

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC135463.3 + phase: 0 /pseudo
(924 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
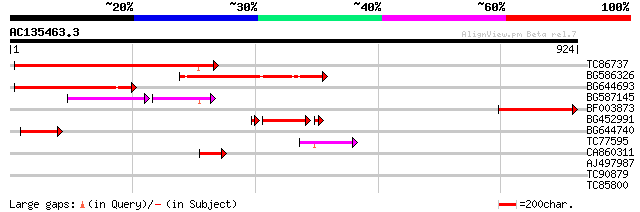
Score E
Sequences producing significant alignments: (bits) Value
TC86737 weakly similar to GP|6683624|dbj|BAA89272.1 Pol {Alterna... 276 2e-74
BG586326 similar to PIR|G84493|G8 probable retroelement pol poly... 206 3e-53
BG644693 weakly similar to GP|18767374|g Putative 22 kDa kafirin... 196 4e-50
BG587145 similar to PIR|H86337|H8 protein F5M15.26 [imported] - ... 87 3e-30
BF003873 similar to GP|14715222|em putative polyprotein {Cicer a... 128 9e-30
BG452991 PIR|A25875|A25 histone H4 - Tetrahymena thermophila, pa... 117 3e-29
BG644740 similar to PIR|A84460|A84 probable retroelement pol pol... 87 4e-17
TC77595 weakly similar to PIR|T18350|T18350 probable pol polypro... 50 4e-06
CA860311 weakly similar to GP|7289872|gb|A CG17427 gene product ... 44 3e-04
AJ497987 weakly similar to GP|9927273|dbj Similar to Arabidopsis... 37 0.027
TC90879 homologue to GP|15809990|gb|AAL06922.1 AT5g60530/muf9_18... 29 7.3
TC85800 similar to GP|11320830|dbj|BAB18313. putative WRKY DNA b... 29 9.5
>TC86737 weakly similar to GP|6683624|dbj|BAA89272.1 Pol {Alternaria
alternata}, partial (21%)
Length = 1540
Score = 276 bits (707), Expect = 2e-74
Identities = 149/339 (43%), Positives = 211/339 (61%), Gaps = 7/339 (2%)
Frame = +1
Query: 8 LKKQLEDLLDKKFVRPSASPWGAPVLLVKKKDGTMRLCIDYRQLNKVTIKNRYPLPRIDD 67
LKK LEDLLDK F++ S S GAPVL V+K G +R C+DYR LN +T K+RYPLP I +
Sbjct: 511 LKKTLEDLLDKGFIKASGSAAGAPVLFVRKPGGGIRFCVDYRALNAITKKDRYPLPLISE 690
Query: 68 LMDQLVGANVFSKIDLRSGYHQIKVKDEDMQKTASRTRYGHYEYKVMPFGVTNAPGVFME 127
+ ++ GA F+K+D+ + +H++++KDED +KTA RTRYG +E+ V PFG+T AP F
Sbjct: 691 TLRRVAGARWFTKLDVVAAFHKMRIKDEDQEKTAFRTRYGLFEWIVCPFGLTGAPATFQR 870
Query: 128 YMNRIFHAYLDKFVVVFIDDILIYSR-TEEEHA*HLKIVL*VLKEKKLYAKLSKCEFWLS 186
Y+N+ H +LD FV +IDD+LIY+ ++++H ++ VL L + L KCEF ++
Sbjct: 871 YINKTLHEFLDDFVTAYIDDVLIYTTGSKKDHEAQVRRVLRRLADAGLSLDPKKCEFSVT 1050
Query: 187 EVSFPGHII-SGSGIAVDPSKVDAVSQWETPKSVTEIRSFLGLAGYYRRFIEGFSKLALP 245
V + G I+ +G G++ DP K+ A+ W P SV RSFLG YY+ FI G+S++ P
Sbjct: 1051TVKYVGFILTAGKGVSCDPLKLAAIRDWLPPGSVKGARSFLGFCNYYKDFIPGYSEITEP 1230
