

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC135465.11 - phase: 0
(295 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
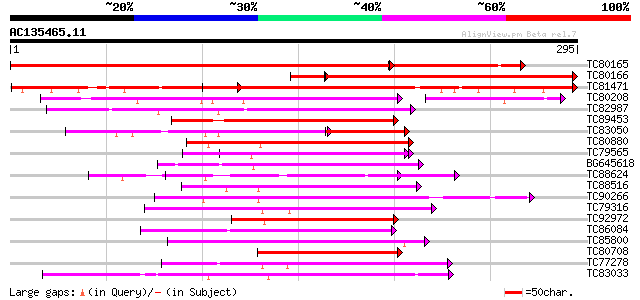
Score E
Sequences producing significant alignments: (bits) Value
TC80165 similar to PIR|A84913|A84913 probable WRKY-type DNA bind... 405 e-141
TC80166 similar to PIR|A84913|A84913 probable WRKY-type DNA bind... 273 8e-74
TC81471 similar to PIR|A84913|A84913 probable WRKY-type DNA bind... 232 1e-61
TC80208 similar to GP|17064158|gb|AAL35286.1 WRKY transcription ... 145 1e-36
TC82987 similar to GP|15991740|gb|AAL13047.1 WRKY transcription ... 149 1e-36
TC89453 similar to GP|9187622|emb|CAB97004.1 WRKY DNA binding pr... 124 3e-29
TC83050 similar to SP|Q9FGZ4|WR48_ARATH Probable WRKY transcript... 124 4e-29
TC80880 similar to SP|Q9ZQ70|WRK3_ARATH Probable WRKY transcript... 119 1e-27
TC79565 similar to SP|Q9ZQ70|WRK3_ARATH Probable WRKY transcript... 116 9e-27
BG645618 similar to GP|18158619|gb WRKY-like drought-induced pro... 116 1e-26
TC88624 similar to PIR|S51529|S51529 SPF1 protein - sweet potato... 112 1e-25
TC88516 similar to GP|14530681|dbj|BAB61053. WRKY DNA-binding pr... 107 4e-24
TC90266 similar to SP|Q9C519|WRK6_ARATH WRKY transcription facto... 100 1e-21
TC79316 similar to GP|11493822|gb|AAG35658.1 transcription facto... 98 4e-21
TC92972 similar to GP|9187622|emb|CAB97004.1 WRKY DNA binding pr... 97 7e-21
TC86084 similar to GP|21537078|gb|AAM61419.1 putaive DNA-binding... 96 1e-20
TC85800 similar to GP|11320830|dbj|BAB18313. putative WRKY DNA b... 96 2e-20
TC80708 weakly similar to GP|14530685|dbj|BAB61055. WRKY DNA-bin... 96 2e-20
TC77278 similar to GP|11493822|gb|AAG35658.1 transcription facto... 93 1e-19
TC83033 similar to PIR|S51529|S51529 SPF1 protein - sweet potato... 90 9e-19
>TC80165 similar to PIR|A84913|A84913 probable WRKY-type DNA binding protein
[imported] - Arabidopsis thaliana, partial (32%)
Length = 1013
Score = 405 bits (1041), Expect(2) = e-141
Identities = 200/200 (100%), Positives = 200/200 (100%)
Frame = +1
Query: 1 MMEKKGLNMEDYANIGSFPSYSFPSVFDLSEERSFMELLGVQQNMNNYSDSLLDLPVVVK 60
MMEKKGLNMEDYANIGSFPSYSFPSVFDLSEERSFMELLGVQQNMNNYSDSLLDLPVVVK
Sbjct: 202 MMEKKGLNMEDYANIGSFPSYSFPSVFDLSEERSFMELLGVQQNMNNYSDSLLDLPVVVK 381
Query: 61 EPPLESDGNGKEYSEVLNSQQQPATPNSSSISSASSEAINDEHNKTVDQTNNQLNKQLKA 120
EPPLESDGNGKEYSEVLNSQQQPATPNSSSISSASSEAINDEHNKTVDQTNNQLNKQLKA
Sbjct: 382 EPPLESDGNGKEYSEVLNSQQQPATPNSSSISSASSEAINDEHNKTVDQTNNQLNKQLKA 561
Query: 121 KKTNQKKPREARIAFMTKSEVDHLEDGYRWRKYGQKAVKNSPFPRSYYRCTSVSCNVKKH 180
KKTNQKKPREARIAFMTKSEVDHLEDGYRWRKYGQKAVKNSPFPRSYYRCTSVSCNVKKH
Sbjct: 562 KKTNQKKPREARIAFMTKSEVDHLEDGYRWRKYGQKAVKNSPFPRSYYRCTSVSCNVKKH 741
Query: 181 VERSLSDPTIVVTTYEGKHT 200
