

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC135465.2 + phase: 0
(194 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
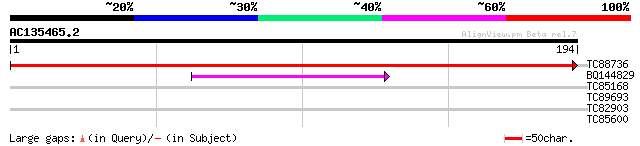
Score E
Sequences producing significant alignments: (bits) Value
TC88736 homologue to GP|13277515|gb|AAK16221.1 PII protein {Medi... 380 e-106
BQ144829 60 5e-10
TC85168 similar to PIR|T06827|T06827 lipoxygenase (EC 1.13.11.12... 28 2.9
TC89693 weakly similar to GP|21104633|dbj|BAB93225. hypothetical... 27 5.0
TC82903 similar to GP|21593294|gb|AAM65243.1 putative carboxyl-t... 27 5.0
TC85600 homologue to SP|P29828|PDI_MEDSA Protein disulfide isome... 26 8.5
>TC88736 homologue to GP|13277515|gb|AAK16221.1 PII protein {Medicago
sativa}, complete
Length = 733
Score = 380 bits (976), Expect = e-106
Identities = 194/194 (100%), Positives = 194/194 (100%)
Frame = +3
Query: 1 MALIAKPNVFNGLNFHINETQFPFSSFSVIRKRFGDSSHRNVVLKSNGNASILPKIRAQN 60
MALIAKPNVFNGLNFHINETQFPFSSFSVIRKRFGDSSHRNVVLKSNGNASILPKIRAQN
Sbjct: 36 MALIAKPNVFNGLNFHINETQFPFSSFSVIRKRFGDSSHRNVVLKSNGNASILPKIRAQN 215
Query: 61 LPDYVPESKFYKVEAILRPWRIPQVSSGLLKMGIRGVTVSDVKGFGAQGGSKERQGGSEF 120
LPDYVPESKFYKVEAILRPWRIPQVSSGLLKMGIRGVTVSDVKGFGAQGGSKERQGGSEF
Sbjct: 216 LPDYVPESKFYKVEAILRPWRIPQVSSGLLKMGIRGVTVSDVKGFGAQGGSKERQGGSEF 395
Query: 121 SEDNFVAKVKMEIVVRKDQVEAVINKIMETARTGEIGDGKIFLIPVSDVIRIRTGERGEQ 180
SEDNFVAKVKMEIVVRKDQVEAVINKIMETARTGEIGDGKIFLIPVSDVIRIRTGERGEQ
Sbjct: 396 SEDNFVAKVKMEIVVRKDQVEAVINKIMETARTGEIGDGKIFLIPVSDVIRIRTGERGEQ 575
Query: 181 AERMAGGLTDALSV 194
AERMAGGLTDALSV
Sbjct: 576 AERMAGGLTDALSV 617
>BQ144829
Length = 320
Score = 60.1 bits (144), Expect = 5e-10
Identities = 29/68 (42%), Positives = 40/68 (58%)
Frame = +2
Query: 63 DYVPESKFYKVEAILRPWRIPQVSSGLLKMGIRGVTVSDVKGFGAQGGSKERQGGSEFSE 122
D P +F++VEAI+RPWR+ V + L + GI VT + V+G GA GG+K R G
Sbjct: 68 DAXPNCEFFRVEAIIRPWRLEPVMAALAEKGIVAVTSTRVRGAGAXGGTKXRYSGKAHXT 247
Query: 123 DNFVAKVK 130
+ V K K
Sbjct: 248 KDLVEKAK 271
>TC85168 similar to PIR|T06827|T06827 lipoxygenase (EC 1.13.11.12) - garden
pea, partial (98%)
Length = 2890
Score = 27.7 bits (60), Expect = 2.9
Identities = 9/22 (40%), Positives = 15/22 (67%)
Frame = -2
Query: 69 KFYKVEAILRPWRIPQVSSGLL 90
+F+K+E++ PW P +S LL
Sbjct: 2565 RFFKLESLFLPWIFPSISDSLL 2500
>TC89693 weakly similar to GP|21104633|dbj|BAB93225. hypothetical
protein~similar to Arabidopsis thaliana chromosome 3
At3g46550, partial (53%)
Length = 1346
Score = 26.9 bits (58), Expect = 5.0
Identities = 25/72 (34%), Positives = 35/72 (47%), Gaps = 5/72 (6%)
Frame = -3
Query: 93 GIRGVTVSDVKGFGAQGGSKERQGGSEFSEDNFVAKVKM-----EIVVRKDQVEAVINKI 147
G+ + ++V G GA GG +E G EF+ V +V E VV +D E ++
Sbjct: 438 GVGDINGAEVVG-GAAGGLEE--SGDEFTGGREVLEVGPGEELEEDVVAEDVGEGGGGEV 268
Query: 148 METARTGEIGDG 159
ME GEIG G
Sbjct: 267 MEVRVGGEIGVG 232
>TC82903 similar to GP|21593294|gb|AAM65243.1 putative carboxyl-terminal
peptidase {Arabidopsis thaliana}, partial (88%)
Length = 1155
Score = 26.9 bits (58), Expect = 5.0
Identities = 12/49 (24%), Positives = 22/49 (44%)
Frame = +2
Query: 32 KRFGDSSHRNVVLKSNGNASILPKIRAQNLPDYVPESKFYKVEAILRPW 80
KR+G H++ + ++ + Q+ YV KFY +A + W
Sbjct: 347 KRYGRKKHKSFAKPRSAEPDLVNQSGHQHAIAYVEGDKFYGAKATINVW 493
>TC85600 homologue to SP|P29828|PDI_MEDSA Protein disulfide isomerase
precursor (PDI) (EC 5.3.4.1). [Alfalfa] {Medicago
sativa}, complete
Length = 1915
Score = 26.2 bits (56), Expect = 8.5
Identities = 14/61 (22%), Positives = 31/61 (49%)
Frame = +2
Query: 42 VVLKSNGNASILPKIRAQNLPDYVPESKFYKVEAILRPWRIPQVSSGLLKMGIRGVTVSD 101
++ ++G P + LP ++ K KVE ++ IP+ ++ +K+ + G T+ D
Sbjct: 1040 IIQHNDGKKFFKPNLELDQLPTWLKAYKDGKVEPFVKSEPIPETNNEPVKV-VVGQTLED 1216
Query: 102 V 102
+
Sbjct: 1217 I 1219
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.318 0.137 0.382
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 4,398,901
Number of Sequences: 36976
Number of extensions: 48474
Number of successful extensions: 258
Number of sequences better than 10.0: 12
Number of HSP's better than 10.0 without gapping: 257
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 258
length of query: 194
length of database: 9,014,727
effective HSP length: 91
effective length of query: 103
effective length of database: 5,649,911
effective search space: 581940833
effective search space used: 581940833
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 56 (26.2 bits)
Medicago: description of AC135465.2


