

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC135416.5 + phase: 0 /pseudo
(653 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
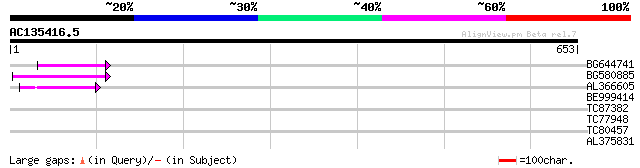
BG644741 69 6e-12
BG580885 57 2e-08
AL366605 46 5e-05
BE999414 34 0.20
TC87382 similar to EGAD|146423|156195 vitellogenin {Anolis pulch... 32 0.59
TC77948 similar to PIR|C85360|C85360 hypothetical protein AT4g30... 24 5.2
TC80457 similar to GP|6682258|gb|AAF23310.1| putative RNA helica... 29 6.5
AL375831 28 8.5
>BG644741
Length = 735
Score = 68.9 bits (167), Expect = 6e-12
Identities = 32/84 (38%), Positives = 46/84 (54%)
Frame = -2
Query: 33 LRLHLFPFSLRDRASAWFHSLEVGSITSWDDMRRALLARFFPPSKTAKLRDQIMRFNQKD 92
L L +FP SL A WF L SI +W+ +R LAR++P SK D++ F
Sbjct: 593 LGLRVFPLSLMGEADIWFTELPYNSIFTWNQLRDVFLARYYPVSKKLNHNDRVNNFVALP 414
Query: 93 GESLYEAWERFKEMLRLCPHHGLE 116
GES+ +W+RF LR P+H ++
Sbjct: 413 GESVSSSWDRFTSFLRSVPNHRID 342
>BG580885
Length = 609
Score = 57.4 bits (137), Expect = 2e-08
Identities = 30/115 (26%), Positives = 62/115 (53%), Gaps = 2/115 (1%)
Frame = +3
Query: 4 FSGSPTEDPNLHISSFLRLSGTIKENQEALRLHLFPFSLRDRASAWFH-SLEVGSIT-SW 61
F G+P E P H+S F ++ + ++ +FP +L + ++ W+ ++E I+ SW
Sbjct: 54 FRGTPNESPITHLSRFNKVCRANNASSVEMQKKIFPVTLEEESALWYDLNIEPYYISLSW 233
Query: 62 DDMRRALLARFFPPSKTAKLRDQIMRFNQKDGESLYEAWERFKEMLRLCPHHGLE 116
D+++ + L ++ +LR ++M +Q + E + + R + +L+ P HGLE
Sbjct: 234 DEIKLSFLQAYYEIEPVEELRSELMGIHQGEKERVRSYFLRLQWILKRWPEHGLE 398
>AL366605
Length = 422
Score = 45.8 bits (107), Expect = 5e-05
Identities = 26/93 (27%), Positives = 49/93 (51%)
Frame = -3
Query: 12 PNLHISSFLRLSGTIKENQEALRLHLFPFSLRDRASAWFHSLEVGSITSWDDMRRALLAR 71
P HI ++R G K+N ++L +H F SL + A+ W+ SL I ++D++ A +
Sbjct: 291 PQNHIIKYVRKMGNYKDN-DSLMIHCFQDSLMEDAAEWYTSLSKNDIHTFDELAAAFKSH 115
Query: 72 FFPPSKTAKLRDQIMRFNQKDGESLYEAWERFK 104
+ ++ R+ + +QK ES E +R++
Sbjct: 114 YGFNTRLKPNREFLRSLSQKKEESFREYAQRWR 16
>BE999414
Length = 613
Score = 33.9 bits (76), Expect = 0.20
Identities = 20/64 (31%), Positives = 32/64 (49%)
Frame = +3
Query: 9 TEDPNLHISSFLRLSGTIKENQEALRLHLFPFSLRDRASAWFHSLEVGSITSWDDMRRAL 68
T DP+ H+ + + + T + + A++ LF +LR A WF +L SI SW D+
Sbjct: 27 T*DPDEHMEN-IEVVLTYRSVRGAVKCKLFVTTLRRGAMTWFKNLRRNSIGSWGDLCHEF 203
Query: 69 LARF 72
F
Sbjct: 204TTHF 215
>TC87382 similar to EGAD|146423|156195 vitellogenin {Anolis pulchellus},
partial (7%)
Length = 2304
Score = 32.3 bits (72), Expect = 0.59
Identities = 22/81 (27%), Positives = 35/81 (43%), Gaps = 13/81 (16%)
Frame = +2
Query: 8 PTEDPNLHISSFLRLSGTI------KENQEALRLHLFPFSLRDRASAWFHSLE------- 54
