

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC135396.1 - phase: 0 /pseudo
(1436 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
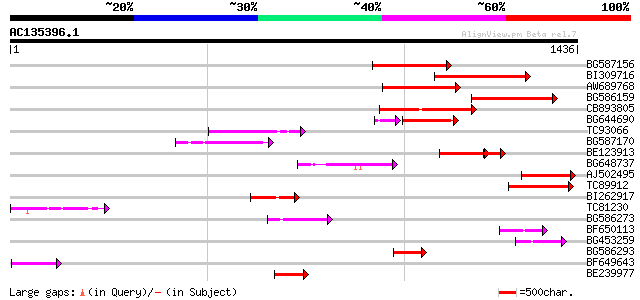
Score E
Sequences producing significant alignments: (bits) Value
BG587156 similar to PIR|G85055|G8 probable polyprotein [imported... 229 6e-60
BI309716 weakly similar to PIR|G96722|G9 hypothetical protein F2... 221 1e-57
AW689768 weakly similar to GP|10177485|d polyprotein {Arabidopsi... 206 4e-53
BG586159 weakly similar to PIR|T47841|T4 hypothetical protein T2... 185 1e-46
CB893805 similar to GP|10177935|d copia-type polyprotein {Arabid... 184 2e-46
BG644690 weakly similar to GP|18542179|gb putative pol protein {... 141 9e-41
TC93066 weakly similar to GP|19920130|gb|AAM08562.1 Putative ret... 152 7e-37
BG587170 similar to PIR|F86470|F8 probable retroelement polyprot... 149 1e-35
BE123913 weakly similar to GP|22093573|d polyprotein {Oryza sati... 140 4e-33
BG648737 weakly similar to GP|21433|emb|CA ORF3 {Solanum tuberos... 126 5e-29
AJ502495 weakly similar to GP|18071369|g putative gag-pol polypr... 126 7e-29
TC89912 weakly similar to PIR|B84512|B84512 probable retroelemen... 112 8e-25
BI262917 weakly similar to GP|19920130|g Putative retroelement {... 112 1e-24
TC81230 100 3e-21
BG586273 weakly similar to PIR|F86470|F8 probable retroelement p... 100 5e-21
BF650113 weakly similar to GP|4753889|emb| Tpv2-1c {Phaseolus vu... 96 1e-19
BG453259 homologue to GP|21434|emb|CA ORF4 {Solanum tuberosum}, ... 87 4e-17
BG586293 weakly similar to PIR|E84473|E84 probable retroelement ... 84 5e-16
BF649643 similar to PIR|H86486|H86 protein Ty1/copia-element pol... 82 1e-15
BE239977 weakly similar to GP|23237899|db polyprotein-like {Oryz... 78 2e-14
>BG587156 similar to PIR|G85055|G8 probable polyprotein [imported] -
Arabidopsis thaliana, partial (17%)
Length = 618
Score = 229 bits (584), Expect = 6e-60
Identities = 108/200 (54%), Positives = 145/200 (72%)
Frame = -1
Query: 918 YDKLSHSHKAYVSRISNLFVPRTIQEALGDPNWKLAVKEEMNALNKNNTWCITDLPYDKK 977
+ + +H A++ ++ +PR+ +EA+ D WK +V E A+ KN+TW ++LP KK
Sbjct: 618 FSQYPEAHCAFMVNLNENHIPRSYEEAMEDKEWKESVGAEAGAMIKNDTWYESELPKGKK 439
Query: 978 AVGCKWVFTVKCKADGSVERYKARLVAKGFTQTHGIDYQETFAPVAKINSIRILLSLAVN 1037
AV +W+FT+K KADGS+ER K RLVA+GFT T+G DY ETFAPVAK+++IRI+LSLAVN
Sbjct: 438 AVSSRWIFTIKYKADGSIERKKTRLVARGFTLTYGEDYIETFAPVAKLHTIRIVLSLAVN 259
Query: 1038 FNWTLHQYDVKNAFLNGELHEEVYMRLPPGFEDKLGRGKVCRLKKSLYGLKQSPRAWFER 1097
W L Q DVKNAFL GEL +EVYM PPG E + RG V RLKK++YGLKQSPRAW+ +
