

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
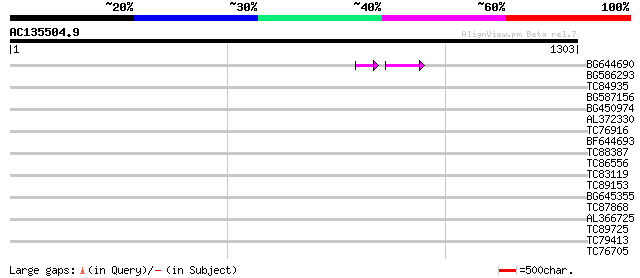
Query= AC135504.9 + phase: 0 /pseudo
(1303 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
BG644690 weakly similar to GP|18542179|gb putative pol protein {... 49 2e-07
BG586293 weakly similar to PIR|E84473|E84 probable retroelement ... 39 0.010
TC84935 similar to PIR|G96631|G96631 probable RNA-binding protei... 39 0.010
BG587156 similar to PIR|G85055|G8 probable polyprotein [imported... 37 0.050
BG450974 similar to PIR|T05112|T05 splicing factor 9G8-like SR p... 36 0.11
AL372330 similar to GP|21322752|dbj cold shock protein-1 {Tritic... 34 0.32
TC76916 MtN4 33 0.55
BF644693 homologue to GP|2598599|emb MtN4 {Medicago truncatula},... 33 0.55
TC88387 similar to GP|13357253|gb|AAK20050.1 putative zinc finge... 33 0.55
TC86556 similar to PIR|T07598|T07598 proline-rich protein GPP1 -... 33 0.94
TC83119 similar to GP|13357253|gb|AAK20050.1 putative zinc finge... 32 1.6
TC89153 similar to GP|18855061|gb|AAL79753.1 putative RNA helica... 32 1.6
BG645355 similar to PIR|G96590|G965 hypothetical protein T24C10.... 32 1.6
TC87868 similar to PIR|T05112|T05112 splicing factor 9G8-like SR... 32 2.1
AL366725 31 2.7
TC89725 similar to PIR|T05494|T05494 glycine-rich protein T19K4.... 31 3.6
TC79413 similar to GP|21439800|emb|CAD35173. unnamed protein pro... 31 3.6
TC76705 similar to SP|Q07202|CORA_MEDSA Cold and drought-regulat... 30 6.1
>BG644690 weakly similar to GP|18542179|gb putative pol protein {Zea mays},
partial (22%)
Length = 629
Score = 49.3 bits (116), Expect(2) = 2e-07
Identities = 35/91 (38%), Positives = 46/91 (50%)
Frame = -1
Query: 863 RIQSTRRH*LH*NICSSCKIGSNQVTSILRS*SWHNIISNGCQECFS*WCH*RRSVCKTT 922
RIQS RR+ L * + C+ GS + + NGC+EC *W R VC+ T
Sbjct: 392 RIQSKRRNRL**GFFTCCQNGSY*NFNSFCCIHGVQAVPNGCEECIY*WRSQRGGVCQAT 213
Query: 923 SWV*GS*AS*PCL*T*EITIWLETSS*SLV* 953
SW+* + C+ TIW E SS S+V*
Sbjct: 212 SWI*RCRGTKSCVQIE*DTIWSEASSKSMV* 120
Score = 25.0 bits (53), Expect(2) = 2e-07
Identities = 22/53 (41%), Positives = 28/53 (52%)
Frame = -2
Query: 794 *A*NC*RSSLR*WMDISYARRAKSVPKE*CVRSDTQT*SEEHY*NKMGIQKQA 846
*A C*RS +D YARR SV K+* + + T* + N +G KQA
Sbjct: 589 *AQEC*RSIA*CRLDQFYARRTPSV*KK*GMVPGSST*RQNSNWN*VGF*KQA 431
>BG586293 weakly similar to PIR|E84473|E84 probable retroelement pol
polyprotein [imported] - Arabidopsis thaliana, partial
(7%)
Length = 763
Score = 39.3 bits (90), Expect = 0.010
Identities = 23/70 (32%), Positives = 35/70 (49%), Gaps = 2/70 (2%)
Frame = +1
Query: 853 NQKQS*TCCTRIQSTRRH*LH*NICSSCKIGSNQVTSILRS*SWHNIISNGCQECFS*W- 911
+Q QS C R++ T RH L ++C+SC ++ T + S W S+ C+ C W
Sbjct: 145 DQIQSKASCKRLRETTRHRLRRSVCTSCSNRNHMTTLGVSSN*WMLDPSHRCKNCIPKWT 324
Query: 912 -CH*RRSVCK 920
C R S C+
Sbjct: 325 LCGNRNSHCR 354
>TC84935 similar to PIR|G96631|G96631 probable RNA-binding protein F8A5.17
