

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
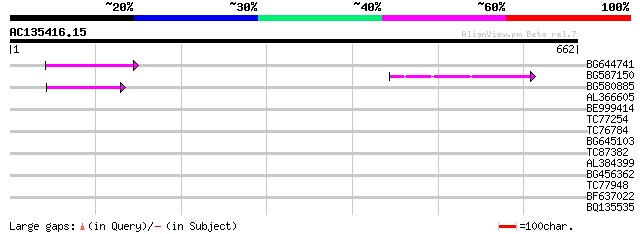
Query= AC135416.15 - phase: 0 /pseudo
(662 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
BG644741 77 2e-14
BG587150 similar to GP|13592175|gb ppg3 {Leishmania major}, part... 56 5e-08
BG580885 49 8e-06
AL366605 37 0.032
BE999414 34 0.21
TC77254 similar to GP|10334497|emb|CAC10210. hypothetical protei... 33 0.27
TC76784 similar to PIR|G84825|G84825 probable CCCH-type zinc fin... 33 0.46
BG645103 30 3.0
TC87382 similar to EGAD|146423|156195 vitellogenin {Anolis pulch... 29 5.1
AL384399 29 6.6
BG456362 homologue to GP|732905|emb| homology with pollen allerg... 29 6.6
TC77948 similar to PIR|C85360|C85360 hypothetical protein AT4g30... 24 8.6
BF637022 similar to GP|7757910|gb| putative translation initiati... 28 8.6
BQ135535 28 8.6
>BG644741
Length = 735
Score = 77.0 bits (188), Expect = 2e-14
Identities = 38/109 (34%), Positives = 56/109 (50%)
Frame = -2
Query: 42 LHLFPFSLRDRARAWFHSLEVGSITSWDDMRRAFLARFFPPSKTAKLRDQITRFNQKDGE 101
L +FP SL A WF L SI +W+ +R FLAR++P SK D++ F GE
Sbjct: 587 LRVFPLSLMGEADIWFTELPYNSIFTWNQLRDVFLARYYPVSKKLNHNDRVNNFVALPGE 408
Query: 102 YLYEAWERFKEMLRLCPHHGLEKWLIVHTFYNGLSYTTKMSVDAAACGA 150
+ +W+RF LR P+H ++ + FY G K+ +D A G+
Sbjct: 407 SVSSSWDRFTSFLRSVPNHRIDDDSLKEYFYRGQDDNNKVVLDTIAGGS 261
>BG587150 similar to GP|13592175|gb ppg3 {Leishmania major}, partial (1%)
Length = 754
Score = 55.8 bits (133), Expect = 5e-08
Identities = 50/170 (29%), Positives = 77/170 (44%)
Frame = +2
Query: 444 ICIKIPFAEALSRMPLYAKFLKEIFSKKKAIDHNETKALTRENSAIIKKPPTKLRDPGSF 503
I ++ PF EA L+ F +E ++ I +A + I K K D G F
Sbjct: 188 IPLEKPFEEAYFTHRLWM-FFRETRETEEDIRRMFCEAREKMKKRITLK---KKSDSGKF 355
Query: 504 AIPCMIGSETLNQALCDLGASVSLLPLPLF*RLGLGELKLTETTLKLADRSDIQPVGNVE 563
AI + ALC+ GASV +LP + LGL ++ ++ + D S G +
Sbjct: 356 AISWTMKGIEFPHALCNTGASVIILPRDMADHLGL-KVDPSKESFTFVDCSQRSSGGIIR 532
Query: 564 DIPVKIEGIDIPTDFMVLDIDEDNECPIILGRPFLATAGAIVDVQNGRII 613
D+ V+I +P DF VLDI + ++L R FL+T V N ++
Sbjct: 533 DLEVQIGNALVPVDFHVLDIKINWNSSLLLRRAFLSTGRGSVQHANQPVL 682
>BG580885
Length = 609
Score = 48.5 bits (114), Expect = 8e-06
Identities = 27/94 (28%), Positives = 52/94 (54%), Gaps = 2/94 (2%)
Frame = +3
Query: 44 LFPFSLRDRARAWFH-SLEVGSIT-SWDDMRRAFLARFFPPSKTAKLRDQITRFNQKDGE 101
+FP +L + + W+ ++E I+ SWD+++ +FL ++ +LR ++ +Q + E
Sbjct: 153 IFPVTLEEESALWYDLNIEPYYISLSWDEIKLSFLQAYYEIEPVEELRSELMGIHQGEKE 332
Query: 102 YLYEAWERFKEMLRLCPHHGLEKWLIVHTFYNGL 135
+ + R + +L+ P HGLE +I F NGL
Sbjct: 333 RVRSYFLRLQWILKRWPEHGLEDDVIKGVFVNGL 434
>AL366605
Length = 422
Score = 36.6 bits (83), Expect = 0.032
