

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC135565.5 + phase: 0 /pseudo
(93 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
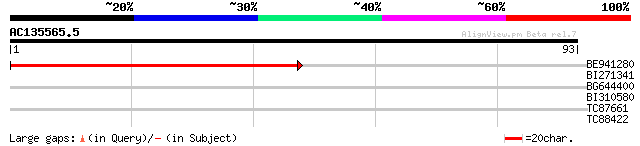
BE941280 weakly similar to GP|20197614|g unknown protein {Arabid... 57 9e-10
BI271341 37 0.001
BG644400 similar to GP|10177409|dbj contains similarity to unkno... 27 1.7
BI310580 25 3.9
TC87661 similar to GP|14517356|gb|AAK62569.1 AT4g15540/dl3810w {... 25 5.1
TC88422 similar to GP|21592839|gb|AAM64789.1 unknown {Arabidopsi... 24 8.6
>BE941280 weakly similar to GP|20197614|g unknown protein {Arabidopsis
thaliana}, partial (8%)
Length = 403
Score = 57.4 bits (137), Expect = 9e-10
Identities = 27/48 (56%), Positives = 38/48 (78%)
Frame = -1
Query: 1 MMNRFCFVALDKTLRNVMGSENKDNKTKPFGGKVFVLVGDFRQVLPVV 48
MM++ CF ALD++LR+V+ + ++ NK PFGGKV VL GDFRQ+L V+
Sbjct: 145 MMHKHCFEALDRSLRDVLKTVDERNKDIPFGGKVVVLGGDFRQILLVM 2
>BI271341
Length = 468
Score = 37.0 bits (84), Expect = 0.001
Identities = 17/52 (32%), Positives = 30/52 (57%)
Frame = +1
Query: 41 FRQVLPVVRVATRQQIVSSAVNASKVWRRCEVLRLTMNMGLSTANNNADQEE 92
FR +LPV+ +R I+ + +N+S + C+V+RL NM L +++ E
Sbjct: 1 FR*ILPVIPRGSRSDIIHATINSSCI*DHCQVVRLKKNMWLQQNGQSSNDPE 156
>BG644400 similar to GP|10177409|dbj contains similarity to unknown
protein~gene_id:K24M7.18~pir||T05785 {Arabidopsis
thaliana}, partial (4%)
Length = 677
Score = 26.6 bits (57), Expect = 1.7
Identities = 10/27 (37%), Positives = 19/27 (70%)
Frame = +1
Query: 55 QIVSSAVNASKVWRRCEVLRLTMNMGL 81
+I+S A+N K W +C++ +T++M L
Sbjct: 385 RILSVALNDRKTWLKCQMDTITVDMSL 465
>BI310580
Length = 297
Score = 25.4 bits (54), Expect = 3.9
Identities = 8/24 (33%), Positives = 17/24 (70%)
Frame = -1
Query: 6 CFVALDKTLRNVMGSENKDNKTKP 29
C++++ T + ++GS+ D+ TKP
Sbjct: 255 CYISVQVTTQQLVGSQKTDHITKP 184
>TC87661 similar to GP|14517356|gb|AAK62569.1 AT4g15540/dl3810w
{Arabidopsis thaliana}, partial (58%)
Length = 1258
Score = 25.0 bits (53), Expect = 5.1
Identities = 13/25 (52%), Positives = 14/25 (56%)
Frame = +3
Query: 22 NKDNKTKPFGGKVFVLVGDFRQVLP 46
NK+N T GG F L D QVLP
Sbjct: 9 NKENMTAESGGLNFDLPEDVMQVLP 83
>TC88422 similar to GP|21592839|gb|AAM64789.1 unknown {Arabidopsis
thaliana}, partial (67%)
Length = 782
Score = 24.3 bits (51), Expect = 8.6
Identities = 13/33 (39%), Positives = 18/33 (54%)
Frame = -2
Query: 3 NRFCFVALDKTLRNVMGSENKDNKTKPFGGKVF 35
N C K +R ++G + K N +PFG KVF
Sbjct: 205 NVLCHTTKLKIIRPLIGMK-KTNYVQPFGRKVF 110
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.322 0.133 0.380
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 2,380,772
Number of Sequences: 36976
Number of extensions: 19774
Number of successful extensions: 110
Number of sequences better than 10.0: 12
Number of HSP's better than 10.0 without gapping: 110
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 110
length of query: 93
length of database: 9,014,727
effective HSP length: 69
effective length of query: 24
effective length of database: 6,463,383
effective search space: 155121192
effective search space used: 155121192
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 51 (24.3 bits)
Medicago: description of AC135565.5