Query: 246 LTQLTCKGKSFVWDAQCESSFNELKQRLPTNPILILPKPEEPFVVYFDASKLGLGVVLMQ 305
LT+LT K F W A+ E++F +LK+ P+L + PE V D S LG VL Q
Sbjct: 1231LTRLTRKDFPFRWGAEQEAAFTKLKRLFAEEPVLRMFDPEAVTTVETDCSGFALGGVLTQ 1410
Query: 306 D-----GKVVAYASRLLRIHEKNYPTHDLELAAVVLVLK 339
+ VA+ S+ L E NYP HD EL AV L+
Sbjct: 1411EDGTGAAHPVAFHSQRLSPAEYNYPIHDKELLAVWACLR 1527
>BG586326 similar to PIR|G84493|G8 probable retroelement pol polyprotein
[imported] - Arabidopsis thaliana, partial (13%)
Length = 736
Score = 206 bits (524), Expect = 3e-53
Identities = 118/243 (48%), Positives = 157/243 (64%), Gaps = 1/243 (0%)
Frame = +2
Query: 277 PILILPKPEEPFVVYFDASKLGLGVVLMQDGKVVAYASRLLRIHEKNYPTHDLELAAVVL 336
PIL+LP+ +VVY DAS GLG VL Q KV+AYASR LR HE NYPTHDLE+AAVV
Sbjct: 17 PILVLPELIT-YVVYTDASITGLGCVLTQHEKVIAYASRQLRKHEGNYPTHDLEMAAVVF 193
Query: 337 VLKIWRHYLYGSRFEVFSDHKSLKYLFDQKELNMRQRRWLELLKDYDFGLNYHPGKANVV 396
LKIWR YLYG++ ++ +DHKSLKY+F Q ELN+RQRRW+E + DYD + Y+PGKAN+V
Sbjct: 194 ALKIWRSYLYGAKVQIHTDHKSLKYIFTQPELNLRQRRWMEFVADYDLDITYYPGKANLV 373
Query: 397 ADALSRKTLHMSALMVKEFDLLEQFRDLSLVCELTPQSVQLGMLKINN-DFLNSIREAQE 455
ADALSR+ + +SA +E D L+ + LT + LG+ +N D IR AQ
Sbjct: 374 ADALSRRRVDVSA--EREADDLDGMVRALRLNVLTKATESLGLEAVNQADLFTRIRLAQG 547
Query: 456 VDVKLVDLMVAGNGTKDSDFKVDDQGVLRFRGRICIPDNDELKKLILEESHKSSLSIRPE 515
D L VA N ++++ G + GRI +P++ LK+ I+ E+HKS S+ P
Sbjct: 548 QDENL--QKVAQNDR--TEYQTAKDGTILVNGRISVPNDRSLKEEIMSEAHKSRFSVHPG 715
Query: 516 ATK 518
A +
Sbjct: 716 APR 724
>BG644693 weakly similar to GP|18767374|g Putative 22 kDa kafirin cluster;
Ty3-Gypsy type {Oryza sativa}, partial (15%)
Length = 716
Score = 196 bits (497), Expect = 4e-50
Identities = 107/199 (53%), Positives = 132/199 (65%)
Frame = +2
Query: 8 LKKQLEDLLDKKFVRPSASPWGAPVLLVKKKDGTMRLCIDYRQLNKVTIKNRYPLPRIDD 67
LK QL+DLL+K F++PS P G VL +KKKDG +R+ IDY QLN V IK +YPLP ID+
Sbjct: 119 LKLQLKDLLEKGFIQPSIYP*GVVVLFLKKKDGFLRMSIDYPQLNNVNIKIKYPLPLIDE 298
Query: 68 LMDQLVGANVFSKIDLRSGYHQIKVKDEDMQKTASRTRYGHYEYKVMPFGVTNAPGVFME 127
L D L G+ F KIDLR G HQ +V ED+ KTA R RYGHYE VM FG TN P FME
Sbjct: 299 LFDNLQGSKWFFKIDLRLG*HQHRVIGEDVPKTAFRIRYGHYEILVMSFG*TNPPMAFME 478
Query: 128 YMNRIFHAYLDKFVVVFIDDILIYSRTEEEHA*HLKIVL*VLKEKKLYAKLSKCEFWLSE 187
MNR+F YLD V+VF +DILIYS+ E EH HL++ L VLK+ L ++S +
