

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC135320.2 + phase: 0
(1662 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
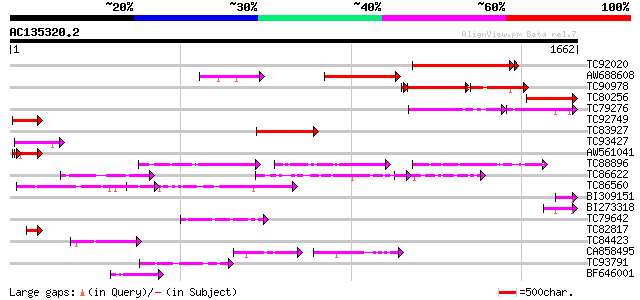
Score E
Sequences producing significant alignments: (bits) Value
TC92020 weakly similar to GP|9279697|dbj|BAB01254.1 centromere p... 591 e-170
AW688608 428 e-120
TC90978 weakly similar to GP|9279697|dbj|BAB01254.1 centromere p... 184 3e-88
TC80256 similar to GP|4894628|gb|AAD32567.1| NT3 {Nicotiana taba... 286 6e-77
TC79276 similar to PIR|F86161|F86161 hypothetical protein AAD258... 132 8e-57
TC92749 similar to GP|13486675|dbj|BAB39912. contains EST AU0626... 147 4e-35
TC83927 weakly similar to GP|9279697|dbj|BAB01254.1 centromere p... 134 2e-31
TC93427 similar to GP|9280668|gb|AAF86537.1| F21B7.9 {Arabidopsi... 111 2e-24
AW561041 similar to GP|13486675|db contains EST AU062670(C30068)... 102 1e-21
TC88896 weakly similar to GP|6728968|gb|AAF26966.1| unknown prot... 74 6e-13
TC86622 weakly similar to GP|20197623|gb|AAM15156.1 putative myo... 69 2e-11
TC86560 similar to GP|8843737|dbj|BAA97285.1 myosin heavy chain-... 64 5e-10
BI309151 similar to GP|4894628|gb|A NT3 {Nicotiana tabacum}, par... 63 8e-10
BI273318 similar to GP|7269756|emb predicted protein of unknown ... 59 2e-08
TC79642 similar to PIR|T07397|T07397 kinesin heavy chain-like pr... 58 4e-08
TC82817 similar to GP|9294158|dbj|BAB02060.1 contains similarity... 54 4e-07
TC84423 similar to PIR|F84730|F84730 probable myosin heavy chain... 52 2e-06
CA858495 50 1e-05
TC93791 similar to GP|15810373|gb|AAL07074.1 unknown protein {Ar... 49 2e-05
BF646001 homologue to GP|20303561|gb| putative kinase {Oryza sat... 48 4e-05
>TC92020 weakly similar to GP|9279697|dbj|BAB01254.1 centromere protein
{Arabidopsis thaliana}, partial (4%)
Length = 934
Score = 591 bits (1523), Expect(2) = e-170
Identities = 299/299 (100%), Positives = 299/299 (100%)
Frame = +2
Query: 1181 TLQFEMKTLLHEVEQHRVREEALNLELLNKENEFKLWENEAAAFYHDLQMSSICLALLES 1240
TLQFEMKTLLHEVEQHRVREEALNLELLNKENEFKLWENEAAAFYHDLQMSSICLALLES
Sbjct: 2 TLQFEMKTLLHEVEQHRVREEALNLELLNKENEFKLWENEAAAFYHDLQMSSICLALLES 181
Query: 1241 KVSELTGVCKILDDESSAKSLENEHMREIISLLESEIGGLKEQLSAYVPLVSSLKEDFNS 1300
KVSELTGVCKILDDESSAKSLENEHMREIISLLESEIGGLKEQLSAYVPLVSSLKEDFNS
Sbjct: 182 KVSELTGVCKILDDESSAKSLENEHMREIISLLESEIGGLKEQLSAYVPLVSSLKEDFNS 361
Query: 1301 LEHISLLWTKRNSVVGNGAQKDVVIETCLRKHSHQSARENEIVLIPDGVSDLLTLQTRIR 1360
LEHISLLWTKRNSVVGNGAQKDVVIETCLRKHSHQSARENEIVLIPDGVSDLLTLQTRIR
Sbjct: 362 LEHISLLWTKRNSVVGNGAQKDVVIETCLRKHSHQSARENEIVLIPDGVSDLLTLQTRIR 541
Query: 1361 AVEKIMMEELKRRVKQKSLTTESTPYSSLEIATYPKVENRKKEIELVEENVFDRNSWRKK 1420
AVEKIMMEELKRRVKQKSLTTESTPYSSLEIATYPKVENRKKEIELVEENVFDRNSWRKK
Sbjct: 542 AVEKIMMEELKRRVKQKSLTTESTPYSSLEIATYPKVENRKKEIELVEENVFDRNSWRKK 721
Query: 1421 PKIRLLMKDIPLDRNVDDQNSKYLKREHRRTNDHVLELCENNEHEPLSAPTVDHAMICH 1479
PKIRLLMKDIPLDRNVDDQNSKYLKREHRRTNDHVLELCENNEHEPLSAPTVDHAMICH
Sbjct: 722 PKIRLLMKDIPLDRNVDDQNSKYLKREHRRTNDHVLELCENNEHEPLSAPTVDHAMICH 898
Score = 27.7 bits (60), Expect(2) = e-170
Identities = 11/11 (100%), Positives = 11/11 (100%)
Frame = +3
Query: 1480 RSDDSGRYLNY 1490
RSDDSGRYLNY
Sbjct: 900 RSDDSGRYLNY 932
>AW688608
Length = 678
Score = 428 bits (1101), Expect = e-120
