

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC135464.5 - phase: 0 /pseudo
(1087 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
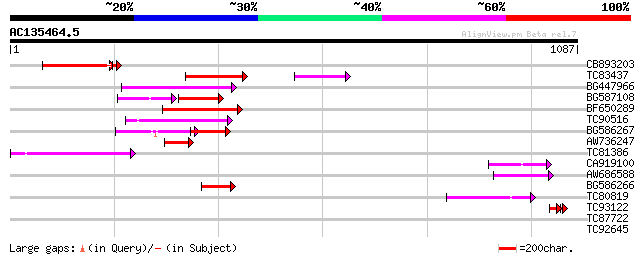
Score E
Sequences producing significant alignments: (bits) Value
CB893203 155 3e-39
TC83437 weakly similar to PIR|D86384|D86384 unknown protein [imp... 140 3e-33
BG447966 weakly similar to PIR|G96509|G96 protein F27F5.21 [impo... 125 7e-29
BG587108 similar to PIR|G96509|G9 protein F27F5.21 [imported] - ... 77 4e-24
BF650289 weakly similar to GP|9049283|dbj| orf129a {Beta vulgari... 107 2e-23
TC90516 106 4e-23
BG586267 weakly similar to PIR|A84888|A8 hypothetical protein At... 102 8e-22
AW736247 weakly similar to PIR|T14619|T146 reverse transcriptase... 79 1e-14
TC81386 similar to GP|10140689|gb|AAG13524.1 putative non-LTR re... 75 1e-13
CA919100 homologue to PIR|G90291|G902 endoglucanase precursor [i... 58 2e-08
AW686588 57 5e-08
BG586266 similar to GP|7267666|em RNA-directed DNA polymerase-li... 52 1e-06
TC80819 weakly similar to GP|10140689|gb|AAG13524.1 putative non... 51 3e-06
TC93122 40 7e-04
TC87722 similar to SP|O24496|GL2C_ARATH Hydroxyacylglutathione h... 32 1.3
TC92645 30 6.6
>CB893203
Length = 800
Score = 155 bits (392), Expect(2) = 3e-39
Identities = 69/138 (50%), Positives = 92/138 (66%)
Frame = -3
Query: 63 LSEEWCLQWPNSFQVALLRGLSDHCPLQLSVDEENWGPRPSRMLKCWQDLPGYNQFVKDK 122
+SE WC++WPN Q AL RGLSD CP+ L++DE NWGPR MLKCW DLPGY+ FVKDK
Sbjct: 798 VSESWCIKWPNMIQRALFRGLSDRCPIMLTIDEGNWGPRLHHMLKCWADLPGYHLFVKDK 619
Query: 123 LKSFQIEGWGGFVLREKLKFIKTALKEWHIGHASNIPGKIDSLKKRQADFDDKGDVGDLS 182
SFQ+ WGG+VL+EK K ++ LK+WH H N+ +I+ +++ A D KG+ L
Sbjct: 618 WNSFQVSRWGGYVLKEK*KMVRNNLKDWHHNHTRNLDARINDIRESIAYLDSKGEDLTLD 439
Query: 183 ADEILELRGITHDLHSLS 200
+E+ DLH+LS
Sbjct: 438 TEEV-------DDLHTLS 406
Score = 26.2 bits (56), Expect(2) = 3e-39
Identities = 10/16 (62%), Positives = 15/16 (93%)
Frame = -1
Query: 198 SLSRVNSSILWQQSRL 213
SLS++N+SI WQ+SR+
Sbjct: 392 SLSKLNTSIQWQKSRI 345
>TC83437 weakly similar to PIR|D86384|D86384 unknown protein [imported] -
Arabidopsis thaliana, partial (6%)
Length = 951
Score = 140 bits (352), Expect = 3e-33
Identities = 71/120 (59%), Positives = 95/120 (79%), Gaps = 1/120 (0%)
Frame = +2
Query: 338 RFISDFHRNGKLTRGINTTFIALISKIDSPQRINDFRPISLVGSLYKILSKVLANRLKLV 397
RF+S+FHRN KL +GIN+TFIALI K+D+PQR+NDFRPISLVGSLYKIL K+LANRL++V