Sbjct: 479 LMNRVFQDYLDSLVIVFSNDILIYSKNENEHENHLRLALKVLKDIGL-CQISYV*ILVEV 655
Query: 188 VSFPGHIISGSGIAVDPSK 206
F H+ISG G+ VD +
Sbjct: 656 GFFSLHVISGEGLKVDSKR 712
>BG587145 similar to PIR|H86337|H8 protein F5M15.26 [imported] - Arabidopsis
thaliana, partial (13%)
Length = 763
Score = 87.0 bits (214), Expect(2) = 3e-30
Identities = 47/133 (35%), Positives = 74/133 (55%)
Frame = +2
Query: 95 EDMQKTASRTRYGHYEYKVMPFGVTNAPGVFMEYMNRIFHAYLDKFVVVFIDDILIYSRT 154
+D++KTA T G Y YKVMPFG+ NA + +NR+F L + V+IDD+L+ S
Sbjct: 11 DDLEKTAFITDRGTYCYKVMPFGLKNAGSTYQRLVNRMFADKLGNTMEVYIDDMLVKSLR 190
Query: 155 EEEHA*HLKIVL*VLKEKKLYAKLSKCEFWLSEVSFPGHIISGSGIAVDPSKVDAVSQWE 214
+H HLK L E + +KC F ++ F G+I++ GI V+P ++ A+
Sbjct: 191 ATDHLNHLKE*FKTLDEYIMKLNPAKCTFGVTSGEFLGYIVTQQGIEVNPKQITAILDLP 370
Query: 215 TPKSVTEIRSFLG 227
+PK+ E++ G
Sbjct: 371 SPKNSREVQRLTG 409
Score = 64.3 bits (155), Expect(2) = 3e-30
Identities = 36/106 (33%), Positives = 54/106 (49%), Gaps = 4/106 (3%)
Frame = +3
Query: 234 RFIEGFSKLALPLTQLTCKGKSFVWDAQCESSFNELKQRLPTNPILILPKPEEPFVVYFD 293
RFI + LP +L C K FVWD +CE +F +LKQ L T P+L P+ + +Y
Sbjct: 429 RFISRSTDKCLPFYKLLCGNKRFVWDEKCEEAFEQLKQYLTTPPVLSKPEAGDTLSLYIA 608
Query: 294 ASKLGLGVVLMQDG----KVVAYASRLLRIHEKNYPTHDLELAAVV 335
S + VL+++ K + Y S+ + E YPT + AV+
Sbjct: 609 ISSTAVSSVLIREDRGEQKPIFYTSKRMTDPETRYPTLEKMAFAVI 746
>BF003873 similar to GP|14715222|em putative polyprotein {Cicer arietinum},
partial (82%)
Length = 559
Score = 128 bits (322), Expect = 9e-30
Identities = 80/128 (62%), Positives = 89/128 (69%)
Frame = +3
Query: 797 DVH*SRRS*LLSSLVRIRFQRGLEQWHIELVCHHIFRICMMYFMCRNFGSM*RILRMLFR 856
DV *S+RS*L SLVRIR+Q+ LE+W IE H IF ICMM FMCRNFGSM RI M R
Sbjct: 9 DVL*SQRS*L*DSLVRIRYQKELERWRIEWDYHRIF*ICMMSFMCRNFGSMFRIHLM*SR 188
Query: 857 EMTCRLETTLQLRPYR*GLKTVR*RS*EARIYL**RLFGVE*LVRV*LGSWRVRCGSRIQ 916
M C+ ETTL+ R YR*GL +*R *EAR YL * FG E +V+ * GS RVR S IQ
Sbjct: 189 VMMCKSETTLR*RLYR*GLMIEK*RH*EARRYLL*ESFGTERMVKA*RGSLRVRWWSLIQ 368
Query: 917 SCLIEVRF 924
SCL EV F
Sbjct: 369 SCLHEVNF 392
>BG452991 PIR|A25875|A25 histone H4 - Tetrahymena thermophila, partial (33%)
Length = 560
Score = 117 bits (293), Expect(3) = 3e-29
Identities = 57/78 (73%), Positives = 69/78 (88%)
Frame = +3