Identities = 222/225 (98%), Positives = 222/225 (98%)
Frame = +3
Query: 922 VLLGSLQQHQSEGEKLKLEKKTVEQEFENMREQNVILQKDKVELLEENRQLRIEVVNGVE 981
VLLGSLQQHQSEGEKLKLEKKTVEQEFENMREQNVILQKDKVELLEENRQLRIEVVNGVE
Sbjct: 3 VLLGSLQQHQSEGEKLKLEKKTVEQEFENMREQNVILQKDKVELLEENRQLRIEVVNGVE 182
Query: 982 KENRSKSTLAALQAEMIELRQTNQVFQEENGKMLDEKNSLCRNVSDLKDAKSSAEDENSV 1041
KENRSKSTLAALQAEMIELRQTNQVFQEENGKMLDEKNSLCRNVSDLKDAKSSAEDENSV
Sbjct: 183 KENRSKSTLAALQAEMIELRQTNQVFQEENGKMLDEKNSLCRNVSDLKDAKSSAEDENSV 362
Query: 1042 MFHDVLALSNLNLVYEIFFTENMVEKRALCEHLGNLSHLNNDLNQEFGVLRKNFEVKEAE 1101
MFHDVLALSNLNLVYEIFFTENMVEKRALCEHLGNLSHLNNDLNQEFGVLR NFEVKEAE
Sbjct: 363 MFHDVLALSNLNLVYEIFFTENMVEKRALCEHLGNLSHLNNDLNQEFGVLRXNFEVKEAE 542
Query: 1102 NVYLNESIERMDKELLEMDKRLKAAETSNAEFSRHIEELKMEQEE 1146
NVYLNESIERMDKELLEMDKRLKAAETSNAEFS HIEEL MEQEE
Sbjct: 543 NVYLNESIERMDKELLEMDKRLKAAETSNAEFSXHIEELXMEQEE 677
Score = 46.2 bits (108), Expect = 1e-04
Identities = 48/215 (22%), Positives = 92/215 (42%), Gaps = 25/215 (11%)
Frame = +3
Query: 557 SVRDLQNENFMLKETCKKEHSEKEALREKS----KDMNEVLMENACMEFSLL-GLNDE-- 609
S++ Q+E LK K E E +RE++ KD E+L EN + ++ G+ E
Sbjct: 15 SLQQHQSEGEKLKLEKKTVEQEFENMREQNVILQKDKVELLEENRQLRIEVVNGVEKENR 194
Query: 610 ----LDGLRGTVKEIQQFCQVLQEEKSILADEKSTLLSQLQIITESMQKILENNTVL--- 662
L L+ + E++Q QV QEE + DEK++L + + ++ + N+V+
Sbjct: 195 SKSTLAALQAEMIELRQTNQVFQEENGKMLDEKNSLCRNVSDLKDAKSSAEDENSVMFHD 374
Query: 663 -----------EKSLSDAKIEFEGLRIKSGDLEDCCKLLNDEKNNLQNERSMLISQLEIV 711
E ++ +E L G+L LN E L+ + ++ +
Sbjct: 375 VLALSNLNLVYEIFFTENMVEKRALCEHLGNLSHLNNDLNQEFGVLRXNFEVKEAENVYL 554
Query: 712 EEKLSNLEKKVTNLEEKYADVEKDKESAVNQVEEL 746
E + ++K++ ++++ E +EEL
Sbjct: 555 NESIERMDKELLEMDKRLKAAETSNAEFSXHIEEL 659
>TC90978 weakly similar to GP|9279697|dbj|BAB01254.1 centromere protein
{Arabidopsis thaliana}, partial (5%)
Length = 1132
Score = 184 bits (467), Expect(3) = 3e-88
Identities = 104/183 (56%), Positives = 132/183 (71%)
Frame = +1
Query: 1167 NHKKEIEHLNEANETLQFEMKTLLHEVEQHRVREEALNLELLNKENEFKLWENEAAAFYH 1226
+ K ++ LNEAN ++ EM+ L EVEQ + REE L+ EL++K NEF+ WE EAA FY
Sbjct: 61 SQK*KLNFLNEANRSIMSEMRLLHQEVEQQKAREETLSSELMDKTNEFQPWEAEAATFYF 240
Query: 1227 DLQMSSICLALLESKVSELTGVCKILDDESSAKSLENEHMREIISLLESEIGGLKEQLSA 1286
DLQ+SSI ALLE+KV+ELTGVC L ES+AKS + E M E + LLESEIGGLK LSA
Sbjct: 241 DLQISSISEALLENKVNELTGVCTRLQGESAAKSSKIEKMTERVGLLESEIGGLKGHLSA 420
Query: 1287 YVPLVSSLKEDFNSLEHISLLWTKRNSVVGNGAQKDVVIETCLRKHSHQSARENEIVLIP 1346
YVP++SSLKEDF SLEH ++L + + S V N QKD VIETCL ++ + S EN ++L
Sbjct: 421 YVPVISSLKEDFASLEH-TILQSNKASAVCNQEQKDYVIETCLGENINPSVIENNLML-- 591
Query: 1347 DGV 1349
DGV
Sbjct: 592 DGV 600
Score = 146 bits (368), Expect(3) = 3e-88
Identities = 84/185 (45%), Positives = 121/185 (65%), Gaps = 13/185 (7%)
Frame = +2
Query: 1350 SDLLTLQTRIRAVEKIMMEELKRRVKQKSLTTESTPYSSLEIATYPKVENRKKEIELVEE 1409
SDL+ ++ RI VE+ M+EE++RRVK+++L++++ P + RK E +L +E
Sbjct: 602 SDLIGMKARIXVVERCMVEEIERRVKEENLSSKANPQK----------DYRKVEKQLKDE 751
Query: 1410 NVFDRNSWRKKPKIRLLMKDIPLDRNVDDQNSKYLKREHRRTNDHVLELCENNEHEPL-- 1467
N+FD N+WR K + LMKDIPLD+ D+ SK +R++R T+D +LEL E E +
Sbjct: 752 NMFDLNTWRTKSQNGSLMKDIPLDQISDNPASKNCRRKNRGTDDGMLELWETAEQDCFDD 931
Query: 1468 -----------SAPTVDHAMICHRSDDSGRYLNYSSELDIEKELGVDKLELSKSVKEKTE 1516