Sbjct: 5 RFVSEFHRNRKLFKGINSTFIALIPKVDNPQRLNDFRPISLVGSLYKILGKLLANRLRVV 184
Query: 398 LGKIISDSQTAFVKDRQILDGILI-ANEVVDEARKNKRN*CCSKLILRRLLTR*IGGTWI 456
+G +ISD+Q+AFVK+RQIL+ + + + R ++ CC + IL+RL+T IG WI
Sbjct: 185 IGSVISDAQSAFVKNRQILEMVFL*QMRLWMRLRN*RKIFCCLRWILKRLITLSIGLIWI 364
Score = 51.2 bits (121), Expect = 2e-06
Identities = 41/106 (38%), Positives = 53/106 (49%)
Frame = +3
Query: 547 NSQMILFFLVLKVGQMCVPYVRFCLYLPRCLV*KLISTKVYW*VSILRIRGYMRRLLF*I 606
N Q L LK+GQ V L+ CLV*+ I +V W V I + G +RRLLF*+
Sbjct: 555 NLQTTLCC*KLKIGQTSVL*GLHLLFFRLCLV*R*IFIRVAWFV*ISLLLG*VRRLLF*V 734
Query: 607 VRWVRFRLCIWAYRLVVILGG*LFGTLC*KQLNLDCWGGKVVSFLM 652
+W R+ C ILG *+FG L * L G VV +L+
Sbjct: 735 GKWARYHFCT*VCLSREILGA*VFGNLL*IA*KLG*LVGIVVFYLL 872
>BG447966 weakly similar to PIR|G96509|G96 protein F27F5.21 [imported] -
Arabidopsis thaliana, partial (7%)
Length = 687
Score = 125 bits (315), Expect = 7e-29
Identities = 80/227 (35%), Positives = 123/227 (53%), Gaps = 5/227 (2%)
Frame = +2
Query: 214 LWLKDGDANSKYFHSVLSGRRRRNSIISLLADS-NLVEGVQPIRNAVLSHFRDHFAARHT 272
+WLKDGD N+K+FHS S RR+ N I L ++ N +G + + ++++F + F + +
Sbjct: 2 VWLKDGDKNTKFFHSKASQRRKVNEIKKLKDETGNWCKGEENVERLLITYFNNLFTSSNP 181
Query: 273 SRVGIDNLPFKC-LSHTDGSGLTKPFSADEVKSAVWDCDSYKSPGPDGVNFGFIKEFWDE 331
+ + K LSH K F+ +EV A+ K+PGPDG+ F +++W
Sbjct: 182 TAIEETCEVVKGKLSHEHIVWCEKEFTEEEVLEAINQMHPVKAPGPDGLPALFFQKYWHI 361
Query: 332 LKEDVMRFISDFHRNGKLTRGINTTFIALISKIDSPQRINDFRPISLVGSLYKILSKVLA 391
+ ++V + + N T +N TFI LI K +P D+RPISL + KI++KV+A
Sbjct: 362 VGKEVQQMVLQVLNNSMETEELNKTFIVLIPKGKNPNTPKDYRPISLCNVVMKIITKVIA 541
Query: 392 NRLKLVLGKIISDSQTAFVKDRQILDGILIANEVV---DEARKNKRN 435
NR+K L +I Q+AFV+ R I D LIA V RK KR+
Sbjct: 542 NRVKQTLPDVIDVEQSAFVQGRLITDNALIAWSVSIG*RXRRKGKRS 682
>BG587108 similar to PIR|G96509|G9 protein F27F5.21 [imported] - Arabidopsis
thaliana, partial (17%)
Length = 677
Score = 76.6 bits (187), Expect(2) = 4e-24
Identities = 39/87 (44%), Positives = 56/87 (63%)
Frame = +3
Query: 324 FIKEFWDELKEDVMRFISDFHRNGKLTRGINTTFIALISKIDSPQRINDFRPISLVGSLY 383
F + W +K D+++ ++ F +GKL +NTT I LI K P R+ + RPISL Y
Sbjct: 411 FFQHSWHIIKMDLLKMVNSFLASGKLDTRLNTTNICLIPKKKRPTRMTELRPISLCNVGY 590
Query: 384 KILSKVLANRLKLVLGKIISDSQTAFV 410
KI+SKVL RLK+ L +IS++Q+AFV
Sbjct: 591 KIISKVLCQRLKVCLPSLISETQSAFV 671
Score = 54.3 bits (129), Expect(2) = 4e-24