Query: 412 VKEFDLLEQFRDLSLVCELTPQSVQLGMLKINNDFLNSIREAQEVDVKLVDLMVAGNGTK 471
++ + LEQFRDLSLVCE++PQSV+LGMLKINN+FL+SI+EAQ+VDVKLVDLM N T+
Sbjct: 51 LESWSCLEQFRDLSLVCEVSPQSVKLGMLKINNEFLDSIKEAQKVDVKLVDLMFGNNQTE 230
Query: 472 DSDFKVDDQGVLRFRGRI 489
D DFKVDDQGVL+FR RI
Sbjct: 231 DGDFKVDDQGVLQFRDRI 284
Score = 25.4 bits (54), Expect(3) = 3e-29
Identities = 11/13 (84%), Positives = 12/13 (91%)
Frame = +1
Query: 395 VVADALSRKTLHM 407
VVAD LSRKTLH+
Sbjct: 1 VVADVLSRKTLHV 39
Score = 25.0 bits (53), Expect(3) = 3e-29
Identities = 9/15 (60%), Positives = 15/15 (100%)
Frame = +2
Query: 497 LKKLILEESHKSSLS 511
+KK+ILEESH+S+++
Sbjct: 284 MKKMILEESHRSNVN 328
>BG644740 similar to PIR|A84460|A84 probable retroelement pol polyprotein
[imported] - Arabidopsis thaliana, partial (4%)
Length = 754
Score = 86.7 bits (213), Expect = 4e-17
Identities = 42/69 (60%), Positives = 51/69 (73%)
Frame = -1
Query: 18 KKFVRPSASPWGAPVLLVKKKDGTMRLCIDYRQLNKVTIKNRYPLPRIDDLMDQLVGANV 77
K+F +PS SP GA +L V+KKDG R+CIDYRQ NKVT KN+YPLPRID+L D++
Sbjct: 274 KRFQQPSISP*GAALLFVRKKDGYFRMCIDYRQFNKVTTKNKYPLPRIDNLFDKIQEDCY 95
Query: 78 FSKIDLRSG 86
F IDLR G
Sbjct: 94 F*NIDLRLG 68
>TC77595 weakly similar to PIR|T18350|T18350 probable pol polyprotein - rice
blast fungus gypsy retroelement (fragment), partial
(14%)
Length = 1708
Score = 50.1 bits (118), Expect = 4e-06
Identities = 31/101 (30%), Positives = 48/101 (46%), Gaps = 7/101 (6%)
Frame = +2
Query: 473 SDFKVDDQGVLRFRGRICIPDND-------ELKKLILEESHKSSLSIRPEATKMYHDLKK 525
S+ ++D L FRGRI +P +D EL+ +++ESH S+ + P + +
Sbjct: 74 SECQLDSLKRLTFRGRIWVPGSDDEESPLNELRTKLVQESHDSTAAGHPGRNGTLEIVSR 253
Query: 526 LFWWSGLKRDVAQFVYACLTCLKSKVEHQKPAELLTPLDVP 566
F+W G + V +FV C C + Q L PL VP
Sbjct: 254 KFFWPGQSQTVRRFVRNCDVCGGIHIWRQAKRGFLKPLPVP 376
>CA860311 weakly similar to GP|7289872|gb|A CG17427 gene product {Drosophila
melanogaster}, partial (20%)
Length = 192
Score = 43.9 bits (102), Expect = 3e-04
Identities = 19/43 (44%), Positives = 29/43 (67%)
Frame = +1
Query: 310 VAYASRLLRIHEKNYPTHDLELAAVVLVLKIWRHYLYGSRFEV 352
+AYASRLL E+NY + E A + ++ +RHYL+G +FE+
Sbjct: 64 IAYASRLLTAAERNYTVVERECLAAIWAIRNFRHYLHGPKFEL 192
>AJ497987 weakly similar to GP|9927273|dbj Similar to Arabidopsis thaliana
chromosome II BAC F26H6; putative retroelement pol