VERSLSDPTIVVTTYEGKHT
Sbjct: 742 VERSLSDPTIVVTTYEGKHT 801
Score = 114 bits (285), Expect(2) = e-141
Identities = 57/71 (80%), Positives = 62/71 (87%)
Frame = +2
Query: 198 KHTHPNPIMSRSSAVRAGSLLPPPAECTTNFASDQNYDISQYYNQQRQQVLFNTLSSLGF 257
++THPNPIMSRSSAVRAGSLLPPPAECTTNFASDQNYDISQYYNQQR + + S GF
Sbjct: 791 ENTHPNPIMSRSSAVRAGSLLPPPAECTTNFASDQNYDISQYYNQQRHKYSSYIVVS-GF 967
Query: 258 PSKNMNATFSQ 268
P K+MNATFSQ
Sbjct: 968 PFKDMNATFSQ 1000
>TC80166 similar to PIR|A84913|A84913 probable WRKY-type DNA binding protein
[imported] - Arabidopsis thaliana, partial (24%)
Length = 829
Score = 273 bits (697), Expect = 8e-74
Identities = 131/131 (100%), Positives = 131/131 (100%)
Frame = +1
Query: 165 RSYYRCTSVSCNVKKHVERSLSDPTIVVTTYEGKHTHPNPIMSRSSAVRAGSLLPPPAEC 224
RSYYRCTSVSCNVKKHVERSLSDPTIVVTTYEGKHTHPNPIMSRSSAVRAGSLLPPPAEC
Sbjct: 235 RSYYRCTSVSCNVKKHVERSLSDPTIVVTTYEGKHTHPNPIMSRSSAVRAGSLLPPPAEC 414
Query: 225 TTNFASDQNYDISQYYNQQRQQVLFNTLSSLGFPSKNMNATFSQDRPLCNPRVQDNGLLQ 284
TTNFASDQNYDISQYYNQQRQQVLFNTLSSLGFPSKNMNATFSQDRPLCNPRVQDNGLLQ
Sbjct: 415 TTNFASDQNYDISQYYNQQRQQVLFNTLSSLGFPSKNMNATFSQDRPLCNPRVQDNGLLQ 594
Query: 285 DVVPSHMFKEE 295
DVVPSHMFKEE
Sbjct: 595 DVVPSHMFKEE 627
Score = 45.1 bits (105), Expect = 3e-05
Identities = 19/20 (95%), Positives = 19/20 (95%)
Frame = +1
Query: 147 GYRWRKYGQKAVKNSPFPRS 166
G RWRKYGQKAVKNSPFPRS
Sbjct: 10 GTRWRKYGQKAVKNSPFPRS 69
>TC81471 similar to PIR|A84913|A84913 probable WRKY-type DNA binding protein
[imported] - Arabidopsis thaliana, partial (43%)
Length = 1274
Score = 232 bits (591), Expect = 1e-61
Identities = 132/215 (61%), Positives = 154/215 (71%), Gaps = 20/215 (9%)
Frame = +2
Query: 101 DEHNKTVDQTNNQLNKQLKAKKTNQKKPREARIAFMTKSEVDHLEDGYRWRKYGQKAVKN 160
D+ + ++ + NKQLK KKTN K+ RE R AFMTKSEVDHLEDGYRWRKYGQKAVKN
Sbjct: 425 DQAEEDEEEEKQKTNKQLKTKKTNLKRQREPRFAFMTKSEVDHLEDGYRWRKYGQKAVKN 604
Query: 161 SPFPRSYYRCTSVSCNVKKHVERSLSDPTIVVTTYEGKHTHPNPIMSRSSAVRAGSLLPP 220
SPFPRSYYRCT+ SCNVKK VERS +DP+IVVTTYEG+HTHP+P MSRS+ AG +P
Sbjct: 605 SPFPRSYYRCTTASCNVKKRVERSYTDPSIVVTTYEGQHTHPSPTMSRSAF--AGVQIPQ 778
Query: 221 PAE------CTTNFAS--DQNYDISQYYNQ--QRQQVLFNTLSSLGFPSKN----MNATF 266
PA TTNF S NY +SQY+ Q Q QQ+L NTLSSL P N N+ F
Sbjct: 779 PAGVVSGGFSTTNFGSVLQGNY-LSQYHQQPYQHQQLLVNTLSSLSHPYNNSSSFKNSPF 955
Query: 267 -SQDR-PLCNPR----VQDNGLLQDVVPSHMFKEE 295
+Q+R LCNP ++DNGLLQDVVPSHM KE+
Sbjct: 956 TTQERVQLCNPGTTAFLRDNGLLQDVVPSHMLKEK 1060
Score = 75.5 bits (184), Expect = 2e-14
Identities = 66/134 (49%), Positives = 81/134 (60%), Gaps = 14/134 (10%)
Frame = +1
Query: 2 MEKK--GLNMEDYANIGSFPS-----YSFPSVFDLSE-ERS----FMELLGVQQNMNNYS 49
MEKK GL+++ + S S Y F SVFD E ERS FMELLG ++YS
Sbjct: 85 MEKKDMGLSVKSEDVVDSSSSLLNNGYQFSSVFDFCEVERSSSLGFMELLGG----HDYS 252