[imported] - Arabidopsis thaliana, partial (41%)
Length = 552
Score = 39.3 bits (90), Expect = 0.010
Identities = 17/44 (38%), Positives = 23/44 (51%)
Frame = +2
Query: 181 ESKFGGRFRYGKVLGRGGYKNSRKEDQKGCFNCKKPGHFIADCP 224
+ ++ G F G GRG Y + Q CF C +PGH+ DCP
Sbjct: 395 DQRYRGGFSSG---GRGSYGAGDRVGQDDCFKCGRPGHWARDCP 517
>BG587156 similar to PIR|G85055|G8 probable polyprotein [imported] -
Arabidopsis thaliana, partial (17%)
Length = 618
Score = 37.0 bits (84), Expect = 0.050
Identities = 20/56 (35%), Positives = 31/56 (54%)
Frame = -3
Query: 854 QKQS*TCCTRIQSTRRH*LH*NICSSCKIGSNQVTSILRS*SWHNIISNGCQECFS 909
++++*T R+ S LH*+IC+S + N L W I++NGC+EC S
Sbjct: 382 EEEN*TSSKRVYSDIWRGLH*DICTSSQATHN*NCFKLGCEPWMGIVANGCEECIS 215
>BG450974 similar to PIR|T05112|T05 splicing factor 9G8-like SR protein
RSZp22 [validated] - Arabidopsis thaliana, partial (54%)
Length = 364
Score = 35.8 bits (81), Expect = 0.11
Identities = 13/29 (44%), Positives = 17/29 (57%)
Frame = +1
Query: 195 GRGGYKNSRKEDQKGCFNCKKPGHFIADC 223
GRGG + R D C+ C +PGHF +C
Sbjct: 274 GRGGGRGGRGGDDLKCYECGEPGHFAREC 360
>AL372330 similar to GP|21322752|dbj cold shock protein-1 {Triticum
aestivum}, partial (33%)
Length = 435
Score = 34.3 bits (77), Expect = 0.32
Identities = 19/62 (30%), Positives = 33/62 (52%), Gaps = 2/62 (3%)
Frame = +1
Query: 260 RQRVFKAHDWRESFVPYPYHERWR-RSEVWWQPNWKNHWYRYYW*FLHFNK-*CVACRWS 317
R+RV+ WR + RWR R +WW+ W R++W +L + + C+ C+++
Sbjct: 175 RRRVW----WRSIRRIWRRRLRWRQRRRIWWRRLWWRWLRRWWWRWLRWRQDGCIGCQFA 342
Query: 318 EA 319
EA
Sbjct: 343 EA 348
>TC76916 MtN4
Length = 1850
Score = 33.5 bits (75), Expect = 0.55
Identities = 15/42 (35%), Positives = 21/42 (49%), Gaps = 4/42 (9%)
Frame = -3
Query: 267 HDWRESFVPYPYHERWRRSEVWWQPNW---KNHW-YRYYW*F 304
H+WR ++ + WR +WW+ NW W YRY W F
Sbjct: 984 HNWRCWYIRWW*WYNWR*RNMWWENNWWLRNIRWCYRYNWWF 859
Score = 32.7 bits (73), Expect = 0.94
Identities = 12/34 (35%), Positives = 18/34 (52%), Gaps = 4/34 (11%)
Frame = -3
Query: 278 YHERWRRSEVWWQPNWKNH----WYRYYW*FLHF 307
Y WR +WW+ NW H WY +W*+ ++
Sbjct: 1038 YRHNWR*RNMWWKNNWWFHNWRCWYIRWW*WYNW 937
Score = 32.0 bits (71), Expect = 1.6
Identities = 14/29 (48%), Positives = 14/29 (48%), Gaps = 3/29 (10%)
Frame = -3
Query: 279 HERWRRSEVWWQPNWKNHWY---RYYW*F 304
H RW R WW NW WY YYW F
Sbjct: 840 HNRWFRDIWWWFHNW-GFWYVRWMYYWRF 757
>BF644693 homologue to GP|2598599|emb MtN4 {Medicago truncatula}, partial
(79%)
Length = 621
Score = 33.5 bits (75), Expect = 0.55
Identities = 15/42 (35%), Positives = 21/42 (49%), Gaps = 4/42 (9%)
Frame = -3
Query: 267 HDWRESFVPYPYHERWRRSEVWWQPNW---KNHW-YRYYW*F 304
H+WR ++ + WR +WW+ NW W YRY W F
Sbjct: 211 HNWRCWYIRWW*WYNWR*RNMWWENNWWLRNIRWCYRYNWWF 86
Score = 32.7 bits (73), Expect = 0.94
Identities = 12/34 (35%), Positives = 18/34 (52%), Gaps = 4/34 (11%)
Frame = -3
Query: 278 YHERWRRSEVWWQPNWKNH----WYRYYW*FLHF 307
