

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
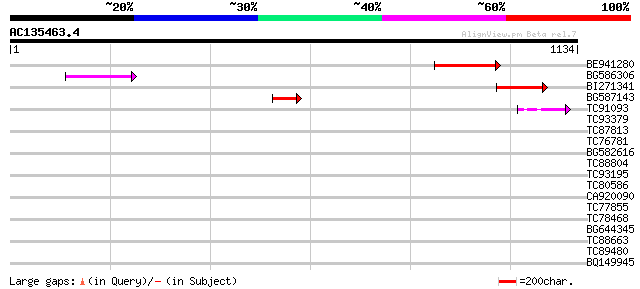
Query= AC135463.4 - phase: 2 /pseudo
(1134 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
BE941280 weakly similar to GP|20197614|g unknown protein {Arabid... 143 4e-34
BG586306 weakly similar to GP|15128241|db helicase-like protein ... 85 1e-16
BI271341 83 7e-16
BG587143 weakly similar to PIR|A96586|A9 hypothetical protein F2... 82 2e-15
TC91093 weakly similar to PIR|T01873|T01873 hypothetical protein... 71 3e-12
TC93379 similar to GP|19569994|gb|AAL92295.1 hypothetical protei... 35 0.13
TC87813 similar to SP|P49299|CYSZ_CUCMA Citrate synthase glyoxy... 32 1.1
TC76781 similar to GP|12539609|gb|AAG59584.1 pathogen-inducible ... 32 1.4
BG582616 weakly similar to GP|11994217|db disease resistance com... 32 1.8
TC88804 similar to GP|7298989|gb|AAF54192.1| CG7878 gene product... 32 1.8
TC93195 weakly similar to GP|10176990|dbj|BAB10222. gb|AAD55291.... 31 3.1
TC80586 30 5.3
CA920090 similar to PIR|F96782|F967 hypothetical protein F22H5.5... 30 5.3
TC77855 similar to SP|O04336|WR21_ARATH Probable WRKY transcript... 30 5.3
TC78468 similar to GP|21594872|gb|AAM66051.1 unknown {Arabidopsi... 30 5.3
BG644345 weakly similar to GP|19697333|gb putative protein poten... 30 5.3
TC88663 similar to GP|8778473|gb|AAF79481.1| F1L3.23 {Arabidopsi... 30 6.9
TC89480 similar to GP|20260684|gb|AAM13240.1 unknown protein {Ar... 30 6.9
BQ149945 weakly similar to GP|2440250|dbj| outer surface protein... 30 6.9
>BE941280 weakly similar to GP|20197614|g unknown protein {Arabidopsis
thaliana}, partial (8%)
Length = 403
Score = 143 bits (360), Expect = 4e-34
Identities = 73/131 (55%), Positives = 92/131 (69%)
Frame = -1
Query: 850 YGGIGKTFLWNALSYHFRSEGKIVLNVASSSIASLLLPRGRTAHSQFSIPLMLLEESGCT 909
YGG KTF+W ALS RSEG+IVL ASS I +LL+P GRTAHS+F IP ++ E S C
Sbjct: 394 YGGTEKTFIWRALSAALRSEGEIVLACASSGIDALLMPGGRTAHSRFGIPFIIDETSMCG 215
Query: 910 IEKEGNKAQLLTMASLIIWDEAPMISRLAFEAFDRTMRDIMSNVVDGASNLPFGGKTVVL 969
+ A L+ A LIIWDEAPM+ + FEA DR++RD++ V + ++PFGGK VVL
Sbjct: 214 VTPNIPLASLVIKAKLIIWDEAPMMHKHCFEALDRSLRDVLKTVDERNKDIPFGGKVVVL 35
Query: 970 GGDFRQILPVV 980
GGDFRQIL V+
Sbjct: 34 GGDFRQILLVM 2
>BG586306 weakly similar to GP|15128241|db helicase-like protein {Oryza
sativa (japonica cultivar-group)}, partial (2%)
Length = 667
Score = 85.1 bits (209), Expect = 1e-16
Identities = 51/142 (35%), Positives = 79/142 (54%)
Frame = +2
Query: 112 LNYHSELSDGNGLDRSLVQDLMKLMDECNLLVKKFRMVRDFREANVDVPVKLRLFRNRNF 171