Query: 50 DSLLDLPVV--VKEPPLESDGNGKEYSEVLNSQQQPATPNSSSISSASSEAINDEHNKTV 107
LLD+P + + SD GKE SEVLN QQQPATPNS SISSASSEA+ND NKT+
Sbjct: 253 P-LLDVPQLSTMSTVKASSDTTGKECSEVLN-QQQPATPNSCSISSASSEAVND--NKTL 420
Query: 108 DQTNNQLNKQLKAK 121
+ + ++ KAK
Sbjct: 421 G*PSRRR*RRRKAK 462
>TC80208 similar to GP|17064158|gb|AAL35286.1 WRKY transcription factor 28
{Arabidopsis thaliana}, partial (31%)
Length = 1294
Score = 145 bits (365), Expect(2) = 1e-36
Identities = 92/207 (44%), Positives = 117/207 (56%), Gaps = 19/207 (9%)
Frame = +1
Query: 17 SFPSYSFPSVFDLSEERSFMELLGVQQNMNNYSDSLLDLPVVVKEPPLE-----SDGNGK 71
S P+Y PS +++S S NM++Y+ L V+ E GN K
Sbjct: 181 SQPNYFDPSSYNMSFTESLQG-----GNMDHYNSPLATSFGVLSPSSSEVFSSVDQGNQK 345
Query: 72 EYS-EVLNSQQQPATPNSSSISSASSEA--INDEHN----KTVDQTNNQLNKQLKA---- 120
+ E+ Q AT NSS +S+SSEA + +E + K + Q + ++
Sbjct: 346 PSAGELGGGDQTLATLNSSISNSSSSEAGGVEEEDSGKRKKNIGQVKEEPGQESSKNGNN 525
Query: 121 ---KKTNQKKPREARIAFMTKSEVDHLEDGYRWRKYGQKAVKNSPFPRSYYRCTSVSCNV 177
KK +KK +E R AFMTKSEVD LEDGYRWRKYGQKAVKNSP+PRSYYRCT+ C V
Sbjct: 526 KVDKKKGEKKQKEPRFAFMTKSEVDQLEDGYRWRKYGQKAVKNSPYPRSYYRCTTQKCTV 705
Query: 178 KKHVERSLSDPTIVVTTYEGKHTHPNP 204
KK VERS DPT V+TTYEG+H P
Sbjct: 706 KKRVERSYEDPTTVITTYEGQHNASCP 786
Score = 25.8 bits (55), Expect(2) = 1e-36
Identities = 22/76 (28%), Positives = 33/76 (42%), Gaps = 3/76 (3%)
Frame = +3
Query: 217 LLPPPAECTTNFASDQNYDISQYYNQQRQQVLFNTLSSLG---FPSKNMNATFSQDRPLC 273
L P P +NF D + + Q +T +++ + N+N +F Q
Sbjct: 840 LTPTPFSYGSNFPQDLLLHMHHHNTFNITQPTTHTTNNVAGSVYSHNNINNSFLQQYQQQ 1019
Query: 274 NPRVQDNGLLQDVVPS 289
P Q GLLQD+VPS
Sbjct: 1020LPPDQ-YGLLQDIVPS 1064
>TC82987 similar to GP|15991740|gb|AAL13047.1 WRKY transcription factor 71
{Arabidopsis thaliana}, partial (33%)
Length = 975
Score = 149 bits (376), Expect = 1e-36
Identities = 92/204 (45%), Positives = 116/204 (56%), Gaps = 12/204 (5%)
Frame = +1
Query: 20 SYSFPSVFDLSEERSFMELLGVQQNMNNYSDSLLDLPVVVKEPPLESDGNGKEYSEV--L 77
S +F S SF + L + N S + DL E D N + S L
Sbjct: 274 SQNFQGFDPSSHSTSFSDYLHGSMDYNTLSKAF-DLSCSSSEVISSIDDNNTKKSSAGDL 450
Query: 78 NSQQQPATPNSSSISSASSEAINDEHNKTV----------DQTNNQLNKQLKAKKTNQKK 127
+S + SS SS+S+EA+ +E + D + + K+ KAKK +KK
Sbjct: 451 SSLVGISETPKSSESSSSNEAVIEEDSTNKKDKQPKLGCEDGDDEKSKKENKAKK-KEKK 627
Query: 128 PREARIAFMTKSEVDHLEDGYRWRKYGQKAVKNSPFPRSYYRCTSVSCNVKKHVERSLSD 187
+E R AF+TKSE+DHLEDGYRWRKYGQKAVKNSP+PRSYYRCTS C VKK VERS D
Sbjct: 628 VKEPRFAFLTKSEIDHLEDGYRWRKYGQKAVKNSPYPRSYYRCTSQKCIVKKRVERSYQD 807
Query: 188 PTIVVTTYEGKHTHPNPIMSRSSA 211
P+IV+TTYEG+H H P R +A
Sbjct: 808 PSIVMTTYEGQHNHHCPATLRGNA 879
>TC89453 similar to GP|9187622|emb|CAB97004.1 WRKY DNA binding protein
{Solanum tuberosum}, partial (56%)
Length = 701
Score = 124 bits (312), Expect = 3e-29
Identities = 61/118 (51%), Positives = 75/118 (62%)