Identities = 36/118 (30%), Positives = 57/118 (47%), Gaps = 5/118 (4%)
Frame = +2
Query: 208 WQQ-SRLLWLKDGDANSKYFHSVLSGRRRRNSIISLLA-DSNLVEGVQPIRNAVLSHFRD 265
WQQ SR +W GD N K++H++ R RN I+ L D N + Q + + +F D
Sbjct: 53 WQQKSRNMWHISGDLNKKFYHALTKQRHARNRIVGLYDYDGNWITEEQGVEKVAVDYFED 232
Query: 266 HFAARHTSRVGIDNLPFKCLSHTD---GSGLTKPFSADEVKSAVWDCDSYKSPGPDGV 320
F + T+ G D + S L + + +EV+ A++ K+PGPDG+
Sbjct: 233 LF--QRTTPTGFDGFLDEITSSITPQMNQRLLRLATEEEVRLALFIMHPEKAPGPDGM 400
>BF650289 weakly similar to GP|9049283|dbj| orf129a {Beta vulgaris}, partial
(69%)
Length = 616
Score = 107 bits (268), Expect = 2e-23
Identities = 58/154 (37%), Positives = 96/154 (61%), Gaps = 1/154 (0%)
Frame = +3
Query: 293 LTKPFSADEVKSAVWDCDSYKSPGPDGVNFGFIKEFWDELKEDVMRFISDFHRNGKLTRG 352
L F+A EVK+A++ DS K+PG DG N F K W+ + + V+ I DF + G + +
Sbjct: 12 LCSEFTAVEVKNALFSMDSSKAPGIDGYNVHFFKCSWNIIGDSVIDAILDFFKTGFMPKI 191
Query: 353 INTTFIALISKIDSPQRINDFRPISLVGSLYKILSKVLANRLKLVLGKIISDSQTAFVKD 412
IN T++ L+ K + + +FRPI+ +YKI+SK+L +R++ VL ++S++Q+AFVK
Sbjct: 192 INCTYVTLLPKEVNVTSVKNFRPIACCSVIYKIISKILTSRMQGVLNSVVSENQSAFVKG 371
Query: 413 RQILDGILIANEVVDE-ARKNKRN*CCSKLILRR 445
R I D I++++E+V +RK C K+ L +
Sbjct: 372 RVIFDNIILSHELVKSYSRKGISPRCMVKIDLXK 473
>TC90516
Length = 983
Score = 106 bits (265), Expect = 4e-23
Identities = 65/205 (31%), Positives = 111/205 (53%)
Frame = -2
Query: 222 NSKYFHSVLSGRRRRNSIISLLADSNLVEGVQPIRNAVLSHFRDHFAARHTSRVGIDNLP 281
N+ YF + + R RNSI + + ++ V + A+++ F HF+ T R + ++
Sbjct: 742 NTTYFLACVKNRGMRNSISAPRVE-RWMDDVAKNKQAIVNFFTHHFSDP*TYRPTMGDID 566
Query: 282 FKCLSHTDGSGLTKPFSADEVKSAVWDCDSYKSPGPDGVNFGFIKEFWDELKEDVMRFIS 341
F +S+ D L+ F +++ V D +SP PDG N F + LK D+
Sbjct: 565 FFHISNLDNVLLSAQFLVSKMELVVSSLDGNESPRPDGFNLNFFIRLRNMLKADIEIMFE 386
Query: 342 DFHRNGKLTRGINTTFIALISKIDSPQRINDFRPISLVGSLYKILSKVLANRLKLVLGKI 401
F+ L + + F+ LISK++ P + DF +S +GSL K+++KVLA RL ++ KI
Sbjct: 385 QFYTPANLLKIFS*YFLTLISKVEYPILLGDFSLMSFLGSL*KLMAKVLALRLAHIMEKI 206
Query: 402 ISDSQTAFVKDRQILDGILIANEVV 426
I +Q+ FV+ RQ +DG++ NE++
Sbjct: 205 IFVNQSTFVRGRQHVDGVVAINEII 131
>BG586267 weakly similar to PIR|A84888|A8 hypothetical protein At2g45230
[imported] - Arabidopsis thaliana, partial (16%)
Length = 794
Score = 102 bits (254), Expect = 8e-22
Identities = 62/168 (36%), Positives = 95/168 (55%), Gaps = 7/168 (4%)
Frame = +3
Query: 203 NSSILW-QQSRLLWLKDGDANSKYFHSVLSGRRRRNSIISLLADSNLVEGVQP-IRNAVL 260