Sbjct: 258 LGWGLWQMDVKNAFLQGELEDEVYMYPPPGLEHLVKRGNVLRLKKAIYGLKQSPRAWYNK 79
Query: 1098 FGSVVKGHGFTQSQADHTMF 1117
+ + G GF +S+ DHT+F
Sbjct: 78 LSTTLNGRGFRKSELDHTLF 19
>BI309716 weakly similar to PIR|G96722|G9 hypothetical protein F20P5.25
[imported] - Arabidopsis thaliana, partial (10%)
Length = 744
Score = 221 bits (564), Expect = 1e-57
Identities = 109/242 (45%), Positives = 158/242 (65%)
Frame = +2
Query: 1076 KVCRLKKSLYGLKQSPRAWFERFGSVVKGHGFTQSQADHTMFFKHSREGKIAILIVYVDD 1135
KVC L+KS+YGLKQ+ R W+ + + G+ QS +D ++F K ++ L+VYVDD
Sbjct: 17 KVCELQKSIYGLKQASRQWYSKLSESLISFGYLQSSSDFSLFTKF-KDSSFTTLLVYVDD 193
Query: 1136 IIMTGDDIGEISDLKRRLEAEFDIKDLGKLKYFLGMEFARSKEGIFLNQRKYILDLLTET 1195
I++ G+DI EI +K L F IKDLG L+YFLG+E ARSK+GI LNQRKY L+LL ++
Sbjct: 194 IVLAGNDISEIQHVKCFLIDRFKIKDLGSLRYFLGLEVARSKQGILLNQRKYTLELLEDS 373
Query: 1196 GMTGCKAAETPMDPNVKLKSVAEDEIIDRERYQRLAGRLIYLSHTRPDIAFAVSVISQFM 1255
G K+ TP D ++KL + D +Y+RL G+LIYL+ TRPDI+FAV +SQF+
Sbjct: 374 GNLAVKSTLTPYDISLKLHNSDSPLYNDETQYRRLIGKLIYLTTTRPDISFAVQQLSQFV 553
Query: 1256 HAPGPAHFEAVFRILRYLKGTPGKGLMFRNRGHIQVEAYTDADWAGNINDRRSTSGYCTF 1315
P H++A R+L+YLK P KGL + ++++ ++ D+DWA R+S +GY F
Sbjct: 554 SKPQQVHYQAAIRVLQYLKTAPAKGLFYSATSNLKLSSFADSDWATCPTTRKSVTGYWVF 733
Query: 1316 VG 1317
+G
Sbjct: 734 LG 739
>AW689768 weakly similar to GP|10177485|d polyprotein {Arabidopsis thaliana},
partial (9%)
Length = 675
Score = 206 bits (525), Expect = 4e-53
Identities = 102/196 (52%), Positives = 133/196 (67%)
Frame = +1
Query: 945 LGDPNWKLAVKEEMNALNKNNTWCITDLPYDKKAVGCKWVFTVKCKADGSVERYKARLVA 1004
L DP W A+K E AL N TW + LP KKA+GCKWV+ VK DGSV ++KARLVA
Sbjct: 82 LSDPRWLQAMKTEYKALIDNKTWDLVPLPPHKKAIGCKWVYRVKENPDGSVNKFKARLVA 261
Query: 1005 KGFTQTHGIDYQETFAPVAKINSIRILLSLAVNFNWTLHQYDVKNAFLNGELHEEVYMRL 1064
KGF+QT G DY ETF+PV K +IR++L++A+ + W + Q D+ NAFLNG L EEVYM
Sbjct: 262 KGFSQTLGCDYTETFSPVIKPVTIRLILTIAITYKWEIQQIDINNAFLNGFLQEEVYMSQ 441
Query: 1065 PPGFEDKLGRGKVCRLKKSLYGLKQSPRAWFERFGSVVKGHGFTQSQADHTMFFKHSREG 1124
P GFE + VC+L KSLYGLKQ+PRAW+E S GFT+S+ D ++ +++ G
Sbjct: 442 PQGFE-AANKSLVCKLNKSLYGLKQAPRAWYEXLTSAQIQFGFTKSRCDPSLLI-YNQNG 615
Query: 1125 KIAILIVYVDDIIMTG 1140
L +YVDDI++TG
Sbjct: 616 ACIYLXIYVDDILITG 663
>BG586159 weakly similar to PIR|T47841|T4 hypothetical protein T2O9.150 -
Arabidopsis thaliana, partial (11%)
Length = 732
Score = 185 bits (469), Expect = 1e-46
Identities = 87/217 (40%), Positives = 139/217 (63%)
Frame = +1
Query: 1171 MEFARSKEGIFLNQRKYILDLLTETGMTGCKAAETPMDPNVKLKSVAEDEIIDRERYQRL 1230
+E +++EGI++ QRKY+ DLL GM + P+ P KL +D +Y+++