P + NL + FL TI KE E ++ + L+ A W+ +L+
Sbjct: 629 PDFEGNLQLDDFLDWLQTIERVFEYKEVPEEQKVKIVAAKLKKHALIWWENLKRRRKREG 808
Query: 55 VGSITSWDDMRRALLARFFPP 75
I +WD MR+ L ++ PP
Sbjct: 809 KSKIKTWDKMRQKLTRKYLPP 871
>TC77948 similar to PIR|C85360|C85360 hypothetical protein AT4g30780
[imported] - Arabidopsis thaliana, partial (28%)
Length = 1916
Score = 24.3 bits (51), Expect(2) = 5.2
Identities = 15/43 (34%), Positives = 19/43 (43%)
Frame = -1
Query: 30 QEALRLHLFPFSLRDRASAWFHSLEVGSITSWDDMRRALLARF 72
Q A +L F S + AW E+G + SW AL RF
Sbjct: 1151 QIAEKLIRFRSSAPETFDAWGFITELGEVISWKLFSTALPIRF 1023
Score = 23.1 bits (48), Expect(2) = 5.2
Identities = 8/24 (33%), Positives = 13/24 (53%)
Frame = -2
Query: 100 WERFKEMLRLCPHHGLEKVAYSPH 123
W + ++ L PHH ++K S H
Sbjct: 1012 WRKMIDLQILIPHHAVQKSQLSLH 941
>TC80457 similar to GP|6682258|gb|AAF23310.1| putative RNA helicase
{Arabidopsis thaliana}, partial (28%)
Length = 1425
Score = 28.9 bits (63), Expect = 6.5
Identities = 25/97 (25%), Positives = 41/97 (41%), Gaps = 3/97 (3%)
Frame = +1
Query: 23 SGTIKENQEA-LRLHLFPFSLRDRASAWFHSLEVGSITSWDDMRRALLARFFPPSKTAKL 81
S + E+ EA L ++ FP + R + + H +G I+ W +FFPP K A
Sbjct: 655 SEALPEHYEAELEINDFPQNARWKVT---HKETLGPISEWTGAAITTRGQFFPPGKVAGP 825
Query: 82 RDQ--IMRFNQKDGESLYEAWERFKEMLRLCPHHGLE 116
D+ + +S+ A K +L H L+
Sbjct: 826 GDRKLYLFIEGPSEQSVKRAKAELKRVLEDITHQALQ 936
>AL375831
Length = 467
Score = 28.5 bits (62), Expect = 8.5
Identities = 12/38 (31%), Positives = 22/38 (57%)
Frame = +3
Query: 26 IKENQEALRLHLFPFSLRDRASAWFHSLEVGSITSWDD 63
I + ++L +H F SL + A+ W+ SL I ++D+
Sbjct: 354 ITKTNDSLMIHCFQDSLMEDAAEWYTSLSKNDIHTFDE 467
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.365 0.161 0.617
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 21,947,044
Number of Sequences: 36976
Number of extensions: 329665
Number of successful extensions: 3732
Number of sequences better than 10.0: 16
Number of HSP's better than 10.0 without gapping: 1055
Number of HSP's successfully gapped in prelim test: 146
Number of HSP's that attempted gapping in prelim test: 2628
Number of HSP's gapped (non-prelim): 1286
length of query: 653
length of database: 9,014,727
effective HSP length: 102
effective length of query: 551
effective length of database: 5,243,175
effective search space: 2888989425
effective search space used: 2888989425
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 14 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 36 (21.6 bits)
S2: 62 (28.5 bits)
Medicago: description of AC135416.5