LN + S LD + + L++++DE N L K FR RD+ + + +RL ++
Sbjct: 227 LNAMGQTSTEGNLDETTLARLIEMIDENNCLAKLFRRARDYYKGSGQ-EFNIRLLSDKG- 400
Query: 172 DSRVYNVPEISEVVALIVGDFDSFEDGRDIVVREKDGGLRRIHETHSKYIPLQYPVLFPL 231
+ Y++P SEV LIVGD S RDIVV+ + L++I + HS Y+ LQYP+LFP
Sbjct: 401 KGKEYDLPSTSEVAGLIVGDMSSTIGVRDIVVQFQSDTLQQIRDYHSLYMSLQYPLLFPY 580
Query: 232 GEDRYEEKIKLNHRTTTGSVKK 253
GE + +I L+ T T ++
Sbjct: 581 GEYGFHPEIPLHLETGTSRTRQ 646
>BI271341
Length = 468
Score = 82.8 bits (203), Expect = 7e-16
Identities = 44/103 (42%), Positives = 64/103 (61%)
Frame = +1
Query: 973 FRQILPVVPKRGRADIVDACINSSMLWRRCRVLKLTKNIRLQFPTNGQDDASLRLFARWI 1032
FR ILPV+P+ R+DI+ A INSS + C+V++L KN+ LQ +D F++ I
Sbjct: 1 FR*ILPVIPRGSRSDIIHATINSSCI*DHCQVVRLKKNMWLQQNGQSSNDPEFEQFSK*I 180
Query: 1033 LDVGDGKLGIDNDGEAVIEIPDDICLKNSGNHVGDIVESTYPN 1075
L VGDGK+ ND A I+IP ++ + N + + IV+STY N
Sbjct: 181 LKVGDGKIYEPNDSYADIDIPPELLISNYDDSLQTIVQSTYQN 309
>BG587143 weakly similar to PIR|A96586|A9 hypothetical protein F20D21.24
[imported] - Arabidopsis thaliana, partial (12%)
Length = 717
Score = 81.6 bits (200), Expect = 2e-15
Identities = 35/58 (60%), Positives = 49/58 (84%)
Frame = -3
Query: 526 VLKNGIQLDNRSVVPYNPKLIMKYHAHINIEFCNKSNSIKYLFKYITKGVDRVTATLE 583
V+KNG L+N +VP+N KL+ KY AHIN+E+CN+++++KYLFKYITKGVDRV+ +E
Sbjct: 265 VVKNGAILNNTFIVPHNIKLLKKYEAHINMEWCNRTSAVKYLFKYITKGVDRVSVVIE 92
Score = 29.3 bits (64), Expect = 9.0
Identities = 12/34 (35%), Positives = 20/34 (58%)
Frame = -3
Query: 314 LLWLDSRDKFDSPKAINSIICAELPDSQMYPELF 347
LLW + + S + ++ II AELP+ + PE +
Sbjct: 541 LLWFGNSSRTPSSEEVDEIISAELPNKKQDPEAY 440
>TC91093 weakly similar to PIR|T01873|T01873 hypothetical protein T24M8.10 -
Arabidopsis thaliana, partial (11%)
Length = 701
Score = 70.9 bits (172), Expect = 3e-12
Identities = 40/107 (37%), Positives = 58/107 (53%)
Frame = +3
Query: 1015 FPTNGQDDASLRLFARWILDVGDGKLGIDNDGEAVIEIPDDICLKNSGNHVGDIVESTYP 1074
FP + + FA ++ GK+ ND A ++ P G+ + IV+STYP
Sbjct: 279 FPNIPRTKVEFKTFA----EILTGKMSEPNDSYAEVDTPP-------GDPIDAIVQSTYP 425
Query: 1075 NLLSNMKESSFFQDRAILAPTLELVEKVNDYVLSLVPGEEKEYLSCD 1121
NL+S F Q RAIL T E+V+++NDYVL L+PGEE+ S +
Sbjct: 426 NLVSQYNNEQFLQSRAILTSTDEVVDQINDYVLKLIPGEERVIYSAN 566
>TC93379 similar to GP|19569994|gb|AAL92295.1 hypothetical protein
{Dictyostelium discoideum}, partial (1%)
Length = 535
Score = 35.4 bits (80), Expect = 0.13
Identities = 12/40 (30%), Positives = 20/40 (50%)
Frame = -2
Query: 403 PHVVNEVLMMMYMMCCLCCYKMMFMMCCLCCYKMMFMMCC 442
P+++ M+ CC+CC +CC CC ++ M C
Sbjct: 267 PYIICCGYMLFMFPCCICCCCSCCCICCCCCCCIIIAMYC 148