Sbjct: 1 VEVIQNEEGIYICQRKYVTDLLERFGMEKSNLSRNPIAPRCKLIKDENGVKVDATKYKQI 180
Query: 1231 AGRLIYLSHTRPDIAFAVSVISQFMHAPGPAHFEAVFRILRYLKGTPGKGLMFRNRGHIQ 1290
G L+YL+ TRPD+ + +S+IS+FM+ P H AV R+LRYL GT G+M++ G +
Sbjct: 181 VGCLMYLAATRPDLMYVLSLISRFMNCPTELHMHAVKRVLRYLNGTINLGIMYKRNGSEK 360
Query: 1291 VEAYTDADWAGNINDRRSTSGYCTFVGGNLVTWRSKKQNVVARSSAEAEFRSVAHGFCEV 1350
+EAYTD+D+AG+++DR+STSGY + V+W SKKQ VV S+ +AEF + A C+
Sbjct: 361 LEAYTDSDYAGDLDDRKSTSGYVFMLSSGAVSWSSKKQPVVTLSTTKAEFIAAAFCACQS 540
Query: 1351 LWIKKFMQELKIAGPTPMKVYCDNKAAISIAHNPVLH 1387
+W+++ +++L + +YCDN + I ++ NPVLH
Sbjct: 541 VWMRRVLEKLGYTQSGSITMYCDNNSTIKLSKNPVLH 651
>CB893805 similar to GP|10177935|d copia-type polyprotein {Arabidopsis
thaliana}, partial (14%)
Length = 778
Score = 184 bits (468), Expect = 2e-46
Identities = 97/249 (38%), Positives = 152/249 (60%), Gaps = 4/249 (1%)
Frame = +3
Query: 938 PRTIQEALGDPNWKLAVKEEMNALNKNNTWCITDLPYDKKAVGCKWVFTVKCKADGSVER 997
P T +EA+ W+ ++ EM A +NNTW +TDL K +G KW+F K +G +E+
Sbjct: 42 PTTFEEAVKSEKWRASMNNEMEATERNNTWELTDLRSGAKTIGLKWIFKTKLNENGEIEK 221
Query: 998 YKARLVAKGFTQTHGIDYQETFAPVAKINSIRILLSLAVNFNWTLHQYDVKNAFLNGELH 1057
YKARLVAKG++Q +G+DY E FAPVA+ ++IR++++LA + D + H
Sbjct: 222 YKARLVAKGYSQQYGVDYTEVFAPVARWDTIRMVIALAAQI-----KRDGVCIS*M*KAH 386
Query: 1058 E--EVYMRLPPGFEDKLGRGKVC--RLKKSLYGLKQSPRAWFERFGSVVKGHGFTQSQAD 1113
E MR ++ +V R+K++LYGLKQ+PRAW+ R + GF + +
Sbjct: 387 SCMEN*MRKFLLINHRVM*RRVIS*RVKRALYGLKQAPRAWYSRIEAYFTKEGFEKCPYE 566
Query: 1114 HTMFFKHSREGKIAILIVYVDDIIMTGDDIGEISDLKRRLEAEFDIKDLGKLKYFLGMEF 1173
HT+F K S GKI I+ +YVDD+I G+D + K+ ++ EF++ DLGK+ YFLG+E
Sbjct: 567 HTLFVKLSEGGKILIISLYVDDLIFIGNDENMFEEFKKSMKKEFNMSDLGKMHYFLGVEV 746
Query: 1174 ARSKEGIFL 1182
++++GI++
Sbjct: 747 TQNEKGIYI 773
>BG644690 weakly similar to GP|18542179|gb putative pol protein {Zea mays},
partial (22%)
Length = 629
Score = 141 bits (356), Expect(2) = 9e-41
Identities = 69/142 (48%), Positives = 100/142 (69%)
Frame = -2
Query: 995 VERYKARLVAKGFTQTHGIDYQETFAPVAKINSIRILLSLAVNFNWTLHQYDVKNAFLNG 1054
+ R K++LV +G+ Q GIDY E F+PVA++ +IRIL++ A + L+Q DVK+AF+NG
Sbjct: 424 ITRNKSKLVVQGYNQKEGIDYDEAFSPVARMEAIRILIAFAAFMGFKLYQMDVKSAFING 245
Query: 1055 ELHEEVYMRLPPGFEDKLGRGKVCRLKKSLYGLKQSPRAWFERFGSVVKGHGFTQSQADH 1114
+L EEV+++ PPGFED V RL K+LYGLKQ+PRAW+ER + +GF + + D+
Sbjct: 244 DLKEEVFVKQPPGFEDAEVPNHVFRLNKTLYGLKQAPRAWYERLSKFLLKNGFKRGKIDN 65
Query: 1115 TMFFKHSREGKIAILIVYVDDI 1136
T+F RE ++ I+ VYVDDI
Sbjct: 64 TLFLL-KRE*ELLIIQVYVDDI 2
Score = 45.4 bits (106), Expect(2) = 9e-41