Identities = 22/85 (25%), Positives = 44/85 (50%)
Frame = -3
Query: 27 LRLSGAIKENQEAVRLHLFPFSLRDRARAWFHSLEVGSITSWDDMRRAFLARFFPPSKTA 86
+R G K+N +++ +H F SL + A W+ SL I ++D++ AF + + ++
Sbjct: 267 VRKMGNYKDN-DSLMIHCFQDSLMEDAAEWYTSLSKNDIHTFDELAAAFKSHYGFNTRLK 91
Query: 87 KLRDQITRFNQKDGEYLYEAWERFK 111
R+ + +QK E E +R++
Sbjct: 90 PNREFLRSLSQKKEESFREYAQRWR 16
>BE999414
Length = 613
Score = 33.9 bits (76), Expect = 0.21
Identities = 17/41 (41%), Positives = 21/41 (50%)
Frame = +3
Query: 39 AVRLHLFPFSLRDRARAWFHSLEVGSITSWDDMRRAFLARF 79
AV+ LF +LR A WF +L SI SW D+ F F
Sbjct: 93 AVKCKLFVTTLRRGAMTWFKNLRRNSIGSWGDLCHEFTTHF 215
>TC77254 similar to GP|10334497|emb|CAC10210. hypothetical protein {Cicer
arietinum}, partial (62%)
Length = 1985
Score = 33.5 bits (75), Expect = 0.27
Identities = 28/93 (30%), Positives = 47/93 (50%), Gaps = 1/93 (1%)
Frame = +2
Query: 256 QFNQGTRPNQKFYKNPQGSYGQVAPPGYTNNQRVAKKSSLEILLENCIANQNKNLQELKN 315
Q T+ + FYKN + + + V KS L +++E+ IA + + L+ KN
Sbjct: 902 QLGYTTKTKKIFYKNKERD-----SKAFLDIVGVKDKSKL-VVMEDPIAQEKRYLEMRKN 1063
Query: 316 -QTGFLNGSLSKLTTKVDSIATHTKMLKTQISQ 347
+ S+SK++ +VDS+A LKT IS+
Sbjct: 1064IKMERAAKSISKISLEVDSLAGQVSALKTIISK 1162
>TC76784 similar to PIR|G84825|G84825 probable CCCH-type zinc finger protein
[imported] - Arabidopsis thaliana, partial (56%)
Length = 2631
Score = 32.7 bits (73), Expect = 0.46
Identities = 21/60 (35%), Positives = 30/60 (50%)
Frame = +1
Query: 450 FAEALSRMPLYAKFLKEIFSKKKAIDHNETKALTRENSAIIKKPPTKLRDPGSFAIPCMI 509
FA+ L+R + + +FS I HN + R A I+K + L DP S A PCM+
Sbjct: 1012 FAKGLARKGILVS-MHMVFSSHGYIPHNTEQGFARTRLAAIEKSASLLTDPKSCA-PCML 1185
>BG645103
Length = 757
Score = 30.0 bits (66), Expect = 3.0
Identities = 19/49 (38%), Positives = 25/49 (50%)
Frame = +2
Query: 426 TKLNLVTQFTKFLNILQKICIKIPFAEALSRMPLYAKFLKEIFSKKKAI 474
TK++ TQF + N QKI IKI E + ++ K L E FS I
Sbjct: 86 TKIHTTTQF*QPKNTQQKIKIKIKIKEGKLALQIHHKSLMEDFSMYNCI 232
>TC87382 similar to EGAD|146423|156195 vitellogenin {Anolis pulchellus},
partial (7%)
Length = 2304
Score = 29.3 bits (64), Expect = 5.1
Identities = 20/85 (23%), Positives = 36/85 (41%), Gaps = 10/85 (11%)
Frame = +2
Query: 8 KISFLDHPLRIQISIFLPSLRLSGAI---KENQEAVRLHLFPFSLRDRARAWFHSLE--- 61
K+ D +Q+ FL L+ + KE E ++ + L+ A W+ +L+
Sbjct: 617 KVDIPDFEGNLQLDDFLDWLQTIERVFEYKEVPEEQKVKIVAAKLKKHALIWWENLKRRR 796
Query: 62 ----VGSITSWDDMRRAFLARFFPP 82
I +WD MR+ ++ PP
Sbjct: 797 KREGKSKIKTWDKMRQKLTRKYLPP 871
>AL384399
Length = 483
Score = 28.9 bits (63), Expect = 6.6
Identities = 25/112 (22%), Positives = 45/112 (39%), Gaps = 2/112 (1%)
Frame = +3
Query: 239 VDCQLGSAANIEQLNYAQFNQGTRPNQ--KFYKNPQGSYGQVAPPGYTNNQRVAKKSSLE 296