N I W Q+SRL WL+ GD N+K+FH+V RR +N I+SL+ D + V+ +
Sbjct: 120 NEEIFWMQKSRLNWLRSGDRNTKFFHAVTKNRRAQNRILSLIDDDDKEWFVEEDLGRLAD 299
Query: 261 SHFRDHFAARHTSRVGI-----DNLPFKCLSHTDGSGLTKPFSADEVKSAVWDCDSYKSP 315
SHF+ +++ VGI +++P ++ + L S +EV+ AV+D + +K P
Sbjct: 300 SHFKLLYSSED---VGITLEDWNSIP-AIVTEEQNAQLMAQISREEVREAVFDINPHKCP 467
Query: 316 GPDGVNFGFIKEFWDELKEDVMRFISDFHRNGKLTRGINTTFIALISK 363
GPDG+N F ++FWD + +D+ +F R GKL GIN T I SK
Sbjct: 468 GPDGMNVFFFQQFWDTMGDDLTSMAQEFLRTGKLEEGINKTNIWFGSK 611
Score = 57.8 bits (138), Expect = 2e-08
Identities = 35/77 (45%), Positives = 47/77 (60%)
Frame = +1
Query: 347 GKLTRGINTTFIALISKIDSPQRINDFRPISLVGSLYKILSKVLANRLKLVLGKIISDSQ 406
G L R L+ K +R+ +FRPISL YKI+SKVL+ RLK VL II+++Q
Sbjct: 562 GNLKRESTKQTSGLVPKKLEAKRLVEFRPISLCNVAYKIVSKVLSKRLKSVLPWIITETQ 741
Query: 407 TAFVKDRQILDGILIAN 423
AF + + I D ILIA+
Sbjct: 742 AAFGRRQLISDNILIAH 792
>AW736247 weakly similar to PIR|T14619|T146 reverse transcriptase - beet
retrotransposon (fragment), partial (4%)
Length = 305
Score = 78.6 bits (192), Expect = 1e-14
Identities = 34/58 (58%), Positives = 44/58 (75%), Gaps = 2/58 (3%)
Frame = +1
Query: 297 FSADEVKSAVWDCDSYKS--PGPDGVNFGFIKEFWDELKEDVMRFISDFHRNGKLTRG 352
FS +EV+ AVWDCDS S PGPDGVNF F+KE+W+ +K D +R + +FH NGKL +G
Sbjct: 121 FSEEEVRKAVWDCDSSHS*NPGPDGVNFTFVKEYWELIKVDFLRVVMEFHTNGKLGKG 294
>TC81386 similar to GP|10140689|gb|AAG13524.1 putative non-LTR retroelement
reverse transcriptase {Oryza sativa (japonica
cultivar-group)}, partial (1%)
Length = 798
Score = 75.5 bits (184), Expect = 1e-13
Identities = 64/247 (25%), Positives = 112/247 (44%), Gaps = 6/247 (2%)
Frame = -1
Query: 1 DFNAVRYAEERKSSRGISGV--IDYHHFSDFIVDNSLIDLSLCGRRFTWF---KGDGNAM 55
DFN + + E + S + + +D+ +SD N+L L G FTW +G N
Sbjct: 726 DFNTILGSHEHQGSHTPARLPMLDFQQWSDV---NNLFHLPTRGSAFTWTNGRRGRNNTR 556
Query: 56 SRIDRFLLSEEWCLQWPNSFQVALLRGLSDHCPLQLSVDEENWGPRPS-RMLKCWQDLPG 114
R+DR ++++ + L + SDH P+ + +N S + +K W P
Sbjct: 555 KRLDRSIVNQLMIDNCESLSACTLTKLRSDHFPILFELQTQNIQFSSSFKFMKMWSAHPD 376
Query: 115 YNQFVKDKLKSFQIEGWGGFVLREKLKFIKTALKEWHIGHASNIPGKIDSLKKRQADFDD 174
+K + Q+ G FVL +KLK +K LK W+ N+ ++D+ K D
Sbjct: 375 CINIIKQCWAN-QVVGCPMFVLNQKLKNLKEVLKVWNKNTFGNVHSQVDNAYKELDDIQV 199
Query: 175 KGDVGDLSADEILELRGITHDLHSLSRVNSSILWQQSRLLWLKDGDANSKYFHSVLSGRR 234
K D S + + + +L S + ++S++ W +GD N+ +FH V + +