Identities = 20/65 (30%), Positives = 36/65 (54%)
Frame = -3
Query: 924 SHKAYVSRISNLFVPRTIQEALGDPNWKLAVKEEMNALNKNNTWCITDLPYDKKAVGCKW 983
S A++S I P+ ++EAL D +W +++EE++ ++ W + P K +G +W
Sbjct: 615 SFSAFISSIE----PKNVKEALRDADWINSMQEELHQFERSKVWYLVPRPEGKTVIGTRW 448
Query: 984 VFTVK 988
VF K
Sbjct: 447 VFRNK 433
>TC93066 weakly similar to GP|19920130|gb|AAM08562.1 Putative retroelement
{Oryza sativa} [Oryza sativa (japonica cultivar-group)],
partial (10%)
Length = 823
Score = 152 bits (385), Expect = 7e-37
Identities = 86/250 (34%), Positives = 133/250 (52%), Gaps = 4/250 (1%)
Frame = +1
Query: 504 RVSFNSRGYSASKPFYLIHSDVWGPSKIKTLSRKKWFVTFIDDHTRVCWVYLMEKKSEVE 563
+VSF++ + IHSD+WGPSK+ + +++ +T IDD R WVY + K+E
Sbjct: 55 KVSFSTATHRTKGILDYIHSDLWGPSKVTSYGGRRYMMTIIDDFPRKVWVYFLRYKNETF 234
Query: 564 QRFQDFFNMIKNQFHTTIGILRSDNGTEYFNKYLSTFLVTNGIIHQSTCRDTPQQNGIAE 623
F+ + +++ Q + L +DN E+ + + F +GI T PQQNG+AE
Sbjct: 235 PTFKKWRILVETQTGKNVKKLITDN*LEFCSSDFNEFCTNHGIARHKTIPRNPQQNGVAE 414
Query: 624 RKNRHLLEVTRAIMFSMNV--PKYLWGNALLTACHLINRMPSRVLQYETPVQVLQNNFPT 681
R R LLE R ++ + + + LW A TACHL+NR P L ++ P + N
Sbjct: 415 RMIRTLLERARCMLSNAGL*N*RDLWVEAASTACHLVNRSPHSALDFKVPEDIWSGN--- 585
Query: 682 SRIITNIPLKVFGCLCYVYIPNIFRSKLDPKAEKCVFLGYASNKKGYK--CFNPVTKKFF 739
++ L++FGC Y + + KL P+A +C+FL YAS KGY+ C +P ++K
Sbjct: 586 --LVDYSNLRIFGCPAYALVND---GKLAPRAGECIFLSYASESKGYRLWCSDPKSQKLI 750
Query: 740 ESMDVHFVED 749
S DV F ED
Sbjct: 751 LSRDVTFNED 780
>BG587170 similar to PIR|F86470|F8 probable retroelement polyprotein
[imported] - Arabidopsis thaliana, partial (13%)
Length = 718
Score = 149 bits (375), Expect = 1e-35
Identities = 92/249 (36%), Positives = 133/249 (52%), Gaps = 1/249 (0%)
Frame = -3
Query: 420 SGKTIGTAREINGLYYFDET-PLGNTAMFGSSRTSPPLVSNKIMLWHKRLGHPSFSYLKH 478
S + IG LY ++ P+ N +S +S NK LWH RLGHP L
Sbjct: 704 SSQLIGEGVTKGDLYMLEKLDPVSNYKCSFTSSSS----LNKDALWHARLGHPHGRALNL 537
Query: 479 LFPEFSKEISSSQFHCDACHLAKDHRVSFNSRGYSASKPFYLIHSDVWGPSKIKTLSRKK 538
+ P E + C+AC L K + F F LI++D+W + K
Sbjct: 536 MLPGVVFENKN----CEACILGKHCKNVFPRTSTVYENCFDLIYTDLWTAPSLSR-DNHK 372
Query: 539 WFVTFIDDHTRVCWVYLMEKKSEVEQRFQDFFNMIKNQFHTTIGILRSDNGTEYFNKYLS 598
+FVTFID+ ++ W+ L+ K V F++F + N +H I ILRSDNG EY +
Sbjct: 371 YFVTFIDEKSKYTWLTLIPSKDRVIDAFKNFQAYVTNHYHAKIKILRSDNGGEYTSYAFK 192
Query: 599 TFLVTNGIIHQSTCRDTPQQNGIAERKNRHLLEVTRAIMFSMNVPKYLWGNALLTACHLI 658
+ L +GI+HQ++C TPQQNG+A+RKN+HL+EV R++MF NV TAC+LI
Sbjct: 191 SHLDHHGILHQTSCPYTPQQNGVAKRKNKHLMEVARSLMFQANVS---------TACYLI 39
Query: 659 NRMPSRVLQ 667
N +P++VL+
Sbjct: 38 NWIPTKVLR 12