Frame = +1
Query: 85 TPNSSSISSASSEAINDEHNKTVDQTNNQLNKQLKAKKTNQKKPREARIAFMTKSEVDHL 144
T N S++ E + E N + Q KK +KK ++ + AF T+S+VD L
Sbjct: 298 TNNEISVTHDDHEDVKGEEGSV-----NVIIDQQDVKKKGEKKAKKPKYAFQTRSQVDIL 462
Query: 145 EDGYRWRKYGQKAVKNSPFPRSYYRCTSVSCNVKKHVERSLSDPTIVVTTYEGKHTHP 202
+DGYRWRKYGQKAVKN+ FPRSYYRCT CNVKK V+R D +VVTTYEG HTHP
Sbjct: 463 DDGYRWRKYGQKAVKNNKFPRSYYRCTHQGCNVKKQVQRLTKDEGVVVTTYEGVHTHP 636
>TC83050 similar to SP|Q9FGZ4|WR48_ARATH Probable WRKY transcription factor
48 (WRKY DNA-binding protein 48). [Mouse-ear cress],
partial (31%)
Length = 974
Score = 124 bits (311), Expect = 4e-29
Identities = 78/162 (48%), Positives = 96/162 (59%), Gaps = 23/162 (14%)
Frame = +2
Query: 30 SEERSFMELLGVQQNMNNYSDSLLD---LPVVVKEP------PLESDGNGKEYSEVLNSQ 80
S FM+LLG +NN +++ L +P V P N + SEV N+
Sbjct: 158 SSSFGFMDLLGSHDYINNNNNTFLLSDWVPTVATTTTTHHTLPSPGSSNIPDSSEVFNT- 334
Query: 81 QQPATPNSSSISSASSEA-IN---DEHNKTV----------DQTNNQLNKQLKAKKTNQK 126
P +PNSSSISS+S+EA +N ++H + DQ ++ KQLK KK NQK
Sbjct: 335 --PVSPNSSSISSSSNEATVNNTLEQHRSKLSKIEAELEGDDQDQDKTKKQLKPKKKNQK 508
Query: 127 KPREARIAFMTKSEVDHLEDGYRWRKYGQKAVKNSPFPRSYY 168
K RE R AFMTKSEVDHL+DGYRWRKYGQKAVKNSPFPR Y
Sbjct: 509 KEREPRFAFMTKSEVDHLDDGYRWRKYGQKAVKNSPFPRYIY 634
Score = 70.5 bits (171), Expect = 7e-13
Identities = 31/44 (70%), Positives = 35/44 (79%)
Frame = +1
Query: 165 RSYYRCTSVSCNVKKHVERSLSDPTIVVTTYEGKHTHPNPIMSR 208
RSYYRCT+ SC VKK VERS D +IV TTYEG+HTHP+P SR
Sbjct: 823 RSYYRCTTASCGVKKRVERSSDDSSIVXTTYEGQHTHPSPATSR 954
>TC80880 similar to SP|Q9ZQ70|WRK3_ARATH Probable WRKY transcription factor
3 (WRKY DNA-binding protein 3). [Mouse-ear cress],
partial (24%)
Length = 907
Score = 119 bits (299), Expect = 1e-27
Identities = 57/125 (45%), Positives = 77/125 (61%), Gaps = 7/125 (5%)
Frame = +2
Query: 93 SASSEAINDE----HNKTVDQTNNQLNKQLKAKKTNQKKPR---EARIAFMTKSEVDHLE 145
++ SE + D H K ++ + + N ++ R E +I T SEVD L+
Sbjct: 59 TSDSEDVGDRETEVHEKRIEPDSKRRNTEVTVSNPTTSSHRTVTEPKIIVQTTSEVDLLD 238
Query: 146 DGYRWRKYGQKAVKNSPFPRSYYRCTSVSCNVKKHVERSLSDPTIVVTTYEGKHTHPNPI 205
DGYRWRKYGQK VK +P+PRSYY+CT+ CNV+KHVER+ +DP V+TTYEGKH H P
Sbjct: 239 DGYRWRKYGQKVVKGNPYPRSYYKCTTPGCNVRKHVERASTDPKAVITTYEGKHNHDVPA 418
Query: 206 MSRSS 210
+S
Sbjct: 419 AKTNS 433
>TC79565 similar to SP|Q9ZQ70|WRK3_ARATH Probable WRKY transcription factor 3
(WRKY DNA-binding protein 3). [Mouse-ear cress], partial
(35%)
Length = 1602
Score = 116 bits (291), Expect = 9e-27
Identities = 57/124 (45%), Positives = 74/124 (58%), Gaps = 4/124 (3%)
Frame = +2
Query: 91 ISSASSEAINDEHNKTVDQTNNQLNKQLKAKKTN----QKKPREARIAFMTKSEVDHLED 146
+S S +H VD ++ +AK N + + +I T S+VD LED
Sbjct: 899 LSGTSDSEEEGDHETEVDYEPGLKRRKTEAKLLNPALSHRTVSKPKIIVQTTSDVDLLED 1078