Sbjct: 198 KIDSIGYSDVLMDQEKAAQLNLESALNIEEVFWHEKSKVNWHCEGDRNTAFFHRV-AKIK 22
Query: 235 RRNSIIS 241
R +S+IS
Sbjct: 21 RTSSLIS 1
>CA919100 homologue to PIR|G90291|G902 endoglucanase precursor [imported] -
Sulfolobus solfataricus, partial (3%)
Length = 789
Score = 58.2 bits (139), Expect = 2e-08
Identities = 47/121 (38%), Positives = 61/121 (49%), Gaps = 1/121 (0%)
Frame = -3
Query: 919 YGISRCRLRFPFLLGDYCVIGCRLRIIFLGVA*STVRFVCVSQGAV-WKRLHLIFLFIAP 977
+G R R+P LLG + VIGC I * + VCV AV W R ++ F +
Sbjct: 583 FGTDRFL*RYPSLLGGFFVIGCLPNRI*SIEV*FQQKLVCVFLVAVPWNRHNICFSLVRI 404
Query: 978 LLVLCGILSGLGSECQVLTLLLLRPISFSLFTLQVTQRLVSHFFNLFGYFVFWWFGTNVI 1037
LL G +G TL+ FSLFT QV RL+S F +LFG+FV ++G NV
Sbjct: 403 LLHF-GRWFVIGLVLLEWTLMFCLITLFSLFTQQVAIRLLSLFSSLFGFFVPGFYGLNVT 227
Query: 1038 I 1038
I
Sbjct: 226 I 224
>AW686588
Length = 567
Score = 56.6 bits (135), Expect = 5e-08
Identities = 44/116 (37%), Positives = 56/116 (47%)
Frame = +2
Query: 927 RFPFLLGDYCVIGCRLRIIFLGVA*STVRFVCVSQGAVWKRLHLIFLFIAPLLVLCGILS 986
RFPF LG IG +L+ I GV R + V AV ++ + FI LL G +S
Sbjct: 152 RFPF*LGV*FGIGFQLKPI**GVGVWQSRQLGVWLDAVLQKPQITCSFIVLLLAQFGSIS 331
Query: 987 GLGSECQVLTLLLLRPISFSLFTLQVTQRLVSHFFNLFGYFVFWWFGTNVIIGFLI 1042
G C L ++L FSL +QV + V + FGYFV WFG N FLI
Sbjct: 332 VRGLACPELIPMILVTTLFSLLLVQVIRGRVDPLCSSFGYFVCGWFGMNATTDFLI 499
>BG586266 similar to GP|7267666|em RNA-directed DNA polymerase-like protein
{Arabidopsis thaliana}, partial (18%)
Length = 789
Score = 52.4 bits (124), Expect = 1e-06
Identities = 24/65 (36%), Positives = 46/65 (69%)
Frame = -3
Query: 368 QRINDFRPISLVGSLYKILSKVLANRLKLVLGKIISDSQTAFVKDRQILDGILIANEVVD 427
+R++++R I+ + YKI++K+L+ R++ +L IIS SQ+AFV R I D +LI ++++
Sbjct: 784 KRVSEYRTIAPCNTQYKIIAKILSKRMQPLLRSIISPSQSAFVPGRAISDNVLITHKILH 605
Query: 428 EARKN 432
R++
Sbjct: 604 YLRQS 590
>TC80819 weakly similar to GP|10140689|gb|AAG13524.1 putative non-LTR
retroelement reverse transcriptase {Oryza sativa
(japonica cultivar-group)}, partial (2%)
Length = 1262
Score = 50.8 bits (120), Expect = 3e-06
Identities = 59/174 (33%), Positives = 78/174 (43%), Gaps = 3/174 (1%)
Frame = +3
Query: 837 LIWLTLVGRVTERCGSGGVR*GRGRMSC*RSVGF*LMVSFCSIISLTGGNGTRILLAVIV 896
L W GR+TE GSG G+ S * SV F + FC I S+T G+G L V
Sbjct: 381 LTWGGWGGRLTEGRGSGHAACLCGKRSV*GSVVFF*ITLFCRITSMTNGDGC*TL*MVTQ 560
Query: 897 WEAPIFILLLHWSLQTFLWETLYGISRCRLRFPFLLGDYCVIGCRLRIIFL--GVA*STV 954