Y WR +WW+ NW H WY +W*+ ++
Sbjct: 265 YRHNWR*RNMWWKNNWWFHNWRCWYIRWW*WYNW 164
>TC88387 similar to GP|13357253|gb|AAK20050.1 putative zinc finger protein
{Oryza sativa (japonica cultivar-group)}, partial (96%)
Length = 1286
Score = 33.5 bits (75), Expect = 0.55
Identities = 11/30 (36%), Positives = 21/30 (69%), Gaps = 6/30 (20%)
Frame = +1
Query: 203 RKEDQKG------CFNCKKPGHFIADCPDL 226
R++ ++G C NCK+PGH++ +CP++
Sbjct: 385 RRDSRRGFSQDNLCKNCKRPGHYVRECPNV 474
Score = 31.6 bits (70), Expect = 2.1
Identities = 10/20 (50%), Positives = 14/20 (70%)
Frame = +1
Query: 206 DQKGCFNCKKPGHFIADCPD 225
++K C NC+K GH DCP+
Sbjct: 718 NEKACNNCRKTGHLARDCPN 777
Score = 31.2 bits (69), Expect = 2.7
Identities = 9/16 (56%), Positives = 13/16 (81%)
Frame = +1
Query: 210 CFNCKKPGHFIADCPD 225
C+NCK+PGH + CP+
Sbjct: 538 CWNCKEPGHMASSCPN 585
Score = 25.4 bits (54), Expect(2) = 1.5
Identities = 7/16 (43%), Positives = 8/16 (49%)
Frame = +3
Query: 278 YHERWRRSEVWWQPNW 293
+ RW R WW P W
Sbjct: 846 WRPRWWRQPSWWLPRW 893
Score = 25.0 bits (53), Expect(2) = 1.5
Identities = 12/38 (31%), Positives = 19/38 (49%)
Frame = +1
Query: 206 DQKGCFNCKKPGHFIADCPDLQKEKSKSRPKKQSELVK 243
D + C NC K GH +C + EK+ + +K L +
Sbjct: 661 DLRLCNNCYKQGHIAVECTN---EKACNNCRKTGHLAR 765
>TC86556 similar to PIR|T07598|T07598 proline-rich protein GPP1 - potato,
partial (24%)
Length = 1250
Score = 32.7 bits (73), Expect = 0.94
Identities = 15/51 (29%), Positives = 23/51 (44%), Gaps = 10/51 (19%)
Frame = -1
Query: 269 WRESFVPYPYHERWR---------RSEVWWQPNWKNHW-YRYYW*FLHFNK 309
WR+ + ERW+ R WW W N W + YYW +++N+
Sbjct: 590 WRQ------WRERWKRWTWWFLEWRKNNWWFVYWNNGWLWNYYWRLIYWNR 456
>TC83119 similar to GP|13357253|gb|AAK20050.1 putative zinc finger protein
{Oryza sativa (japonica cultivar-group)}, partial (16%)
Length = 421
Score = 32.0 bits (71), Expect = 1.6
Identities = 13/39 (33%), Positives = 24/39 (61%), Gaps = 8/39 (20%)
Frame = +3
Query: 196 RGGYKNS--RKEDQKG------CFNCKKPGHFIADCPDL 226
R Y+++ R++ +G C NCK+PGH+ +CP++
Sbjct: 282 RHSYRDAPYRRDSSRGFSRDNLCKNCKRPGHYARECPNV 398
>TC89153 similar to GP|18855061|gb|AAL79753.1 putative RNA helicase {Oryza
sativa}, partial (3%)
Length = 737
Score = 32.0 bits (71), Expect = 1.6
Identities = 18/57 (31%), Positives = 26/57 (45%), Gaps = 1/57 (1%)
Frame = +3
Query: 170 LGKGKIEKKPLESKFGGRFRYG-KVLGRGGYKNSRKEDQKGCFNCKKPGHFIADCPD 225
LG+ ++ + GGR G R N R CF+C +PGH +DCP+
Sbjct: 24 LGREVLKPVMMTWLIGGRRSSGYSSSNRSSSPNRRGSYGGACFSCGQPGHRASDCPN 194
>BG645355 similar to PIR|G96590|G965 hypothetical protein T24C10.5 [imported]
- Arabidopsis thaliana, partial (5%)
Length = 627
Score = 32.0 bits (71), Expect = 1.6
Identities = 15/50 (30%), Positives = 24/50 (48%)
Frame = -1
Query: 177 KKPLESKFGGRFRYGKVLGRGGYKNSRKEDQKGCFNCKKPGHFIADCPDL 226
+ P E GG + V G GG C+ C++PGH+ ++CP +
Sbjct: 576 RNPPEQTAGGAY-VNTVSGSGGASGK-------CYKCQQPGHWASNCPSM 451
