

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC135504.16 - phase: 0 /pseudo
(343 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
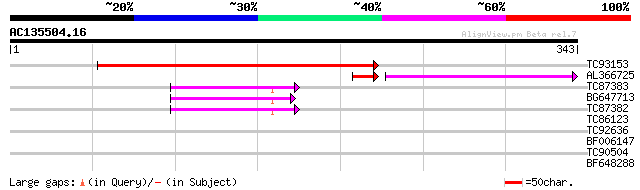
Score E
Sequences producing significant alignments: (bits) Value
TC93153 similar to GP|14715220|emb|CAC44106. gag polyprotein {Ci... 281 3e-76
AL366725 63 4e-14
TC87383 similar to GP|19168656|emb|CAD26175. DNA-DIRECTED RNA PO... 50 2e-06
BG647713 homologue to GP|15042313|gb| 232R {Chilo iridescent vir... 49 4e-06
TC87382 similar to EGAD|146423|156195 vitellogenin {Anolis pulch... 47 8e-06
TC86123 homologue to PIR|S71770|S71770 calcium-dependent protein... 28 3.9
TC92636 homologue to GP|15042313|gb|AAK82093.1 232R {Chilo iride... 28 5.1
BF006147 similar to GP|12322790|gb| unknown protein; 108050-1057... 28 5.1
TC90504 similar to PIR|T48329|T48329 hypothetical protein F15A17... 28 6.7
BF648288 homologue to GP|19918388|gb| ribosomal protein L1p {Met... 27 8.8
>TC93153 similar to GP|14715220|emb|CAC44106. gag polyprotein {Cicer
arietinum}, partial (8%)
Length = 516
Score = 281 bits (719), Expect = 3e-76
Identities = 135/170 (79%), Positives = 146/170 (85%)
Frame = +2
Query: 54 MAGRNDPAIAAALEAVA*AVGQQPNAAAGNDGVRMLETFLRNHPPTFKGRYDPDGTQKWL 113
M G +D A+ AALEAVA AV Q P G+DG RMLETFLRNHPPTFKGRY PDG KWL
Sbjct: 2 MTGSSDAALVAALEAVAQAVQQLPKVDTGSDGTRMLETFLRNHPPTFKGRYAPDGA*KWL 181
Query: 114 KEVERIFRVMQCSEVQKVRFGTHMLAEEADDWWVSLLPVLEQDGAVVTWAVFRREFLNRY 173
KE+ERIFRVMQC E QKV+FGTHMLAEEADDWW+SLLPVLEQD AVVTWA+FR+EFL RY
Sbjct: 182 KEIERIFRVMQCFETQKVQFGTHMLAEEADDWWISLLPVLEQDDAVVTWAMFRKEFLGRY 361
Query: 174 FPEDVRGKNEIEFLELKHGDMSVTKYAAKFVELAKFYPYYTAEIVEFSKC 223
FPEDVRGK EIEFLELK GDMSVT+YAAKFVELA FYP+Y+AE EFSKC
Sbjct: 362 FPEDVRGKKEIEFLELKQGDMSVTEYAAKFVELATFYPHYSAETAEFSKC 511
>AL366725
Length = 485
Score = 63.2 bits (152), Expect(2) = 4e-14
Identities = 50/116 (43%), Positives = 63/116 (54%)
Frame = +1
Query: 228 EWLEG*HQESHWLSENKKLL*FDK*LQDL*GGHESSLQGDE*TQRQGTTESSEAV*CSC* 287
EW E HQE + + *F + LQ+L GG+ S QG E* + QG ES A+*C C*
Sbjct: 61 EWFEARHQEGNRIPTAPSFP*FSEYLQNLRGGY*GS*QGCE*AEDQGPIESP*AL*CPC* 240
Query: 288 QG*AEVK**EEAQEERCSYEYCLLQV**EGP*E*CL*QR*EEVLQVWSERTHSS*L 343
G E * + EE CS CL Q+ E P E*CL R +E+ V E ++ S*L
Sbjct: 241 *GQTENGR**AS*EEGCSCGDCLFQLWRERPQE*CLS*RDQEMCPV*QEGSYCS*L 408
Score = 32.0 bits (71), Expect(2) = 4e-14
Identities = 12/16 (75%), Positives = 13/16 (81%)
Frame = +2
Query: 208 KFYPYYTAEIVEFSKC 223
KFYP+Y AE EFSKC
Sbjct: 2 KFYPHYAAETAEFSKC 49
>TC87383 similar to GP|19168656|emb|CAD26175. DNA-DIRECTED RNA POLYMERASE II
{Encephalitozoon cuniculi}, partial (0%)
Length = 1247
Score = 49.7 bits (117), Expect = 2e-06
Identities = 24/80 (30%), Positives = 41/80 (51%), Gaps = 2/80 (2%)
Frame = -2
Query: 98 PTFKGRYDPDGTQKWLKEVERIFRVMQCSEVQKVRFGTHMLAEEADDWWVSLLPVLEQDG 157
P F+G PD WL+ +ER+F+ + E QKV+ L + A WW ++ +++G
Sbjct: 724 PDFEGNLQPDDLLDWLQIMERLFKYKEVLEEQKVKIVAAKLKKLASIWWENVKRRRKREG 545
Query: 158 --AVVTWAVFRREFLNRYFP 175
+ TW R++ +Y P
Sbjct: 544 KSKIKTWEKMRQKLTRKYLP 485
>BG647713 homologue to GP|15042313|gb| 232R {Chilo iridescent virus}, partial
(1%)
Length = 726
Score = 48.5 bits (114), Expect = 4e-06
Identities = 24/78 (30%), Positives = 39/78 (49%), Gaps = 2/78 (2%)
Frame = +2
Query: 98 PTFKGRYDPDGTQKWLKEVERIFRVMQCSEVQKVRFGTHMLAEEADDWWVSLLPVLEQDG 157