S PT D ++CH+SD+SGR LN SSEL+ EKELGVDKL LSKS+K++T+
Sbjct: 932 GLMVGEAMKRSSDPTED-VIMCHQSDNSGRCLNTSSELEAEKELGVDKLHLSKSIKDRTQ 1108
Query: 1517 DDKRR 1521
D KRR
Sbjct: 1109 DGKRR 1123
Score = 37.0 bits (84), Expect(3) = 3e-88
Identities = 16/20 (80%), Positives = 17/20 (85%)
Frame = +2
Query: 1150 IKENLDRQILEQSENCMNHK 1169
I ENL+RQILE SE CMNHK
Sbjct: 8 INENLERQILELSEGCMNHK 67
>TC80256 similar to GP|4894628|gb|AAD32567.1| NT3 {Nicotiana tabacum}, partial
(3%)
Length = 746
Score = 286 bits (731), Expect = 6e-77
Identities = 147/147 (100%), Positives = 147/147 (100%)
Frame = +1
Query: 1516 EDDKRRILERLSSDGQKLAILKMALQDLKKKTETKKKSKQGNDIEYETVKRHIEEVEEAV 1575
EDDKRRILERLSSDGQKLAILKMALQDLKKKTETKKKSKQGNDIEYETVKRHIEEVEEAV
Sbjct: 1 EDDKRRILERLSSDGQKLAILKMALQDLKKKTETKKKSKQGNDIEYETVKRHIEEVEEAV 180
Query: 1576 MQQVSINDQMAKNVEEGASSLDREIPRRGSEQIGKLQFEVQNIQYILLKLAEENNNKVKN 1635
MQQVSINDQMAKNVEEGASSLDREIPRRGSEQIGKLQFEVQNIQYILLKLAEENNNKVKN
Sbjct: 181 MQQVSINDQMAKNVEEGASSLDREIPRRGSEQIGKLQFEVQNIQYILLKLAEENNNKVKN 360
Query: 1636 RISRKTGILLRRKLRVCGCSGPSTNED 1662
RISRKTGILLRRKLRVCGCSGPSTNED
Sbjct: 361 RISRKTGILLRRKLRVCGCSGPSTNED 441
>TC79276 similar to PIR|F86161|F86161 hypothetical protein AAD25801.1
[imported] - Arabidopsis thaliana, partial (3%)
Length = 1748
Score = 132 bits (333), Expect(2) = 8e-57
Identities = 91/242 (37%), Positives = 128/242 (52%), Gaps = 36/242 (14%)
Frame = +3
Query: 1457 ELCENNEHEPLSA--PTVDHAMICHRSDDSGRYLNYSSELDIEKELGVDKLELSKSV-KE 1513
EL E ++ P A P DH R+ + S + +EKELGVDKLE+S+ + +
Sbjct: 864 ELWETSDKTPKMAAEPAEDHHQ--RRASKETYNKHPSGDSLVEKELGVDKLEISRRMSRP 1037
Query: 1514 KTEDDKRRILERLSSDGQKLAILKMALQDLKKKTETKKKSKQGNDIEYETVKRHIEEVEE 1573
+ E +K R+LERL SD QKL L++ +QDL KK ET +KS +G EY+TVK +E +E
Sbjct: 1038 REEGNKSRVLERLDSDSQKLTNLQITIQDLMKKVETIEKSTKGKSAEYDTVKEQLEASQE 1217
Query: 1574 AVMQQVSINDQMAKNVEEGA-SSLDR-----------------EIPRRGSEQIGKLQFEV 1615
VM+ N ++ KNVEEGA SS R E +RGSE+IG+LQ EV
Sbjct: 1218 TVMKLFDANRKLVKNVEEGALSSAGRASSESDEIGSVSRRRFSEQAQRGSEKIGQLQLEV 1397
Query: 1616 QNIQYILLKLAEENNNKVKNRIS---------------RKTGILLRRKLRVCGCSGPSTN 1660
Q +Q++LLKL + +K K +++ KT ++K C C P T
Sbjct: 1398 QRLQFLLLKLNDAKESKEKTKMADQSRRVRLRDYLYGGTKTNNQKKKKTPFCACVRPPTK 1577
Query: 1661 ED 1662
D
Sbjct: 1578 GD 1583
Score = 108 bits (269), Expect(2) = 8e-57
Identities = 82/288 (28%), Positives = 147/288 (50%), Gaps = 1/288 (0%)
Frame = +2
Query: 1170 KEIEHLNEANETLQFEMKTLLHEVEQHRVREEALNLELLNKENEFKLWENEAAAFYHDLQ 1229
+E+ L+E N+ L+ EMK L E+E+ ++RE+ L+ E+ NE + WE +AA + +LQ
Sbjct: 68 EELGCLSEVNQKLESEMKCLHQELEEIKLREKKLSYEVHEGINEIEQWETQAAVLFAELQ 247
Query: 1230 MSSICLALLESKVSELTGVCKILDDESSAKSLENEHMREIISLLESEIGGLKEQLSAYVP 1289
+S++ LL+ K EL C+ L+ + +K +E E ++E++S LE E G + +QL+AYVP
Sbjct: 248 VSAVNETLLQGKACELADTCEHLESINYSKDMEREQLKELVSKLEGENGKMCDQLAAYVP 427
Query: 1290 LVSSLKEDFNSLEHISLLWTKRNSVVGNGAQKDVVIETCLRKHSHQSARENEIVLIPDGV 1349
+S+L + SLE + L ++ K++V + + Q E + V PD +
Sbjct: 428 AISALNDCVTSLE-VQTLGHPKHHDYEKPEVKNLVNHQYI--ENGQQIDEYQSVTAPDPL 598
Query: 1350 SDLLTLQTRIRAVEKIMMEELKRRVKQKSLTTESTPYSSLEIATYPKVENRK-KEIELVE 1408
D LQ RI E+ VK + ++++ VE R+ +E + +E
Sbjct: 599 LDFQDLQRRI--------NEISMAVKNFNASSKA------------NVEMREIQEAKEIE 718