++ L A EQLN + N+ + +K Q S + N + L
Sbjct: 147 INLGLADGAQSEQLNILSEKMDSESNEIIESFKLLQNSTREYLDSLKENKENPDLSYELS 326
Query: 297 ILLENCIANQNKNLQELKNQTGFLNGSLSKLTTKVDSIATHTKMLKTQISQV 348
+ ++N KNLQ L + +S+LTTK + +A + ++S +
Sbjct: 327 LKVKNASQAWQKNLQNLAIKFTSAQSKISELTTKNEFLAKANETDSNKVSDL 482
>BG456362 homologue to GP|732905|emb| homology with pollen allergens {Pisum
sativum}, partial (79%)
Length = 655
Score = 28.9 bits (63), Expect = 6.6
Identities = 20/59 (33%), Positives = 30/59 (49%)
Frame = +2
Query: 333 SIATHTKMLKTQISQVAQQVATSSQTPGVFPGQPETNPKAHVNAIFLGGSKLEETVAKA 391
S+A +L T S + AT+++ PGV+ G P T+ +A F GGS T+ A
Sbjct: 20 SLAMAGILLLTIASLASLFAATTARIPGVYSGGPWTS----AHATFYGGSDASGTMGGA 184
>TC77948 similar to PIR|C85360|C85360 hypothetical protein AT4g30780
[imported] - Arabidopsis thaliana, partial (28%)
Length = 1916
Score = 24.3 bits (51), Expect(2) = 8.6
Identities = 10/27 (37%), Positives = 16/27 (59%), Gaps = 1/27 (3%)
Frame = -2
Query: 107 WERFKEMLRLCPHHGLEK-WLIVHTFY 132
W + ++ L PHH ++K L +H FY
Sbjct: 1012 WRKMIDLQILIPHHAVQKSQLSLHFFY 932
Score = 22.3 bits (46), Expect(2) = 8.6
Identities = 14/43 (32%), Positives = 18/43 (41%)
Frame = -1
Query: 37 QEAVRLHLFPFSLRDRARAWFHSLEVGSITSWDDMRRAFLARF 79
Q A +L F S + AW E+G + SW A RF
Sbjct: 1151 QIAEKLIRFRSSAPETFDAWGFITELGEVISWKLFSTALPIRF 1023
>BF637022 similar to GP|7757910|gb| putative translation initiation factor 2B
beta subunit {Nicotiana tabacum}, partial (38%)
Length = 542
Score = 28.5 bits (62), Expect = 8.6
Identities = 11/29 (37%), Positives = 19/29 (64%)
Frame = -3
Query: 211 FEKLNVNAVTPSSASPPCEICGISGHIGV 239
FE L+ + + PS + P ++CG++ IGV
Sbjct: 474 FELLSTDLLFPSESPPEEDVCGMAAEIGV 388
>BQ135535
Length = 765
Score = 28.5 bits (62), Expect = 8.6
Identities = 15/34 (44%), Positives = 19/34 (55%)
Frame = +2
Query: 118 PHHGLEKWLIVHTFYNGLSYTTKMSVDAAACGAL 151
PHHGL+K L + YN YTT+ D+ C L
Sbjct: 635 PHHGLKKHLPSLSAYN---YTTRFPHDSPRCTTL 727
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.319 0.135 0.394
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 19,477,482
Number of Sequences: 36976
Number of extensions: 257281
Number of successful extensions: 1273
Number of sequences better than 10.0: 28
Number of HSP's better than 10.0 without gapping: 1261
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1272
length of query: 662
length of database: 9,014,727
effective HSP length: 102
effective length of query: 560
effective length of database: 5,243,175
effective search space: 2936178000
effective search space used: 2936178000
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 62 (28.5 bits)
Medicago: description of AC135416.15