>BE123913 weakly similar to GP|22093573|d polyprotein {Oryza sativa (japonica
cultivar-group)}, partial (8%)
Length = 503
Score = 140 bits (353), Expect = 4e-33
Identities = 74/127 (58%), Positives = 85/127 (66%)
Frame = +1
Query: 1089 QSPRAWFERFGSVVKGHGFTQSQADHTMFFKHSREGKIAILIVYVDDIIMTGDDIGEISD 1148
QSPR WF+RF VVK G+ Q Q DH MF KHS K AILIVYVDDI +TGD I
Sbjct: 1 QSPRDWFDRFT*VVKKFGYIQCQTDHAMFIKHSSTVKKAILIVYVDDIFLTGDHGK*IKR 180
Query: 1149 LKRRLEAEFDIKDLGKLKYFLGMEFARSKEGIFLNQRKYILDLLTETGMTGCKAAETPMD 1208
LK L EF+IKDLG LKYFLGME AR K+G ++QRKY+LDLL ET M GCK P
Sbjct: 181 LKNLLAEEFEIKDLGNLKYFLGMEVARWKKGSSISQRKYVLDLLKETRMIGCKTIRDPYG 360
Query: 1209 PNVKLKS 1215
N + ++
Sbjct: 361 CNCEARN 381
Score = 72.0 bits (175), Expect = 2e-12
Identities = 33/55 (60%), Positives = 42/55 (76%)
Frame = +2
Query: 1201 KAAETPMDPNVKLKSVAEDEIIDRERYQRLAGRLIYLSHTRPDIAFAVSVISQFM 1255
K +ETPMD VKL ++ ++D+ RYQRL G+LIYLSHTRPDI+F V +SQFM
Sbjct: 338 KPSETPMDATVKLGTLDNGTLVDKGRYQRLVGKLIYLSHTRPDISFVVCTMSQFM 502
>BG648737 weakly similar to GP|21433|emb|CA ORF3 {Solanum tuberosum}, partial
(10%)
Length = 804
Score = 126 bits (317), Expect = 5e-29
Identities = 88/294 (29%), Positives = 132/294 (43%), Gaps = 39/294 (13%)
Frame = +3
Query: 728 YKCFNPVTKKFFESMDVHFVEDQPFFRENSLQGESQSSNEEDNFWEILPVLDDIVTNDHL 787
YKC++P+TKKF+ SMDV F EDQPF+ + +QGES + EE W+I
Sbjct: 3 YKCYSPITKKFYNSMDVTFFEDQPFYTKIGIQGESFT--EEYQLWDI------------- 137
Query: 788 DAKITEPRNPNLELLEHMVFETGGEEGETLIKNPELRVYVRKKF-----HKDGTNPIVFP 842
++LE + E + NP + ++ K + P +FP
Sbjct: 138 ------------KVLESIHGENSDVSPQVNQTNPIVSYHLSSKIPPPVSDQPNIQPSIFP 281
Query: 843 AEVQSDSPSEGPTDNPSSSSSGNSSHSSNDLP-------------------------DLS 877
Q D + T P S S S++ P D S
Sbjct: 282 MSNQFDLSNSQSTIMPLSEQQPPSQRSTSTQPAEICDFRVFTTRYKKQEGKRMQQIQDCS 461
Query: 878 FPDINLPIA------VRKNIPDLNIP---IAERKRTRTCTKHPMSNYLSYDKLSHSHKAY 928
+ N I + + IP + I+ RK R+CT HP+ ++SYDKLS H A+
Sbjct: 462 DSNSNFEIENTQDNEICEPIPTVEFDDRLISLRKDKRSCTSHPICKFISYDKLSPHHLAF 641
Query: 929 VSRISNLFVPRTIQEALGDPNWKLAVKEEMNALNKNNTWCITDLPYDKKAVGCK 982
VS + + +P ++ E+L +P WK A+ E++AL KN TW ++DLP K +GCK
Sbjct: 642 VSNLDKIQIPNSVHESLTNP*WKKAILGEIHALEKNETWELSDLPSGKHPMGCK 803
>AJ502495 weakly similar to GP|18071369|g putative gag-pol polyprotein {Oryza
sativa}, partial (9%)
Length = 542
Score = 126 bits (316), Expect = 7e-29
Identities = 56/137 (40%), Positives = 89/137 (64%)
Frame = +2
Query: 1297 ADWAGNINDRRSTSGYCTFVGGNLVTWRSKKQNVVARSSAEAEFRSVAHGFCEVLWIKKF 1356
+DWAG+ R+STSGY +G ++W SKKQ VVA S+AEAE+ + + +W+++