Query: 1409 ENVFDRNSWRKKPKIRLLMKDIPLDRNVDDQNSKYLKREHRRTNDHVL 1456
+ + +I +L KDI LD+ + + + ++DH+L
Sbjct: 719 QKMGSLRPDNPVTEIEVLPKDIMLDQISECSSYGVSRGGTLESDDHML 862
Score = 30.8 bits (68), Expect = 4.6
Identities = 47/231 (20%), Positives = 95/231 (40%), Gaps = 20/231 (8%)
Frame = +2
Query: 295 LNDQIERTELEVETLRKNLAEMNEERDSLSVLYHHCLEKISKMENE--ILHVQENAEQLK 352
L++ ++ E E++ L + L E+ LS H + +I + E + +L + +
Sbjct: 83 LSEVNQKLESEMKCLHQELEEIKLREKKLSYEVHEGINEIEQWETQAAVLFAELQVSAVN 262
Query: 353 NKI-EKEAEKLEISEKHRGMLEKSNQNLQLEAENLVQRIASKDHE--------------- 396
+ + +A +L + +H LE N + +E E L + ++ + E
Sbjct: 263 ETLLQGKACELADTCEH---LESINYSKDMEREQLKELVSKLEGENGKMCDQLAAYVPAI 433
Query: 397 --LLEKHTEIERLQTLMHGEHSNFIQIESALQALQKLYSQSQKEQRNLALELKYGLLLLK 454
L + T +E +QTL H +H ++ + E + Q+ +Y +
Sbjct: 434 SALNDCVTSLE-VQTLGHPKHHDYEKPEVKNLVNHQYIENGQQID-------EYQSVTAP 589
Query: 455 DLELSKQDFKEEMQGIVEENKTLHELNFSSTRSLKKQQMEISKLKEIKEKL 505
D L QD + + I K NF+++ + EI + KEI++K+
Sbjct: 590 DPLLDFQDLQRRINEISMAVK-----NFNASSKANVEMREIQEAKEIEQKM 727
Score = 30.4 bits (67), Expect = 6.0
Identities = 28/126 (22%), Positives = 55/126 (43%), Gaps = 7/126 (5%)
Frame = +3
Query: 49 EEADSFARRAEMYYKKRPELMKLVEEFYRAYRALAERYDHAMGELRHAHKTMPEAFPNSA 108
E DS +++ +LMK VE ++ + + YD +L + +T+ + F +
Sbjct: 1068 ERLDSDSQKLTNLQITIQDLMKKVETIEKSTKGKSAEYDTVKEQLEASQETVMKLFDANR 1247
Query: 109 YYILNDD----SPCGSLGPDAESHTSARPTHRSKKNERSSEESNG---EVQTLREALAKM 161
+ N + S G +++ S S++ +R SE+ EVQ L+ L K+
Sbjct: 1248 KLVKNVEEGALSSAGRASSESDEIGSVSRRRFSEQAQRGSEKIGQLQLEVQRLQFLLLKL 1427
Query: 162 QSDKDA 167
K++
Sbjct: 1428 NDAKES 1445
>TC92749 similar to GP|13486675|dbj|BAB39912. contains EST
AU062670(C30068)~similar to Arabidopsis thaliana
chromosome 2 At2g30500~unknown, partial (17%)
Length = 688
Score = 147 bits (370), Expect = 4e-35
Identities = 66/87 (75%), Positives = 76/87 (86%)
Frame = +2
Query: 8 ESRRLYSWWWDSHNSPKNSKWLLENLTDIDTKVKSMIKLIEEEADSFARRAEMYYKKRPE 67
ESR+ +SWWWDSH SPKNSKWL ENL ++D VK M+KLIEE+ADSFA++AEMYYKKRPE
Sbjct: 98 ESRKSHSWWWDSHISPKNSKWLFENLEEMDRNVKRMLKLIEEDADSFAKKAEMYYKKRPE 277
Query: 68 LMKLVEEFYRAYRALAERYDHAMGELR 94
L+ LVEEFYR YR+LAERYDH GELR
Sbjct: 278 LVALVEEFYRGYRSLAERYDHVTGELR 358
>TC83927 weakly similar to GP|9279697|dbj|BAB01254.1 centromere protein
{Arabidopsis thaliana}, partial (3%)
Length = 743
Score = 134 bits (338), Expect = 2e-31
Identities = 68/183 (37%), Positives = 120/183 (65%)
Frame = +1
Query: 723 TNLEEKYADVEKDKESAVNQVEELFASILVQKENHSNHKHSSEARLANLENIVRVLQEEQ 782
+ LE + +++ ++ES++ +VEEL S+ Q+E H +E +AN E + +L+E+
Sbjct: 7 SELELMHLELKGERESSLKKVEELLVSLYSQREEHCRVLKLNEDEVANKELQIDILKEDA 186
Query: 783 RLGKVEFEQELDRVVNAQIEMFILQNCIEELELKNFVLLTECEKLVEASKFSDKVISELE 842
+ K E+E+ELDR +NAQIE+FILQ CI++LE +NF LL EC++L+EASK SDK+IS LE
Sbjct: 187 KCRKQEYEEELDRSLNAQIEIFILQKCIQDLEKRNFSLLVECQRLLEASKMSDKIISNLE 366
Query: 843 SENLMQLIEEEFLLHRIRKFKMDIHKVCGVLQIDSDGGGDNEIKKEEIPISRILDKIESL 902
+EN+ + + + L +I+ ++ +H+V L I+ D ++ + + + ++ I K++
Sbjct: 367 TENIQKQDDVDSLSDKIKILRVGLHQVLKTLDINGDNFFEDMLDENQTLLNHIHGKLKER 546
Query: 903 ESS 905
+ S
Sbjct: 547 KKS 555
Score = 35.0 bits (79), Expect = 0.24
Identities = 20/64 (31%), Positives = 34/64 (52%)
Frame = +2
Query: 896 LDKIESLESSLVKSQEENQQLLVENSVLLGSLQQHQSEGEKLKLEKKTVEQEFENMREQN 955