Score = 35.4 bits (80), Expect = 0.13
Identities = 11/33 (33%), Positives = 17/33 (51%)
Frame = -2
Query: 410 LMMMYMMCCLCCYKMMFMMCCLCCYKMMFMMCC 442
++ M+ C CC + CC CC ++ M CC
Sbjct: 243 MLFMFPCCICCCCSCCCICCCCCCCIIIAMYCC 145
Score = 31.2 bits (69), Expect = 2.4
Identities = 9/17 (52%), Positives = 11/17 (63%)
Frame = -2
Query: 417 CCLCCYKMMFMMCCLCC 433
CC CC ++ M CC CC
Sbjct: 186 CCCCCCIIIAMYCCCCC 136
Score = 29.6 bits (65), Expect = 6.9
Identities = 10/27 (37%), Positives = 13/27 (48%)
Frame = -2
Query: 416 MCCLCCYKMMFMMCCLCCYKMMFMMCC 442
+CC CC ++ M C CC CC
Sbjct: 192 ICCCCCCCIIIAMYCCCCCCC*CCCCC 112
Score = 29.3 bits (64), Expect = 9.0
Identities = 10/25 (40%), Positives = 13/25 (52%)
Frame = -2
Query: 409 VLMMMYMMCCLCCYKMMFMMCCLCC 433
+++ MY CC CC CC CC
Sbjct: 168 IIIAMYCCCCCCC------*CCCCC 112
Score = 29.3 bits (64), Expect = 9.0
Identities = 11/34 (32%), Positives = 14/34 (40%)
Frame = -2
Query: 409 VLMMMYMMCCLCCYKMMFMMCCLCCYKMMFMMCC 442
+ M +CC CC CC CC + CC
Sbjct: 240 LFMFPCCICC-CCSCCCICCCCCCCIIIAMYCCC 142
>TC87813 similar to SP|P49299|CYSZ_CUCMA Citrate synthase glyoxysomal
precursor (EC 4.1.3.7) (GCS). [Pumpkin Winter squash],
partial (57%)
Length = 1235
Score = 32.3 bits (72), Expect = 1.1
Identities = 11/32 (34%), Positives = 18/32 (55%), Gaps = 2/32 (6%)
Frame = -1
Query: 409 VLMMMYMMCCL--CCYKMMFMMCCLCCYKMMF 438
V+ + + CC CC +F+ CLCC+ + F
Sbjct: 209 VVHLRFRCCCR*HCCTMFVFLCLCLCCFSLFF 114
>TC76781 similar to GP|12539609|gb|AAG59584.1 pathogen-inducible
alpha-dioxygenase {Nicotiana attenuata}, partial (94%)
Length = 2232
Score = 32.0 bits (71), Expect = 1.4
Identities = 14/46 (30%), Positives = 22/46 (47%), Gaps = 1/46 (2%)
Frame = +3
Query: 404 HVVNEVLMMMYMMCCLCCYKMMFMMCCLCCYKMMFM-MCCYKMMFM 448
H N L + + +K+ +CC+C Y FM MC Y +F+
Sbjct: 1929 HQTNTTLFQFTCVFLVNQFKLALYLCCICVYTYTFMFMCMYWCLFI 2066
>BG582616 weakly similar to GP|11994217|db disease resistance comples protein
{Arabidopsis thaliana}, partial (3%)
Length = 679
Score = 31.6 bits (70), Expect = 1.8
Identities = 13/17 (76%), Positives = 13/17 (76%)
Frame = +3
Query: 503 PLCQHDVEFKCFPLHLR 519
PLCQHDV FKCF LR
Sbjct: 570 PLCQHDV*FKCFSAALR 620
>TC88804 similar to GP|7298989|gb|AAF54192.1| CG7878 gene product
{Drosophila melanogaster}, partial (2%)
Length = 1355
Score = 31.6 bits (70), Expect = 1.8
Identities = 11/28 (39%), Positives = 17/28 (60%)
Frame = -2
Query: 418 CLCCYKMMFMMCCLCCYKMMFMMCCYKM 445
C+CC M+ M C+CC M+M C ++
Sbjct: 178 CICCMTMLGMDMCICCCCCMWMG*CIRI 95
>TC93195 weakly similar to GP|10176990|dbj|BAB10222.