Sbjct: 2 SDWAGDTETRKSTSGYAFHLGTGAISWSSKKQPVVAFSTAEAEYIASTSCATQTVWLRRI 181
Query: 1357 MQELKIAGPTPMKVYCDNKAAISIAHNPVLHDRTKHVEVDKHFIKEKIDSGEICMSYIPT 1416
++ + TP K+YCDNK+AI+++ NPV H R+KH+++ H I+E I E+ + Y PT
Sbjct: 182 LEVMHHEQNTPTKIYCDNKSAIALSKNPVFHGRSKHIDIQFHKIRELIAEKEVVIEYCPT 361
Query: 1417 KSQVADVLTKSLPKRQF 1433
+ ++AD+ TK L F
Sbjct: 362 EEKIADIFTKPLKIESF 412
>TC89912 weakly similar to PIR|B84512|B84512 probable retroelement pol
polyprotein [imported] - Arabidopsis thaliana, partial
(10%)
Length = 814
Score = 112 bits (281), Expect = 8e-25
Identities = 55/165 (33%), Positives = 103/165 (62%), Gaps = 2/165 (1%)
Frame = +1
Query: 1264 EAVFRILRYLKGTPGKGLMFRNRGHIQ--VEAYTDADWAGNINDRRSTSGYCTFVGGNLV 1321
+A+ +L+YL + L + + +E Y DAD+AGN++ R+S SG+ + G +
Sbjct: 1 QALKWVLKYLNESLKSSLKYTKAAQEEDALEGYVDADYAGNVDTRKSLSGFVFTLYGTTI 180
Query: 1322 TWRSKKQNVVARSSAEAEFRSVAHGFCEVLWIKKFMQELKIAGPTPMKVYCDNKAAISIA 1381
+W++ +Q+VV S+ +AE+ + G + +W+K + EL I +K++CD+++AI +A
Sbjct: 181 SWKANQQSVVTLSTTQAEYIAFVEGVKDAIWLKGMIGELGIT-QEYVKIHCDSQSAIHLA 357
Query: 1382 HNPVLHDRTKHVEVDKHFIKEKIDSGEICMSYIPTKSQVADVLTK 1426
++ V H+RTKH+++ HFI++ I+S EI + + ++ ADV TK
Sbjct: 358 NHQVYHERTKHIDIRLHFIRDMIESKEIVVEKMASEENPADVFTK 492
>BI262917 weakly similar to GP|19920130|g Putative retroelement {Oryza
sativa} [Oryza sativa (japonica cultivar-group)],
partial (8%)
Length = 426
Score = 112 bits (280), Expect = 1e-24
Identities = 57/124 (45%), Positives = 79/124 (62%)
Frame = +1
Query: 610 STCRDTPQQNGIAERKNRHLLEVTRAIMFSMNVPKYLWGNALLTACHLINRMPSRVLQYE 669
S TPQQNG+AER NR LLE TRA++ + + K W A+ TAC++INR PS V+ +
Sbjct: 70 SXVAHTPQQNGVAERMNRTLLERTRAMLKTAGMAKSFWAEAVKTACYVINRSPSTVIDLK 249
Query: 670 TPVQVLQNNFPTSRIITNIPLKVFGCLCYVYIPNIFRSKLDPKAEKCVFLGYASNKKGYK 729
TP+++ + + + L VFGC YV + R+KLDPK+ KC+FLGYA N KGY
Sbjct: 250 TPMEMWK-----GKPVDYSSLHVFGCPVYVMYNSQERTKLDPKSRKCIFLGYADNVKGYX 414
Query: 730 CFNP 733
++P
Sbjct: 415 LWDP 426
>TC81230
Length = 958
Score = 100 bits (250), Expect = 3e-21
Identities = 73/260 (28%), Positives = 122/260 (46%), Gaps = 11/260 (4%)
Frame = +1
Query: 3 TIRLNGANYFRWSQSVQLYIRGRGKIGYITGDKKQPDKVGAD--------YDTWDAENSM 54
+I LNG+NY W++S+ +++GR Y+TGDKK P K D + WD++N
Sbjct: 229 SIILNGSNYNHWAESMCGFLKGRRLWRYVTGDKKCPTKGKDDTADAFADKLEEWDSKNHQ 408
Query: 55 VMTWLVNSLTEEIGANYLCYATAKDLWENVSQMY--SDLGNQSQIYELTLQLGEIR-QAE 111
++TW N+ I + + AK++W+++ Q Y SDL +Q Y+L L ++ Q+
Sbjct: 409 IITWFRNTSIPSIHMQFGRFENAKEVWDHLKQRYTISDLSHQ---YQLLKDLSNLKQQSG 579
Query: 112 DSVTKYFGCLKRIWQDLDLFNEYEWKSPEDCKHYQKMVDVSRVFKFLAGLNVEFDEVRGR 171
V ++ ++ IW L E K D K Y+ + R+ +FL L E++ VR