++ + S L + E+ L VENSVL+ L+Q + E L +EK +++E + +Q
Sbjct: 527 MENLRRERSLLTRFSRESHHLTVENSVLITFLEQLKMTVENLVIEKGALDEESKIQSKQF 706
Query: 956 VILQ 959
LQ
Sbjct: 707 TALQ 718
>TC93427 similar to GP|9280668|gb|AAF86537.1| F21B7.9 {Arabidopsis
thaliana}, partial (25%)
Length = 710
Score = 111 bits (278), Expect = 2e-24
Identities = 68/172 (39%), Positives = 94/172 (54%), Gaps = 24/172 (13%)
Frame = +3
Query: 14 SWWW--DSH-NSPKNSKWLLENLTDIDTKVKSMIKLIEEEADSFARRAEMYYKKRPELMK 70
+WWW +SH N+ K S WL L++++ K +M+KLIEE+ADSFA+RAEMYYKKRPEL+
Sbjct: 111 NWWWLDNSHTNNTKRSPWLQSTLSELNEKTNAMLKLIEEDADSFAKRAEMYYKKRPELVS 290
Query: 71 LVEEFYRAYRALAERYDHAMGELRHAHKTMPEAFPNSAYYILNDDSPCGSLG-------- 122
+VE+FYR++R+LAERYD + + H + + SA Y L G
Sbjct: 291 MVEDFYRSHRSLAERYDQVKPDTGNGHLVLGGSPFASAKYQLEKLMSFADTGYDTYSENF 470
Query: 123 ------------PDAESHTSARPTHRSKKNER-SSEESNGEVQTLREALAKM 161
P+ E A+ H K ER SS N EV LR+ + K+
Sbjct: 471 DVDESMESEVDDPEHEEKDLAKFHHNYTKEERVSSVAVNDEVMRLRDEIKKI 626
>AW561041 similar to GP|13486675|db contains EST AU062670(C30068)~similar to
Arabidopsis thaliana chromosome 2 At2g30500~unknown,
partial (17%)
Length = 694
Score = 102 bits (255), Expect = 1e-21
Identities = 53/93 (56%), Positives = 64/93 (67%), Gaps = 13/93 (13%)
Frame = +1
Query: 15 WWWDS-HNSPKNSKWLL------------ENLTDIDTKVKSMIKLIEEEADSFARRAEMY 61
W+W HNS N +LL +++D VK M+KLIEE+ADSFA++AEMY
Sbjct: 247 WFWQKLHNSGLNDYYLL**FFCW*IEC*LSLTSEMDRNVKRMLKLIEEDADSFAKKAEMY 426
Query: 62 YKKRPELMKLVEEFYRAYRALAERYDHAMGELR 94
YKKRPEL+ LVEEFYR YR+LAERYDH GELR
Sbjct: 427 YKKRPELVALVEEFYRGYRSLAERYDHVTGELR 525
Score = 49.3 bits (116), Expect = 1e-05
Identities = 20/26 (76%), Positives = 22/26 (83%)
Frame = +1
Query: 8 ESRRLYSWWWDSHNSPKNSKWLLENL 33
ESR+ +SW WDSH SPKNSKWL ENL
Sbjct: 40 ESRKSHSWCWDSHISPKNSKWLFENL 117
>TC88896 weakly similar to GP|6728968|gb|AAF26966.1| unknown protein
{Arabidopsis thaliana}, partial (27%)
Length = 1097
Score = 73.6 bits (179), Expect = 6e-13
Identities = 92/385 (23%), Positives = 180/385 (45%), Gaps = 28/385 (7%)
Frame = +2
Query: 379 LQLEAENL---VQRIASKDHELLEKHTEIERLQTLMHGEHSNFIQIESALQALQKLYSQS 435
LQ E E L +++ D +L EK T IE+L +ES + + Y++S
Sbjct: 8 LQTEIEALKHELEKAKGYDEKLAEKETLIEQLN------------VESEAAKMAESYARS 151
Query: 436 -----QKEQRNLALELKYGLLLLKDLELSKQDFKEEMQGIVEENKTLHELNFSSTRSLKK 490
+K+ L ++++ L + LS + ++++G +N+ LH+ S SLK+
Sbjct: 152 VLDECRKKVEELEMKVEEANQLERSASLSLETATKQLEG---KNELLHDAE-SEISSLKE 319
Query: 491 Q----QMEISKLKEIKEKLEREFHTSTEESNVLQRETHQIKDDIQHLNERYQAMLEQLQS 546
+ +M + + + E ER + EE+ + ++ ++ +I+ +++ E+ Q+
Sbjct: 320 KLGMLEMTVGRQRGDLEDAERCLLAAKEENIEMSKKIESLESEIETVSK------EKAQA 481
Query: 547 LGLNPNSFAASVRDLQNENFML---KETCKKEHSE-KEALREKSKDMNEVLMENACMEFS 602
L N A+SV+ L E L E C+ E + K A+ + ++EV E +
Sbjct: 482 LN-NEKLSASSVQTLLEEKNKLINELEICRDEEEKTKLAMDSLASALHEVSAEARDTKEK 658
Query: 603 LLGLNDELDGLRGTVKEIQQFCQVLQEE-KSILAD---EKSTLLSQLQIITESMQKILEN 658
LL E + +++++ + +E+ +S+L D E L S L+ E + +L +
Sbjct: 659 LLANQAEHESYETQIEDLKSDLEASKEKYESMLNDAHHEIEDLKSDLEASKEKYESMLND 838
Query: 659 N----TVLEKSLSDAKIEFEGLRI----KSGDLEDCCKLLNDEKNNLQNERSMLISQLEI 710
VL S+ ++K++ + K DL +C K +E ++L NE + LIS L+
Sbjct: 839 AHHEIDVLTSSIENSKMDILNSKAEWEQKEHDLVECIKRTEEENSSLGNEVNRLISLLKK 1018