gb|AAD55291.1~gene_id:MYH19.18~similar to unknown
protein {Arabidopsis thaliana}, partial (21%)
Length = 737
Score = 30.8 bits (68), Expect = 3.1
Identities = 11/24 (45%), Positives = 12/24 (49%)
Frame = -2
Query: 410 LMMMYMMCCLCCYKMMFMMCCLCC 433
L MY CC CC + CC CC
Sbjct: 289 LRSMYGDCCFCCCWVRV*RCCCCC 218
>TC80586
Length = 1044
Score = 30.0 bits (66), Expect = 5.3
Identities = 10/26 (38%), Positives = 17/26 (64%)
Frame = +2
Query: 400 VLYPHVVNEVLMMMYMMCCLCCYKMM 425
V +PH+ +VLM+MY C +C + +
Sbjct: 542 VAFPHIYVDVLMLMYCECYVCFFNFL 619
>CA920090 similar to PIR|F96782|F967 hypothetical protein F22H5.5 [imported]
- Arabidopsis thaliana, partial (12%)
Length = 859
Score = 30.0 bits (66), Expect = 5.3
Identities = 13/40 (32%), Positives = 17/40 (42%)
Frame = +3
Query: 405 VVNEVLMMMYMMCCLCCYKMMFMMCCLCCYKMMFMMCCYK 444
++N + M + CC CC CC CC CC K
Sbjct: 501 ILNMLFMGSIIGCCECC-----CCCCCCCC*WWSSSCCCK 605
>TC77855 similar to SP|O04336|WR21_ARATH Probable WRKY transcription factor
21 (WRKY DNA-binding protein 21). [Mouse-ear cress],
partial (38%)
Length = 1507
Score = 30.0 bits (66), Expect = 5.3
Identities = 8/21 (38%), Positives = 13/21 (61%)
Frame = -1
Query: 414 YMMCCLCCYKMMFMMCCLCCY 434
++ CC CC + + CC CC+
Sbjct: 673 FICCCCCCCCINNLCCCSCCW 611
>TC78468 similar to GP|21594872|gb|AAM66051.1 unknown {Arabidopsis
thaliana}, partial (78%)
Length = 1249
Score = 30.0 bits (66), Expect = 5.3
Identities = 11/18 (61%), Positives = 12/18 (66%), Gaps = 2/18 (11%)
Frame = +2
Query: 420 CCYKMMF--MMCCLCCYK 435
CCY F +CCLCCYK
Sbjct: 737 CCYWYSF*SSLCCLCCYK 790
>BG644345 weakly similar to GP|19697333|gb putative protein potential
transcriptional repressor Not4hp - Mus musculus, partial
(7%)
Length = 635
Score = 30.0 bits (66), Expect = 5.3
Identities = 11/28 (39%), Positives = 11/28 (39%)
Frame = -1
Query: 417 CCLCCYKMMFMMCCLCCYKMMFMMCCYK 444
CC CC CC CC CC K
Sbjct: 392 CCCCCC*SCCCCCCCCC*N*SCCCCCEK 309
>TC88663 similar to GP|8778473|gb|AAF79481.1| F1L3.23 {Arabidopsis
thaliana}, partial (34%)
Length = 946
Score = 29.6 bits (65), Expect = 6.9
Identities = 11/27 (40%), Positives = 11/27 (40%)
Frame = +3
Query: 417 CCLCCYKMMFMMCCLCCYKMMFMMCCY 443
CC C Y CC CC CCY
Sbjct: 564 CCFCWYS-----CCCCCCCCSGQCCCY 629
>TC89480 similar to GP|20260684|gb|AAM13240.1 unknown protein {Arabidopsis
thaliana}, partial (43%)
Length = 1068
Score = 29.6 bits (65), Expect = 6.9
Identities = 10/27 (37%), Positives = 12/27 (44%)
Frame = -2
Query: 417 CCLCCYKMMFMMCCLCCYKMMFMMCCY 443
CC CC + CC CC CC+
Sbjct: 320 CCSCCCCCCRIRCCCCC**SCCCCCCW 240
>BQ149945 weakly similar to GP|2440250|dbj| outer surface protein A {Borrelia
garinii}, partial (10%)
Length = 648
Score = 29.6 bits (65), Expect = 6.9
Identities = 9/34 (26%), Positives = 16/34 (46%)
Frame = +2
Query: 410 LMMMYMMCCLCCYKMMFMMCCLCCYKMMFMMCCY 443
L M++ C C + + C+CC +CC+
Sbjct: 332 LFFMFLCDCRVCVVLCVYLLCVCCVLRRAFICCF 433
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.337 0.148 0.482
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 37,533,872
Number of Sequences: 36976
Number of extensions: 572905
Number of successful extensions: 6402
Number of sequences better than 10.0: 41
Number of HSP's better than 10.0 without gapping: 3136
Number of HSP's successfully gapped in prelim test: 282
Number of HSP's that attempted gapping in prelim test: 2292
Number of HSP's gapped (non-prelim): 4219
length of query: 1134
length of database: 9,014,727
effective HSP length: 107
effective length of query: 1027
effective length of database: 5,058,295
effective search space: 5194868965
effective search space used: 5194868965
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.7 bits)
S2: 64 (29.3 bits)
Medicago: description of AC135463.4