Query: 147 GYRWRKYGQKAVKNSPFPRSYYRCTSVSCNVKKHVERSLSDPTIVVTTYEGKHTHPNPIM 206
GYRWRKYGQK VK +P+PRSYY+CT+ CNV+KHVER +DP V+TTYEGKH H P
Sbjct: 1079 GYRWRKYGQKVVKGNPYPRSYYKCTTPGCNVRKHVERVSTDPKAVLTTYEGKHNHDVPAA 1258
Query: 207 SRSS 210
+S
Sbjct: 1259 KTNS 1270
Score = 79.3 bits (194), Expect = 2e-15
Identities = 42/100 (42%), Positives = 52/100 (52%), Gaps = 1/100 (1%)
Frame = +2
Query: 110 TNNQLNKQLKAKKTNQKKPREARIAFMTKSEVDHL-EDGYRWRKYGQKAVKNSPFPRSYY 168
T N + +QL E R+ + VD +DGY WRKYGQK VK FPRSYY
Sbjct: 566 TFNNIAQQLIPNSVEYSSNSEQRLQKSSFVNVDKANDDGYNWRKYGQKQVKGCEFPRSYY 745
Query: 169 RCTSVSCNVKKHVERSLSDPTIVVTTYEGKHTHPNPIMSR 208
+CT SC V K VER D + Y+G+H H P S+
Sbjct: 746 KCTHPSCLVTKKVERDPVDGHVTAIIYKGEHIHQRPRPSK 865
>BG645618 similar to GP|18158619|gb WRKY-like drought-induced protein {Retama
raetam}, partial (37%)
Length = 747
Score = 116 bits (290), Expect = 1e-26
Identities = 63/144 (43%), Positives = 84/144 (57%), Gaps = 6/144 (4%)
Frame = -1
Query: 78 NSQQQPATPNSSSISSASSEAINDEHNKTVDQTNNQLNKQLKAKKTNQ------KKPREA 131
NS +Q ++ +S + ++ N VD ++ +KQ K + N K RE
Sbjct: 597 NSPEQSVVA-TNDLSPEGAGFVSTRTNDGVDD-DDPFSKQRKMELGNADIIPVVKPIREP 424
Query: 132 RIAFMTKSEVDHLEDGYRWRKYGQKAVKNSPFPRSYYRCTSVSCNVKKHVERSLSDPTIV 191
R+ T SE+D L+DGYRWRKYGQK V+ +P PRSYY+CT+ C V+KHVER+ DP V
Sbjct: 423 RVVVQTMSEIDILDDGYRWRKYGQKVVRGNPNPRSYYKCTNAGCPVRKHVERASHDPKAV 244
Query: 192 VTTYEGKHTHPNPIMSRSSAVRAG 215
+TTYEGKH H P SS AG
Sbjct: 243 ITTYEGKHNHDVPAARSSSHDMAG 172
Score = 35.0 bits (79), Expect = 0.035
Identities = 31/102 (30%), Positives = 42/102 (40%), Gaps = 3/102 (2%)
Frame = -1
Query: 190 IVVTTYEGKHTHPNPIMSRSSAVRAGSLLPPPAECTTNFASDQNYDISQYYNQQRQQVL- 248
I+ Y+G H HP P S+ AGS++ E + N AS + N Q V+
Sbjct: 744 IIEIIYKGTHDHPKP--QPSNRYSAGSVMSTQGERSDNRASSLAVRDDKASNSPEQSVVA 571
Query: 249 FNTLS--SLGFPSKNMNATFSQDRPLCNPRVQDNGLLQDVVP 288
N LS GF S N D P R + G D++P
Sbjct: 570 TNDLSPEGAGFVSTRTNDGVDDDDPFSKQRKMELG-NADIIP 448
>TC88624 similar to PIR|S51529|S51529 SPF1 protein - sweet potato, partial
(50%)
Length = 1496
Score = 112 bits (281), Expect = 1e-25
Identities = 65/183 (35%), Positives = 94/183 (50%), Gaps = 20/183 (10%)
Frame = +2
Query: 42 QQNMNNYSDSLLDLPV----VVKEPPLESDGNGKEYSEVLNSQQQPATPNSSSISSASSE 97
++N ++ S++L+ +PV + + S GNG+ ATP +SSIS +
Sbjct: 953 RRNSSSSSNALVVVPVNPINEIHDQSYASHGNGQ--------MDSAATPENSSISIGGDD 1108
Query: 98 AIN----------------DEHNKTVDQTNNQLNKQLKAKKTNQKKPREARIAFMTKSEV 141
DE + N+ + + + + RE R+ T S++
Sbjct: 1109 DFEQSSHQRSRSGGAGDEFDEEEPEAKRWKNEGENEGISAQPASRTVREPRVVVQTTSDI 1288
Query: 142 DHLEDGYRWRKYGQKAVKNSPFPRSYYRCTSVSCNVKKHVERSLSDPTIVVTTYEGKHTH 201
D L+DGYRWRKYGQK VK +P PRSYY+CT +C V+KHVER+ D V+TTYEGKH H
Sbjct: 1289 DILDDGYRWRKYGQKVVKGNPNPRSYYKCTHPNCPVRKHVERASHDLRAVITTYEGKHNH 1468
Query: 202 PNP 204
P
Sbjct: 1469 DVP 1477