Query: 711 VEEKLSNLEKKVTNLEEKYADVEKD 735
EE+ + ++ T L+E +VE +
Sbjct: 1019TEEEANVKREEETQLKENMKEVEAE 1093
Score = 49.3 bits (116), Expect = 1e-05
Identities = 85/365 (23%), Positives = 161/365 (43%), Gaps = 24/365 (6%)
Frame = +2
Query: 776 RVLQEEQRLGKVEFEQELDRVVNAQIEMFILQNC---IEELELKNFVLLTECEKLVEASK 832
++ ++E + ++ E E ++ + +L C +EELE+K + E +L ++
Sbjct: 68 KLAEKETLIEQLNVESEAAKMAESYARS-VLDECRKKVEELEMK----VEEANQLERSAS 232
Query: 833 FS-DKVISELESEN-LMQLIEEEF--LLHRIRKFKMDIHKVCGVLQIDSDGGGDNE---- 884
S + +LE +N L+ E E L ++ +M + + G L+ D E
Sbjct: 233 LSLETATKQLEGKNELLHDAESEISSLKEKLGMLEMTVGRQRGDLE-------DAERCLL 391
Query: 885 -IKKEEIPISRILDKIESLESSLVKSQEENQQLLVENSVLLGSLQQHQSEGEKLKLEKKT 943
K+E I +S+ KIESLES + +E Q L + S+Q E KL E +
Sbjct: 392 AAKEENIEMSK---KIESLESEIETVSKEKAQALNNEKLSASSVQTLLEEKNKLINELEI 562
Query: 944 VEQEFENMREQNVILQKDKVELLEENRQLRIEVVNGVEKENRSKSTLAALQAEMIELRQT 1003
E E+ L D L ++ E + EK +++ + + ++ +L+
Sbjct: 563 CRDE-----EEKTKLAMD--SLASALHEVSAEARDTKEKLLANQAEHESYETQIEDLKSD 721
Query: 1004 NQVFQEENGKMLDEKNSLCRNV-SDLKDAKSSAEDENSVMFH--DVLALSNLNLVYEIFF 1060
+ +E+ ML++ + ++ SDL+ +K E + H DVL S N +I
Sbjct: 722 LEASKEKYESMLNDAHHEIEDLKSDLEASKEKYESMLNDAHHEIDVLTSSIENSKMDILN 901
Query: 1061 TENMVEKRA--LCEHLGNL----SHLNNDLNQEFGVLRKNFE---VKEAENVYLNESIER 1111
++ E++ L E + S L N++N+ +L+K E VK E L E+++
Sbjct: 902 SKAEWEQKEHDLVECIKRTEEENSSLGNEVNRLISLLKKTEEEANVKREEETQLKENMKE 1081
Query: 1112 MDKEL 1116
++ E+
Sbjct: 1082 VEAEV 1096
Score = 45.4 bits (106), Expect = 2e-04
Identities = 95/407 (23%), Positives = 167/407 (40%), Gaps = 13/407 (3%)
Frame = +2
Query: 1182 LQFEMKTLLHEVEQHRVREEALN-----LELLNKENEF-KLWENEAAAFYHDLQMSSICL 1235
LQ E++ L HE+E+ + +E L +E LN E+E K+ E+ A + + + +
Sbjct: 8 LQTEIEALKHELEKAKGYDEKLAEKETLIEQLNVESEAAKMAESYARSVLDECRKK---V 178
Query: 1236 ALLESKVSELTGVCKI--LDDESSAKSLENEHMREIISLLESEIGGLKEQLSAYVPLVSS 1293
LE KV E + + L E++ K LE ++ E++ ESEI LKE+L V
Sbjct: 179 EELEMKVEEANQLERSASLSLETATKQLEGKN--ELLHDAESEISSLKEKLGMLEMTVGR 352
Query: 1294 LKEDFNSLEHISLLWTKRNSVVGNGAQKDVVIETCLRKHSHQSARE-NEIVLIPDGVSDL 1352
+ D E L + N + ++K +E+ + S + A+ N L V L
Sbjct: 353 QRGDLEDAERCLLAAKEENIEM---SKKIESLESEIETVSKEKAQALNNEKLSASSVQTL 523
Query: 1353 LTLQTR-IRAVEKIMMEELKRRVKQKSLTTESTPYSSLEIATYPKVENRKKEIELVEENV 1411
L + + I +E EE K ++ SL + S+ T K+ + E E E +
Sbjct: 524 LEEKNKLINELEICRDEEEKTKLAMDSLASALHEVSAEARDTKEKLLANQAEHESYETQI 703
Query: 1412 FDRNS--WRKKPKIRLLMKDIPLDRNVDDQNSKYLKREHRRTNDHVLELCENNEHE-PLS 1468
D S K K ++ D ++D LK + + + + + HE +
Sbjct: 704 EDLKSDLEASKEKYESMLND--AHHEIED-----LKSDLEASKEKYESMLNDAHHEIDVL 862
Query: 1469 APTVDHAMICHRSDDSGRYLNYSSELDIEKELGVDKLELSKSVKEKTEDDKRRILERLSS 1528
+++ +S++DI SK+ E+ E D ++R
Sbjct: 863 TSSIE-----------------NSKMDILN---------SKAEWEQKEHDLVECIKRTEE 964
Query: 1529 DGQKLAILKMALQDLKKKTETKKKSKQGNDIEYETVKRHIEEVEEAV 1575
+ L L L KKTE + K+ + + +K +++EVE V
Sbjct: 965 ENSSLGNEVNRLISLLKKTEEEANVKREEETQ---LKENMKEVEAEV 1096
>TC86622 weakly similar to GP|20197623|gb|AAM15156.1 putative myosin heavy
chain {Arabidopsis thaliana}, partial (17%)
Length = 2546
Score = 68.9 bits (167), Expect = 2e-11
Identities = 103/466 (22%), Positives = 207/466 (44%), Gaps = 10/466 (2%)
Frame = +2
Query: 722 VTNLEEKYADVEKDKESAVNQVEELFASILVQKENHSNHKHSSEARLANLENIVRVLQEE 781
