFRS-------DLKCDFHQGALGHDVEGCYALKHIVKKLINQGKL 258
>TC80104 similar to GP|20197699|gb|AAD20910.3 putative LRR receptor protein
kinase {Arabidopsis thaliana}, partial (28%)
Length = 1549
Score = 28.1 bits (61), Expect = 7.0
Identities = 30/115 (26%), Positives = 45/115 (39%), Gaps = 5/115 (4%)
Frame = +1
Query: 22 IQQVQQQEKTDSHHGDLDEPEPQPFSAKIWNAPVPDNFKPPHLPTFDGRGDPSEHVTVFN 81
+Q++ K H D+ PE +S P P PP +PT G +P+ ++
Sbjct: 886 VQRMATIAKPVDHEIDISTPEV--YSVPPPPPPPPPPPPPPSIPTKKGIVEPTTSHSLPT 1059
Query: 82 TRMSVYGVADSLKCKLLAGTFADAALRWYMSLP-----CFSIMGYQDMIKKFTQQ 131
R+ V GT D +LR S P CF+I Q F+Q+
Sbjct: 1060KRVIVEPTTSH------RGTTVDPSLR--SSSPPTFAKCFTIASLQQYTNSFSQE 1200
>TC79456 similar to PIR|T51518|T51518 ubiquitin-fusion degradation
protein-like - Arabidopsis thaliana, partial (31%)
Length = 1392
Score = 28.1 bits (61), Expect = 7.0
Identities = 21/60 (35%), Positives = 30/60 (50%), Gaps = 3/60 (5%)
Frame = -2
Query: 249 RGTFKKPYERNQNRYVPEHF--TPLNTRPERILKEVFESKIIPPPP-FSRARVMGQDKDA 305
R T K Y + QN Y + TP+++ ++IL + I PPPP F R D+DA
Sbjct: 737 RTTLSKFYPKFQNHYP*KTHPKTPVDSSLQQILVSPPNTPIPPPPPHFDIIRGSNGDRDA 558
>AW690258 GP|22900886|gb amt2-like protein {Medicago truncatula}, partial
(23%)
Length = 450
Score = 27.7 bits (60), Expect = 9.1
Identities = 22/60 (36%), Positives = 27/60 (44%), Gaps = 3/60 (5%)
Frame = +1
Query: 135 TRHRKVLSTSLFNVRQGPSESLREYLARFNDSTIKVSNPNQEVF---VGAFQNGLWAGQF 191
+RHR T LF+VRQ + L+ R N P +F VGAF LW G F
Sbjct: 19 SRHRHSHRTRLFSVRQNRTLILQSIPQRMNIKAWMAFVPLWLIFSYTVGAF--SLWGGGF 192
>TC87868 similar to PIR|T05112|T05112 splicing factor 9G8-like SR protein
RSZp22 [validated] - Arabidopsis thaliana, partial (91%)
Length = 860
Score = 27.7 bits (60), Expect = 9.1
Identities = 18/60 (30%), Positives = 25/60 (41%)
Frame = +3
Query: 211 ECYVKGEESNAEKKAREAKERGNSGGERRNHYVPPNRDRGTFKKPYERNQNRYVPEHFTP 270
+CY GE + ARE + RG G RR PP F++ + Y P +P
Sbjct: 333 KCYECGEPGHF---ARECRNRGGGGAGRRRSRSPPR-----FRRSPSYGRRSYSPRGRSP 488
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.317 0.134 0.407
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 14,274,127
Number of Sequences: 36976
Number of extensions: 213066
Number of successful extensions: 1191
Number of sequences better than 10.0: 36
Number of HSP's better than 10.0 without gapping: 1172
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1190
length of query: 437
length of database: 9,014,727
effective HSP length: 99
effective length of query: 338
effective length of database: 5,354,103
effective search space: 1809686814
effective search space used: 1809686814
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 60 (27.7 bits)
Medicago: description of AC135566.1