>TC87868 similar to PIR|T05112|T05112 splicing factor 9G8-like SR protein
RSZp22 [validated] - Arabidopsis thaliana, partial (91%)
Length = 860
Score = 31.6 bits (70), Expect = 2.1
Identities = 27/104 (25%), Positives = 43/104 (40%), Gaps = 7/104 (6%)
Frame = +3
Query: 185 GGRFRYGKVLGRGGYKNSRKEDQKGCFNCKKPGHFIADCPD-------LQKEKSKSRPKK 237
GGR R G G GG + + C+ C +PGHF +C + ++ +S R ++
Sbjct: 288 GGRGRGG---GGGGGSDLK------CYECGEPGHFARECRNRGGGGAGRRRSRSPPRFRR 440
Query: 238 QSELVKTAGVPKGKRETKVMVPRQRVFKAHDWRESFVPYPYHER 281
+ + P+G+ PR+R S P PY R
Sbjct: 441 SPSYGRRSYSPRGRS------PRRRSVSPRGRSYSRSPPPYRGR 554
>AL366725
Length = 485
Score = 31.2 bits (69), Expect = 2.7
Identities = 14/36 (38%), Positives = 19/36 (51%)
Frame = +2
Query: 188 FRYGKVLGRGGYKNSRKEDQKGCFNCKKPGHFIADC 223
F YG+ +G N ++ K C C K GH +ADC
Sbjct: 311 FNYGE---KGHKSNVCPKEIKKCVRCDKKGHIVADC 409
>TC89725 similar to PIR|T05494|T05494 glycine-rich protein T19K4.150 -
Arabidopsis thaliana, partial (17%)
Length = 378
Score = 30.8 bits (68), Expect = 3.6
Identities = 16/40 (40%), Positives = 19/40 (47%)
Frame = +1
Query: 185 GGRFRYGKVLGRGGYKNSRKEDQKGCFNCKKPGHFIADCP 224
GG RYG G GG C++C + GHF DCP
Sbjct: 73 GGGGRYGGGGGGGGGGGGGGS----CYSCGESGHFARDCP 180
>TC79413 similar to GP|21439800|emb|CAD35173. unnamed protein product
{Glycine max}, partial (42%)
Length = 971
Score = 30.8 bits (68), Expect = 3.6
Identities = 14/40 (35%), Positives = 20/40 (50%)
Frame = -2
Query: 277 PYHERWRRSEVWWQPNWKNHWYRYYW*FLHFNK*CVACRW 316
P + W R WW+ W N W+ Y+W* + + C RW
Sbjct: 580 P*FDYWSR---WWR--WSNWWFNYWW*LI---RRCFHNRW 485
>TC76705 similar to SP|Q07202|CORA_MEDSA Cold and drought-regulated protein
CORA. [Alfalfa] {Medicago sativa}, partial (80%)
Length = 854
Score = 30.0 bits (66), Expect = 6.1
Identities = 10/26 (38%), Positives = 14/26 (53%), Gaps = 3/26 (11%)
Frame = +1
Query: 280 ERWRRSEVWWQP---NWKNHWYRYYW 302
+RWR + WW+P W W R +W
Sbjct: 313 QRWRWLQWWWRPRRSRWLQSWRRSWW 390
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.382 0.170 0.717
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 42,271,596
Number of Sequences: 36976
Number of extensions: 649317
Number of successful extensions: 10380
Number of sequences better than 10.0: 36
Number of HSP's better than 10.0 without gapping: 1871
Number of HSP's successfully gapped in prelim test: 365
Number of HSP's that attempted gapping in prelim test: 8084
Number of HSP's gapped (non-prelim): 2852
length of query: 1303
length of database: 9,014,727
effective HSP length: 108
effective length of query: 1195
effective length of database: 5,021,319
effective search space: 6000476205
effective search space used: 6000476205
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 13 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 35 (21.8 bits)
S2: 64 (29.3 bits)
Medicago: description of AC135504.9