Score = 85.9 bits (211), Expect = 2e-17
Identities = 51/153 (33%), Positives = 78/153 (50%)
Frame = +2
Query: 82 QPATPNSSSISSASSEAINDEHNKTVDQTNNQLNKQLKAKKTNQKKPREARIAFMTKSEV 141
Q N+SS+ S S E + + N T N ++ +N ++P++ +++
Sbjct: 614 QTTENNNSSMQSFSPEIASVQTNNT--------NNGFQSDYSNYQQPQQQPTQTLSRRS- 766
Query: 142 DHLEDGYRWRKYGQKAVKNSPFPRSYYRCTSVSCNVKKHVERSLSDPTIVVTTYEGKHTH 201
+DGY WRKYGQK VK S PRSYY+CT +C KK VERS+ + + Y+G H H
Sbjct: 767 ---DDGYNWRKYGQKQVKGSENPRSYYKCTYPNCPTKKKVERSI-EGQVTEIVYKGTHNH 934
Query: 202 PNPIMSRSSAVRAGSLLPPPAECTTNFASDQNY 234
P P +R ++ + + L N DQ+Y
Sbjct: 935 PKPQCTRRNSSSSSNALVVVPVNPINEIHDQSY 1033
>TC88516 similar to GP|14530681|dbj|BAB61053. WRKY DNA-binding protein
{Nicotiana tabacum}, partial (17%)
Length = 1085
Score = 107 bits (268), Expect = 4e-24
Identities = 57/141 (40%), Positives = 80/141 (56%), Gaps = 16/141 (11%)
Frame = +2
Query: 90 SISSASSEAINDEHNKTVDQTN------NQLNKQLKAKKTNQKKP----------REARI 133
SI SS ++ +E + QT+ N L + K K + + +E R+
Sbjct: 71 SIQEDSSASVGEEEFEQTSQTSYSGGNDNALVPEAKRWKGDNENEGYCASASRTVKEPRV 250
Query: 134 AFMTKSEVDHLEDGYRWRKYGQKAVKNSPFPRSYYRCTSVSCNVKKHVERSLSDPTIVVT 193
T SE+D L+DG+RWRKYGQK VK +P RSYY+CT+ CNV+KHVER+ D V+T
Sbjct: 251 VVQTTSEIDILDDGFRWRKYGQKVVKGNPNARSYYKCTAPGCNVRKHVERAAHDIKAVIT 430
Query: 194 TYEGKHTHPNPIMSRSSAVRA 214
TYEGKH H P S+ +++
Sbjct: 431 TYEGKHNHDVPAARGSAGLQS 493
>TC90266 similar to SP|Q9C519|WRK6_ARATH WRKY transcription factor 6 (WRKY
DNA-binding protein 6) (AtWRKY6). [Mouse-ear cress],
partial (15%)
Length = 1376
Score = 99.8 bits (247), Expect = 1e-21
Identities = 69/205 (33%), Positives = 103/205 (49%), Gaps = 7/205 (3%)
Frame = +2
Query: 76 VLNSQQQPATPNSSSISSASSEAI--NDEHNKTVDQTNNQLNK-QLKAKKTNQKKP---R 129
VL ++Q P+ S+ S + AI N+E+N + +N+ ++ +K +Q R
Sbjct: 221 VLPTRQFFDEPSPSNCSKNNGFAIVENNENNMGRNLACEYINEGEINSKIEDQSSEVGCR 400
Query: 130 EARIAFMTKSEVDHLEDGYRWRKYGQKAVKNSPFPRSYYRCT-SVSCNVKKHVERSLSDP 188
AR++ +S+ + DG +WRKYGQK K +P PR+YYRC+ SC V+K V+R D
Sbjct: 401 RARVSIRARSDFAFMVDGCQWRKYGQKTAKGNPCPRAYYRCSMGTSCPVRKQVQRCFKDE 580
Query: 189 TIVVTTYEGKHTHPNPIMSRSSAVRAGSLLPPPAECTTNFASDQNYDISQYYNQQRQQVL 248
++ +TTYEG H H P ++ A S L F + + QY N L
Sbjct: 581 SVFITTYEGNHNHQLPPAAKPIANLTSSAL-------NTFLPTSSTTLQQYGNNLTNTFL 739
Query: 249 FNTLSSLGFPSKNMNATFSQDRPLC 273
F+ S L P+ N ATFS P C
Sbjct: 740 FS--SPLSPPNSNAIATFSPS-PTC 805
>TC79316 similar to GP|11493822|gb|AAG35658.1 transcription factor WRKY4
{Petroselinum crispum}, partial (49%)
Length = 1282
Score = 97.8 bits (242), Expect = 4e-21
Identities = 55/166 (33%), Positives = 89/166 (53%), Gaps = 14/166 (8%)
Frame = +3
Query: 71 KEYSEVLNSQQQPATPNSSSISSASSEAINDEHNKTVDQ-TNNQLNKQLKAKKTNQKKPR 129
+ Y+ + N+ + N S+ I+ +N T N + + KKPR