++ LE++ +D E++ ++ V +L + + S + R ++LEN ++ E
Sbjct: 2 ISALEKRCSDSEENSNKYLDNVSDLTSEL-----------ESFKVRTSSLENTLQTANES 148
Query: 782 QRLGKVEFEQELDRVVNAQIEMFILQNCIEELELKNFVLLTECEKLVEA-SKFSDKVISE 840
+ +E ++ L+ V + E +KL +A + S+K+
Sbjct: 149 E----IELKESLNAVTD------------------------EKKKLEDALNSLSEKLA-- 238
Query: 841 LESENLMQLIEEEFLLHRIRKFKMDIHKVCGVLQIDSDGGGDNEIKKEEIPISRILDKIE 900
ESENL++++ ++ L +++ LQ +N++K E+ S I +K
Sbjct: 239 -ESENLLEIVRDDLNLTQVK------------LQ-----STENDLKAAELRESEIREKHN 364
Query: 901 SLESSLVKSQEENQQLLVENSVLLGSLQQHQSEGEKLKLEKKTVEQEFENMREQNVILQK 960
++E +L + + +L ++ L SL + + + KL++ E N ++ V
Sbjct: 365 AIEENLAV-RGRDIELTSARNLELESLHESLTRDSEQKLQEAI---EKFNSKDSEVQSLL 532
Query: 961 DKVELLEENRQLRIEVVNGVEKE-NRSKSTLAALQAEMIELRQ--------TNQVFQEEN 1011
+K+++LEEN E ++ E S S LA+LQ+E +L++ T+Q F E N
Sbjct: 533 EKIKILEENIAGAGEQSISLKSEFEESLSKLASLQSENEDLKRQIVEAEKKTSQSFSE-N 709
Query: 1012 GKMLDEKNSLCRNVSDLKDAKSSAEDENSVMFHDVLALSNLNLVYEIFFTENMVEKRALC 1071
++ L + +L+++ +S E V +++ S+ NL+ E+ N V+ ++
Sbjct: 710 ELLVGTNIQLKTKIDELQESLNSVVSEKEVTAQELV--SHKNLLAEL----NDVQSKSSE 871
Query: 1072 EHLGNLSHLNNDLNQEFGVLRKNFEVKEAENVYLNESIERMDKELLEMDKRLKAAETSNA 1131
H N + ++ L+K+ E KE+E LNE + ++ ++ K E
Sbjct: 872 IHSANEVRILEVESKLQEALQKHTE-KESETKELNEKLNTLEGQI-------KIYEEQAH 1027
Query: 1132 EFSRHIEELKMEQEESTKIKENLDRQILEQSENCMNHKKEIEHLNE 1177
E E K E EES ++L+ + EQ + + E +NE
Sbjct: 1028 EAVAAAENRKAELEESLIKLKHLEAAVEEQQNKSLERETETAGINE 1165
Score = 46.2 bits (108), Expect = 1e-04
Identities = 67/277 (24%), Positives = 119/277 (42%), Gaps = 8/277 (2%)
Frame = +3
Query: 1127 ETSNAEFSRHIEELKMEQEESTKIKENLDRQILEQSENCMNHKKEIEHLNEANETLQFE- 1185
+T ++ S I++ + E + +K+ L+ IL+ E H+K + L ETL+ E
Sbjct: 1332 QTLKSQISSVIDDRNLLNETNQNLKKELESIILDLEEKLKEHQKNEDSLKSEVETLKIEI 1511
Query: 1186 -----MKTLLHEVEQHRVREEALNLELLNKENEFKLWENEAAAFYHDLQMSSICLALLES 1240
+++ LHE+E + E+ E ++ +AAA ++ +SS E
Sbjct: 1512 AEKSALQSRLHEIEAQLAKAESRLHE--------EVGSVQAAASQREVDLSS-KFEDYEQ 1664
Query: 1241 KVSELTGVCKILDDESSAKSLENEHMREIISLLESEIGGLKEQLSAYVPLVSSLKEDFNS 1300
K+SE+ + LE E + L + I K + S + L ++LK
Sbjct: 1665 KISEINVL------NGKVAELEKE-----LHLAQDTIANQKGEESQKLELEAALKNSVEE 1811
Query: 1301 LEHISLLWTKRNSVVGNGAQKDVV-IETCLRKHSHQSARENEIVLIPDGVSDLLTLQTRI 1359
LE TK+N + + QK V+ E L++ + + + E + D L +++R
Sbjct: 1812 LE------TKKNEI--SLLQKQVIEFEQKLQQADEKISVKGEEAV---DKKDALEVKSRD 1958
Query: 1360 RAVEKIMMEELKRRVKQKS-LTTESTPYSSLEIATYP 1395
++ KR+ K+KS TT T SS E P
Sbjct: 1959 FSIS----SPSKRKSKKKSEATTPQTSTSSSETHIQP 2057
Score = 46.2 bits (108), Expect = 1e-04
Identities = 56/280 (20%), Positives = 120/280 (42%), Gaps = 4/280 (1%)
Frame = +2
Query: 148 NGEVQTLREALAKMQSDKDALFLQYQESLENLSKMETDLNKAQNNARGLDDRASEAEIQV 207
N E+++L E+L + K QE++E + ++++ + L++ + A Q
Sbjct: 422 NLELESLHESLTRDSEQK------LQEAIEKFNSKDSEVQSLLEKIKILEENIAGAGEQS 583
Query: 208 EILK----ESLMQLKADKDAGEVLYNQCLETIARLESMLSQKDNIEAKNLKQELTRVVVQ 263
LK ESL +L + + E L Q +E + S+ + + N++
Sbjct: 584 ISLKSEFEESLSKLASLQSENEDLKRQIVEAEKKTSQSFSENELLVGTNIQ--------- 736
Query: 264 KDTVLLQYKQCLEKIPMLENKIALAEENSRMLNDQIERTELEVETLRKNLAEMNEERDSL 323
L+ KI +E+ + + E T E+ + + LAE+N+ +
Sbjct: 737 -----------------LKTKIDELQESLNSVVSEKEVTAQELVSHKNLLAELNDVQSKS 865