Sbjct: 580 QPVYEFLAQMEVIWNQLTSC-EPSLKDATDMKTYETHRNRVRLIQFLMALTDEYEPVRAS 756
Query: 172 ILGRNPIPPIGEVFAEVRREESRRQVMLGKKKAAVPPPVEGSALAVPQVNHKSFRNPRGG 231
L +NP+P + ++ EE+R Q++ PP A AV K
Sbjct: 757 SLHQNPLPTLENALPCLKSEETRLQLV---------PPKADLAFAVTNNATKP------- 888
Query: 232 DKTHLVCDYCGRNRHTQETC 251
C +C ++ H+ C
Sbjct: 889 ------CRHCQKSGHSFSDC 930
>BG586273 weakly similar to PIR|F86470|F8 probable retroelement polyprotein
[imported] - Arabidopsis thaliana, partial (7%)
Length = 705
Score = 100 bits (248), Expect = 5e-21
Identities = 53/168 (31%), Positives = 93/168 (54%), Gaps = 5/168 (2%)
Frame = -2
Query: 654 ACHLINRMPSRVLQYETPVQVLQNNFPTSRIITNIPLKVFGCLCYVYIPNIFRSKLDPKA 713
AC+LINR+P+RVL+ + P +VL P+ ++VFGCLCYV +P R+KL+ ++
Sbjct: 704 ACYLINRIPTRVLKDQAPFEVLNQRKPSLTY-----MRVFGCLCYVLVPGELRNKLEARS 540
Query: 714 EKCVFLGYASNKKGYKCFNPVTKKFFESMDVHFVEDQPFFRENSLQGESQSSNEEDNFWE 773
K +F+GY++ +KGYKC++P ++ S DV F+E++ ++ E + + ++++
Sbjct: 539 RKAMFIGYSTTQKGYKCYDPEARRVLVSRDVKFIEERGYYEEKNQEDLRDLTSDKAGVLR 360
Query: 774 ILPVLDDIVTNDHLDAKITEPRNPNLE-----LLEHMVFETGGEEGET 816
++ I N + +P + E H+ E GG E ET
Sbjct: 359 VILEGLGIKMNQDQSTRSRQPEESSNEPRRAAQTPHLDHE-GGSEPET 219
>BF650113 weakly similar to GP|4753889|emb| Tpv2-1c {Phaseolus vulgaris},
partial (13%)
Length = 494
Score = 95.9 bits (237), Expect = 1e-19
Identities = 54/124 (43%), Positives = 73/124 (58%), Gaps = 3/124 (2%)
Frame = +1
Query: 1241 RPDIAFAVSVISQFMHAPGPAHFEAVFRILRYLKGTPGKGLMFRNRGHIQVE---AYTDA 1297
RPDI ++VSVIS+FMH P H A RILRY++GT GL+F +V Y+D+
Sbjct: 121 RPDICYSVSVISKFMHDPRKPHLIAANRILRYVRGTMEYGLLFPYGAKSEVYELICYSDS 300
Query: 1298 DWAGNINDRRSTSGYCTFVGGNLVTWRSKKQNVVARSSAEAEFRSVAHGFCEVLWIKKFM 1357
DW G DRRSTSGY ++W +KKQ + A SS EAE+ + + LW+ +
Sbjct: 301 DWCG---DRRSTSGYVFKFNDAAISWCTKKQPITALSSYEAEYIAGTFATFQALWLDSVI 471
Query: 1358 QELK 1361
+ELK
Sbjct: 472 KELK 483
>BG453259 homologue to GP|21434|emb|CA ORF4 {Solanum tuberosum}, partial (5%)
Length = 657
Score = 87.4 bits (215), Expect = 4e-17
Identities = 55/131 (41%), Positives = 75/131 (56%), Gaps = 2/131 (1%)
Frame = -2
Query: 1281 LMFRNRGHIQVEAYTDADWAGNINDRRSTSGYCTFVGGNLVTWRSKKQNVVARSSAEAEF 1340
++F+ + +E Y + AG I DR STSGY F+GGN+V KQNVVAR F
Sbjct: 530 IVFKRE*KLSMEVYXNTVCAGWIVDRGSTSGY*MFLGGNMV---E*KQNVVAR*VQRHNF 360
Query: 1341 RSVAHGFCEVL--WIKKFMQELKIAGPTPMKVYCDNKAAISIAHNPVLHDRTKHVEVDKH 1398
+ G V+ +K + +L I PM ++ +N IAHNPV H RTKH+E+D+H
Sbjct: 359 ELCSQGL*RVMDEELKIKLDDLIINYKDPMTLF*NNNFVSRIAHNPVQHYRTKHIEIDQH 180
Query: 1399 FIKEKIDSGEI 1409
FI EK+ SG I
Sbjct: 179 FIIEKLYSGLI 147