Sbjct: 339 ENYNTLRNNLMEYMRKNPDHKEKRKSDEISSSNNLMGGGVTGNNSESSSSDEDQSCKKPR 518
Query: 130 E----------ARIAFMTKSEVDHL--EDGYRWRKYGQKAVKNSPFPRSYYRCT-SVSCN 176
+ +R+ F T+S L +DGY+WRKYGQK +++P PR+Y++C+ + SC
Sbjct: 519 QIVETCIKAKVSRVYFKTESSDSSLIVKDGYQWRKYGQKVTRDNPSPRAYFKCSFAPSCT 698
Query: 177 VKKHVERSLSDPTIVVTTYEGKHTHPNPIMSRSSAVRAGSLLPPPA 222
VKK V+RS+ DP++VV TYEG+H HP+P + + R + + P A
Sbjct: 699 VKKKVQRSVDDPSVVVATYEGEHNHPHPSQTEPTNNRCRNQISPVA 836
>TC92972 similar to GP|9187622|emb|CAB97004.1 WRKY DNA binding protein
{Solanum tuberosum}, partial (46%)
Length = 672
Score = 97.1 bits (240), Expect = 7e-21
Identities = 43/90 (47%), Positives = 61/90 (67%), Gaps = 3/90 (3%)
Frame = +2
Query: 116 KQLKAKKTNQKKPREA---RIAFMTKSEVDHLEDGYRWRKYGQKAVKNSPFPRSYYRCTS 172
K K + ++QK +E + F T+S++D L+DG+RWRKYG+K VK++ FPRSYY+CT
Sbjct: 134 KTKKKETSSQKGGKEIMKHKYVFQTRSQIDILDDGFRWRKYGEKMVKDNKFPRSYYKCTY 313
Query: 173 VSCNVKKHVERSLSDPTIVVTTYEGKHTHP 202
C VKK ++R D +V T+YEG H HP
Sbjct: 314 QGCKVKKQIQRHSKDEQVVETSYEGMHIHP 403
>TC86084 similar to GP|21537078|gb|AAM61419.1 putaive DNA-binding protein
{Arabidopsis thaliana}, partial (56%)
Length = 1372
Score = 96.3 bits (238), Expect = 1e-20
Identities = 51/136 (37%), Positives = 75/136 (54%), Gaps = 3/136 (2%)
Frame = +3
Query: 69 NGKEYSEV-LNSQQQPATPNSSS-ISSASSEAINDEHNKTVDQTNNQLNKQLKAKKTNQK 126
NGK+ S + LN PA +S+ S+ +H + D + NK K+ +
Sbjct: 639 NGKQGSSIFLNPAATPAISGGKPPLSAVPSKKRCHDHGEHSDDVSGS-NKCHCVKRRKNR 815
Query: 127 KPREARIAFMTKSEVDHLEDGYRWRKYGQKAVKNSPFPRSYYRCTSV-SCNVKKHVERSL 185
R R+ ++ D D Y WRKYGQK +K SP+PR YY+C++V C +KHVER+
Sbjct: 816 VKRTVRVPAISSKTADIPPDEYSWRKYGQKPIKGSPYPRGYYKCSTVRGCPARKHVERAT 995
Query: 186 SDPTIVVTTYEGKHTH 201
DPT+++ TYEG+H H
Sbjct: 996 DDPTMLIVTYEGEHRH 1043
>TC85800 similar to GP|11320830|dbj|BAB18313. putative WRKY DNA binding
protein {Oryza sativa (japonica cultivar-group)},
partial (37%)
Length = 2028
Score = 95.9 bits (237), Expect = 2e-20
Identities = 53/142 (37%), Positives = 78/142 (54%), Gaps = 6/142 (4%)
Frame = +1
Query: 83 PATPNSSSISSASSEAINDEHNKTVDQTNNQLNKQLKAKKTNQKKP-REARIAFMTKSEV 141
P NS+ + ++ +++ LN A + N + R+AR++ +SE
Sbjct: 832 PQMRNSNGKTGREDSPESETQGWGPNKSQKILNSSNVADQANTEATMRKARVSVRARSEA 1011
Query: 142 DHLEDGYRWRKYGQKAVKNSPFPRSYYRCT-SVSCNVKKHVERSLSDPTIVVTTYEGKHT 200
+ DG +WRKYGQK K +P PR+YYRCT +V C V+K V+R D TI+VTTYEG H
Sbjct: 1012SMISDGCQWRKYGQKMAKGNPCPRAYYRCTMAVGCPVRKQVQRCAEDKTILVTTYEGTHN 1191
Query: 201 HPNP----IMSRSSAVRAGSLL 218
HP P M+ +++ A LL
Sbjct: 1192HPLPPAAMAMASTTSAAASMLL 1257
>TC80708 weakly similar to GP|14530685|dbj|BAB61055. WRKY DNA-binding
protein {Nicotiana tabacum}, partial (20%)
Length = 864
Score = 95.9 bits (237), Expect = 2e-20
Identities = 43/75 (57%), Positives = 53/75 (70%)
Frame = +2
Query: 130 EARIAFMTKSEVDHLEDGYRWRKYGQKAVKNSPFPRSYYRCTSVSCNVKKHVERSLSDPT 189