P F G+ PD WL+ +ER+F+ + +E QKV+ L + A WW +L +G
Sbjct: 347 PDF*GKLQPDEFVDWLQTIERVFKYKEVAEEQKVKIVAAKLKKHASIWWKNLKRKRNCEG 526
Query: 158 --AVVTWAVFRREFLNRY 173
+ TW R++ +Y
Sbjct: 527 KSKIKTWDKMRQKLTRKY 580
>TC87382 similar to EGAD|146423|156195 vitellogenin {Anolis pulchellus},
partial (7%)
Length = 2304
Score = 47.4 bits (111), Expect = 8e-06
Identities = 24/80 (30%), Positives = 39/80 (48%), Gaps = 2/80 (2%)
Frame = +2
Query: 98 PTFKGRYDPDGTQKWLKEVERIFRVMQCSEVQKVRFGTHMLAEEADDWWVSLLPVLEQDG 157
P F+G D WL+ +ER+F + E QKV+ L + A WW +L +++G
Sbjct: 629 PDFEGNLQLDDFLDWLQTIERVFEYKEVPEEQKVKIVAAKLKKHALIWWENLKRRRKREG 808
Query: 158 --AVVTWAVFRREFLNRYFP 175
+ TW R++ +Y P
Sbjct: 809 KSKIKTWDKMRQKLTRKYLP 868
>TC86123 homologue to PIR|S71770|S71770 calcium-dependent protein kinase (EC
2.7.1.-) - mung bean, complete
Length = 2448
Score = 28.5 bits (62), Expect = 3.9
Identities = 10/20 (50%), Positives = 12/20 (60%)
Frame = -1
Query: 37 CCLCCCSELLFLM*NFEMAG 56
CC CCC LF + N + AG
Sbjct: 363 CCCCCCQR*LFALKNGQQAG 304
>TC92636 homologue to GP|15042313|gb|AAK82093.1 232R {Chilo iridescent
virus}, partial (1%)
Length = 772
Score = 28.1 bits (61), Expect = 5.1
Identities = 10/23 (43%), Positives = 14/23 (60%)
Frame = +3
Query: 98 PTFKGRYDPDGTQKWLKEVERIF 120
P F+G PD WL+ +ER+F
Sbjct: 438 PDFEGELQPDEFVDWLQAIERVF 506
>BF006147 similar to GP|12322790|gb| unknown protein; 108050-105786
{Arabidopsis thaliana}, partial (11%)
Length = 551
Score = 28.1 bits (61), Expect = 5.1
Identities = 12/37 (32%), Positives = 18/37 (48%)
Frame = -2
Query: 83 NDGVRMLETFLRNHPPTFKGRYDPDGTQKWLKEVERI 119
NDG+++LE FL P YD Q W+ + +
Sbjct: 199 NDGLQILEVFLPKEHPQLHLTYDEHDIQFWISSQQHM 89
>TC90504 similar to PIR|T48329|T48329 hypothetical protein F15A17.110 -
Arabidopsis thaliana, partial (54%)
Length = 611
Score = 27.7 bits (60), Expect = 6.7
Identities = 15/45 (33%), Positives = 21/45 (46%), Gaps = 5/45 (11%)
Frame = +3
Query: 135 THMLAEEADDW-----WVSLLPVLEQDGAVVTWAVFRREFLNRYF 174
TH+ E D W+SL+PV G + +FRRE +F
Sbjct: 123 THVRYERGDPLGHFLAWISLIPVFISFGGFFSHFIFRREIQTIFF 257
>BF648288 homologue to GP|19918388|gb| ribosomal protein L1p
{Methanosarcina acetivorans str. C2A} [Methanosarcina
acetivorans C2A], partial (5%)
Length = 181
Score = 27.3 bits (59), Expect = 8.8
Identities = 9/15 (60%), Positives = 12/15 (80%)
Frame = +2
Query: 31 IECQL*CCLCCCSEL 45
IE ++*CC CCC +L
Sbjct: 77 IETEM*CC*CCCCDL 121
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.355 0.158 0.577
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 10,856,952
Number of Sequences: 36976
Number of extensions: 143081
Number of successful extensions: 2864
Number of sequences better than 10.0: 20
Number of HSP's better than 10.0 without gapping: 1817
Number of HSP's successfully gapped in prelim test: 46
Number of HSP's that attempted gapping in prelim test: 531
Number of HSP's gapped (non-prelim): 2242
length of query: 343
length of database: 9,014,727
effective HSP length: 97
effective length of query: 246
effective length of database: 5,428,055
effective search space: 1335301530
effective search space used: 1335301530
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 14 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 37 (21.6 bits)
S2: 59 (27.3 bits)
Medicago: description of AC135504.16