polyprotein, partial (1%)
Length = 636
Score = 37.4 bits (85), Expect = 0.027
Identities = 24/76 (31%), Positives = 38/76 (49%), Gaps = 3/76 (3%)
Frame = -2
Query: 333 AVVLVLKIWRHYLYGSRFEVFSDHKSLKYLFDQKELNMRQRRWLELLKDYDFGLNYHPGK 392
A+ K RHY+ + S +KY+F++ L R RW LL +YD + Y K
Sbjct: 623 ALAWAAKRLRHYMINHTTWLVSKMDPIKYIFEKPALTGRIARWQMLLSEYD--IEYRSQK 450
Query: 393 A---NVVADALSRKTL 405
A +++AD L+ + L
Sbjct: 449 AIKGSILADHLAHQPL 402
>TC90879 homologue to GP|15809990|gb|AAL06922.1 AT5g60530/muf9_180
{Arabidopsis thaliana}, partial (2%)
Length = 760
Score = 29.3 bits (64), Expect = 7.3
Identities = 26/83 (31%), Positives = 32/83 (38%)
Frame = +2
Query: 204 PSKVDAVSQWETPKSVTEIRSFLGLAGYYRRFIEGFSKLALPLTQLTCKGKSFVWDAQCE 263
PS + S W S I SF L+ RF FS + C SF
Sbjct: 497 PSILSFASSWVFNSSFLSIISFSPLSF---RFTSSFSLAKPSSVKHCCVANSFF--CLSF 661
Query: 264 SSFNELKQRLPTNPILILPKPEE 286
SSF + P N +L+LP P E
Sbjct: 662 SSFTSSNIKPPPNSLLLLPSPFE 730
>TC85800 similar to GP|11320830|dbj|BAB18313. putative WRKY DNA binding
protein {Oryza sativa (japonica cultivar-group)},
partial (37%)
Length = 2028
Score = 28.9 bits (63), Expect = 9.5
Identities = 17/59 (28%), Positives = 26/59 (43%), Gaps = 4/59 (6%)
Frame = -2
Query: 236 IEGFSKLA----LPLTQLTCKGKSFVWDAQCESSFNELKQRLPTNPILILPKPEEPFVV 290
+EGFS + L + LTC W CE L ++ T+P++ P P V+
Sbjct: 422 VEGFSSVETISFLMMLDLTC------WLCSCEGDGLFLSEKKSTSPVIFRSPPSSPLVL 264
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.341 0.151 0.492
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 29,205,182
Number of Sequences: 36976
Number of extensions: 422714
Number of successful extensions: 2919
Number of sequences better than 10.0: 24
Number of HSP's better than 10.0 without gapping: 1622
Number of HSP's successfully gapped in prelim test: 127
Number of HSP's that attempted gapping in prelim test: 1264
Number of HSP's gapped (non-prelim): 1808
length of query: 924
length of database: 9,014,727
effective HSP length: 105
effective length of query: 819
effective length of database: 5,132,247
effective search space: 4203310293
effective search space used: 4203310293
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.9 bits)
S2: 63 (28.9 bits)
Medicago: description of AC135463.3