E+R+ T SE + DGYRWRKYGQK VK + PR+YYRC+S C VKKHVE+S + T
Sbjct: 146 ESRVIVRTTSESGIVNDGYRWRKYGQKMVKGNTNPRNYYRCSSPGCPVKKHVEKSSQNTT 325
Query: 190 IVVTTYEGKHTHPNP 204
V+TTYEG+H H P
Sbjct: 326 TVITTYEGQHDHAPP 370
>TC77278 similar to GP|11493822|gb|AAG35658.1 transcription factor WRKY4
{Petroselinum crispum}, partial (40%)
Length = 1604
Score = 93.2 bits (230), Expect = 1e-19
Identities = 54/162 (33%), Positives = 91/162 (55%), Gaps = 11/162 (6%)
Frame = +2
Query: 80 QQQPATPNSSSISSASSEAINDEHNKTVDQTNNQLNKQLKAKKTNQKKPRE-----ARIA 134
++ P S+S SE+ N+ ++ + NN ++ + + KKPRE A+I+
Sbjct: 467 KRNPEKEVSTSSKKRKSESSNNNNSNLIG-INNGNSESSSTDEESCKKPREEENIKAKIS 643
Query: 135 -FMTKSEVDH----LEDGYRWRKYGQKAVKNSPFPRSYYRCT-SVSCNVKKHVERSLSDP 188
++EV ++DGY WRKYGQK +++P PR+Y++C+ + SC VKK V+RS+ D
Sbjct: 644 RAYVRTEVSDTGLIVKDGYHWRKYGQKVTRDNPCPRAYFKCSFAPSCPVKKKVQRSVDDQ 823
Query: 189 TIVVTTYEGKHTHPNPIMSRSSAVRAGSLLPPPAECTTNFAS 230
+++V TYEG+H HP P S++ S+ C+ + S
Sbjct: 824 SMLVATYEGEHNHPQPPQIESTSGSGRSVNHSSVPCSASLTS 949
>TC83033 similar to PIR|S51529|S51529 SPF1 protein - sweet potato, partial
(17%)
Length = 900
Score = 90.1 bits (222), Expect = 9e-19
Identities = 68/229 (29%), Positives = 106/229 (45%), Gaps = 15/229 (6%)
Frame = +3
Query: 18 FPSYS-FPSVFDLSEERSFMELLGVQQNMNNYSDSLLDLPVVVKEPPLESDGNGKEYSEV 76
+PSYS FP F SE + L ++Y++ + + ++ E + + +SE
Sbjct: 165 YPSYSDFPPGFSPSELLNSPSFLSSSNIPSSYTNEPSTVQSMNQQHSKELEE--RNFSEP 338
Query: 77 LNSQQQPATPNSSSISSASSEAINDE--------HNKTVDQTNNQLNKQLKAK-KTNQKK 127
+ Q Q T + SS S + + N++ QTN K + +TN
Sbjct: 339 -SFQTQKQTMHDLFQSSTSMFQVEEPLKTHDTLIFNESTKQTNFSFEGTTKHEAQTNSSV 515
Query: 128 PREARI-----AFMTKSEVDHLEDGYRWRKYGQKAVKNSPFPRSYYRCTSVSCNVKKHVE 182
P + + + + EDGY+WRKYG+K VK S PRSYY+CT+ C++ K VE
Sbjct: 516 PNSSYFNNTSASMSIREQRKSEEDGYKWRKYGEKQVKGSENPRSYYKCTNPICSMXKKVE 695
Query: 183 RSLSDPTIVVTTYEGKHTHPNPIMSRSSAVRAGSLLPPPAECTTNFASD 231
RSL D I Y+G H HP P ++ A + + P + C + SD
Sbjct: 696 RSLDDGQITQIVYKGSHNHPKPQCTKRRA-NSQYIHQPSSSCNNSMXSD 839
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.310 0.127 0.359
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 9,191,589
Number of Sequences: 36976
Number of extensions: 134259
Number of successful extensions: 799
Number of sequences better than 10.0: 102
Number of HSP's better than 10.0 without gapping: 756
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 769
length of query: 295
length of database: 9,014,727
effective HSP length: 95
effective length of query: 200
effective length of database: 5,502,007
effective search space: 1100401400
effective search space used: 1100401400
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.8 bits)
S2: 58 (26.9 bits)
Medicago: description of AC135465.11