++ I LL QT LW ++GI+ RF +L G G + I+ G +
Sbjct: 561 *KSSIDTLLPQVIFQTGLWLMMFGINIFLQRFLYLCGASFATGFLQKTIWCIEGFFLQQM 740
Query: 955 RFVCVSQGAVWKR-LHLIFLFIAPLLVLCGILSGLGSECQVLTLLLLRPISFSL 1007
R V V GA+ LH F +IA V G+L +G L+ I FSL
Sbjct: 741 RHVYV--GALTPNPLHTSF-YIATFFVHFGLL*EIGLVFHQCLLVSFEHILFSL 893
Score = 29.6 bits (65), Expect = 6.6
Identities = 14/26 (53%), Positives = 16/26 (60%)
Frame = +2
Query: 723 GYGNLIALC*VNGVGGCWWIEADFGI 748
G+ LI C* NGVG WWI+ GI
Sbjct: 38 GWELLIYRC*ENGVGVFWWIKRGCGI 115
>TC93122
Length = 795
Score = 40.0 bits (92), Expect(2) = 7e-04
Identities = 20/23 (86%), Positives = 20/23 (86%)
Frame = -1
Query: 1035 NVIIGFLIISKPQLTI*QIKFDF 1057
NVII FLIISKPQL *QIKFDF
Sbjct: 117 NVIIDFLIISKPQLNY*QIKFDF 49
Score = 21.9 bits (45), Expect(2) = 7e-04
Identities = 9/13 (69%), Positives = 10/13 (76%)
Frame = -2
Query: 1057 FILYGGFKLITPI 1069
FILYGGF+L I
Sbjct: 50 FILYGGFQLTKQI 12
>TC87722 similar to SP|O24496|GL2C_ARATH Hydroxyacylglutathione hydrolase
cytoplasmic (EC 3.1.2.6) (Glyoxalase II) (Glx II).,
partial (84%)
Length = 876
Score = 32.0 bits (71), Expect = 1.3
Identities = 10/22 (45%), Positives = 15/22 (67%)
Frame = +1
Query: 734 NGVGGCWWIEADFGIGSW*RGM 755
+ +G CWW D GIG+W +G+
Sbjct: 172 SSLGSCWWKREDKGIGAWNQGL 237
>TC92645
Length = 561
Score = 29.6 bits (65), Expect = 6.6
Identities = 14/36 (38%), Positives = 20/36 (54%)
Frame = +3
Query: 293 LTKPFSADEVKSAVWDCDSYKSPGPDGVNFGFIKEF 328
LT PFS DE+ +A++ K+ G DG+N F
Sbjct: 306 LTAPFSVDEIWTAIFQMHHDKAFGLDGLNLAVYYRF 413
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.344 0.155 0.538
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 40,145,941
Number of Sequences: 36976
Number of extensions: 681368
Number of successful extensions: 6378
Number of sequences better than 10.0: 32
Number of HSP's better than 10.0 without gapping: 2621
Number of HSP's successfully gapped in prelim test: 372
Number of HSP's that attempted gapping in prelim test: 3638
Number of HSP's gapped (non-prelim): 3254
length of query: 1087
length of database: 9,014,727
effective HSP length: 106
effective length of query: 981
effective length of database: 5,095,271
effective search space: 4998460851
effective search space used: 4998460851
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 38 (21.5 bits)
S2: 64 (29.3 bits)
Medicago: description of AC135464.5