>BG586293 weakly similar to PIR|E84473|E84 probable retroelement pol
polyprotein [imported] - Arabidopsis thaliana, partial
(7%)
Length = 763
Score = 83.6 bits (205), Expect = 5e-16
Identities = 39/82 (47%), Positives = 55/82 (66%)
Frame = +2
Query: 973 PYDKKAVGCKWVFTVKCKADGSVERYKARLVAKGFTQTHGIDYQETFAPVAKINSIRILL 1032
P K +G +W++ +K DG++ +YKARLVAKG+ + GID+ E FAPV +I +I +LL
Sbjct: 77 PTGVKPIGLRWIYKIKRNEDGTLIKYKARLVAKGYVKQQGIDFDEVFAPVVRIETI*LLL 256
Query: 1033 SLAVNFNWTLHQYDVKNAFLNG 1054
+LA +H DVK AFLNG
Sbjct: 257 ALAATNGC*IHHIDVKIAFLNG 322
>BF649643 similar to PIR|H86486|H86 protein Ty1/copia-element polyprotein
[imported] - Arabidopsis thaliana, partial (2%)
Length = 608
Score = 82.0 bits (201), Expect = 1e-15
Identities = 40/129 (31%), Positives = 70/129 (54%), Gaps = 2/129 (1%)
Frame = +1
Query: 5 RLNGANYFRWSQSVQLYIRGRGKIGYITGDKKQPDKVG--ADYDTWDAENSMVMTWLVNS 62
+LNG NY WS+S+ + + K+G+I G K P +V A+Y W+ NSM+++WL +S
Sbjct: 172 KLNGTNYSSWSRSMVHALTAKNKVGFINGSIKTPSEVDQPAEYALWNRCNSMILSWLTHS 351
Query: 63 LTEEIGANYLCYATAKDLWENVSQMYSDLGNQSQIYELTLQLGEIRQAEDSVTKYFGCLK 122
+ ++ + TA +WE+ +S N IY++ L + Q S + YF +K
Sbjct: 352 VEPDLAKGVIHAKTACQVWEDFKDQFSQ-KNIPAIYQIQKSLASLSQGTVSASTYFTKIK 528
Query: 123 RIWQDLDLF 131
+W +L+ +
Sbjct: 529 GLWDELETY 555
>BE239977 weakly similar to GP|23237899|db polyprotein-like {Oryza sativa
(japonica cultivar-group)}, partial (2%)
Length = 514
Score = 78.2 bits (191), Expect = 2e-14
Identities = 34/85 (40%), Positives = 57/85 (67%)
Frame = -2
Query: 672 VQVLQNNFPTSRIITNIPLKVFGCLCYVYIPNIFRSKLDPKAEKCVFLGYASNKKGYKCF 731
++VL + +P N+ ++FGC +V+I + RSK D +A KCVF+ Y+S +KGY+C+
Sbjct: 513 IEVLPSFYPYVPTSNNLTPRIFGCTSFVHIHSDGRSKFDHRALKCVFIRYSSTQKGYRCY 334
Query: 732 NPVTKKFFESMDVHFVEDQPFFREN 756
+P ++K+F S DV F E + +F +N
Sbjct: 333 HPPSRKYFVSRDVTFHEQESYFGQN 259
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.319 0.136 0.412
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 47,862,060
Number of Sequences: 36976
Number of extensions: 727199
Number of successful extensions: 3974
Number of sequences better than 10.0: 86
Number of HSP's better than 10.0 without gapping: 3848
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 3938
length of query: 1436
length of database: 9,014,727
effective HSP length: 108
effective length of query: 1328
effective length of database: 5,021,319
effective search space: 6668311632
effective search space used: 6668311632
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 65 (29.6 bits)
Medicago: description of AC135396.1