Query: 324 SVLYHHCLEKISKMENEILHVQENAEQLKNKIEKEAEKLEISEKHRGMLEKSNQNLQLEA 383
S ++ +I ++E+++ E L+ EKE+E E++EK LE + + +A
Sbjct: 866 SEIHSANEVRILEVESKL------QEALQKHTEKESETKELNEK-LNTLEGQIKIYEEQA 1024
Query: 384 ENLVQRIASKDHELLEKHTEIERLQTLMHGEHSNFIQIES 423
V ++ EL E +++ L+ + + + ++ E+
Sbjct: 1025HEAVAAAENRKAELEESLIKLKHLEAAVEEQQNKSLERET 1144
Score = 37.7 bits (86), Expect = 0.038
Identities = 43/176 (24%), Positives = 84/176 (47%), Gaps = 28/176 (15%)
Frame = +2
Query: 1482 DDSGRYLNYSSELDIEKE---------------LGVDKLELSKSVKEKTEDDKRRILERL 1526
++S +YL+ S+L E E ++EL +S+ T D+K+++ + L
Sbjct: 38 ENSNKYLDNVSDLTSELESFKVRTSSLENTLQTANESEIELKESLNAVT-DEKKKLEDAL 214
Query: 1527 SSDGQKLA----ILKMALQDLKKKTETKKKSKQGNDIEYETVKRHIEEVEEAVMQQVSIN 1582
+S +KLA +L++ DL T+ K +S + + E + I E A+ + +++
Sbjct: 215 NSLSEKLAESENLLEIVRDDL-NLTQVKLQSTENDLKAAELRESEIREKHNAIEENLAVR 391
Query: 1583 DQ-----MAKNVEEGA--SSLDREIPRRGSEQIGKLQFEVQNIQYIL--LKLAEEN 1629
+ A+N+E + SL R+ ++ E I K + +Q +L +K+ EEN
Sbjct: 392 GRDIELTSARNLELESLHESLTRDSEQKLQEAIEKFNSKDSEVQSLLEKIKILEEN 559
>TC86560 similar to GP|8843737|dbj|BAA97285.1 myosin heavy chain-like
{Arabidopsis thaliana}, partial (28%)
Length = 2450
Score = 63.9 bits (154), Expect = 5e-10
Identities = 116/530 (21%), Positives = 217/530 (40%), Gaps = 28/530 (5%)
Frame = +2
Query: 342 LHVQENAEQLKNKIEKEAEKLEISEKHRGMLEKSNQNLQL-EAENLVQRIASKDHELLEK 400
+H + A + + KE + L ++EK L++ +N + +A+ LV+ + ++
Sbjct: 443 IHKKPKAYSAERVLAKETQ-LHVAEKELNKLKEQVKNAETTKAQALVE--LERAQRTVDD 613
Query: 401 HTEIERLQTLMHGEHSNFIQIESALQALQKLYSQSQ------KEQRNLALELKYGLLLLK 454
T+ +L+ + S E+A ++ Y +S KE+ A++ +Y ++
Sbjct: 614 LTQ--KLKLITESRESAVKATEAAKSQAKQKYGESDGVNGAWKEELENAVQ-RYASIMT- 781
Query: 455 DLELSKQDFKEEMQGIVEENKTLHELNFSSTRSLKKQQMEISKLKEIKEKLEREFHTSTE 514
+L+ +KQD + KT E + SS +S +K +E E +TE
Sbjct: 782 ELDAAKQDLR----------KTRQEYDSSS-------DARVSAVKRTEEA-ENAMKENTE 907
Query: 515 ESNVLQRETHQIKDDIQHLNERYQAMLEQLQSLGLNPNSFAASVRDLQNENFMLKETCKK 574
+ L +E +K+ I+ Y +Q Q+L L E L+++ K
Sbjct: 908 RVSELSKEISAVKESIEQTKLAYVESQQQ-QALVLT-------------EKDALRQSYKA 1045
Query: 575 --EHSEKEALREKSKDMNEVLMENACMEFSLLGLNDELDGLRGTVKEIQQFCQVLQEEKS 632
E S+K+ L K K+ N + ++ + L T+ EI L+ ++S
Sbjct: 1046TLEQSKKKLLALK-KEFNPEITKSLEAQ------------LAETMNEIAALHTELENKRS 1186
Query: 633 ILADEKSTLLSQLQIITESMQKILENNTVLEKSLSDAKIEFEGLRIKSGDLEDCCKLLND 692
D T+ S+L ES+QK+++ L + K+E E ++ + +L++ L
Sbjct: 1187SDLDSVKTVTSELDGAKESLQKVVDEENTLRSLVETLKVELENVKKEHSELKEKESELES 1366
Query: 693 EKNNLQNERSMLISQLEIV-----------EEKLSNLEKKVTNLEEKYADVEKDKESAVN 741
NL + S+LE EE + L + + EE +VE K
Sbjct: 1367TVGNLHVKLRKSKSELEACSADESKVRGASEEMILTLSRLTSETEEARREVEDMKNKTDE 1546
Query: 742 QVEELFASILVQKENHSNHKHSSE----ARLANLENI--VRVLQEEQRLGKVEFEQELDR 795
+E A+ L +E K ++E A+ A I + VL E + E
Sbjct: 1547LKKEAEATKLALEEAEKKLKEATEEAEAAKAAEASAIEQITVLTERTSARRASTSSESGA 1726
Query: 796 VVNAQIEMF-ILQNCIEELELKNFVLLTECEKLVEASKFSD-KVISELES 843
+ E F L+ +EE + + + + VEA K S+ +V+ +LE+
Sbjct: 1727AMTISTEEFESLKRKVEESDKLADMKVDAAKAQVEAVKASENEVLKKLEA 1876
Score = 46.6 bits (109), Expect = 8e-05
Identities = 85/451 (18%), Positives = 185/451 (40%), Gaps = 33/451 (7%)
Frame = +2
Query: 20 HNSPKNSKWLLENLTDIDT-KVKSMIKL--IEEEADSFARRAEMYYKKRPELMKLVEEFY 76
H + K L E + + +T K +++++L + D ++ ++ + R +K E
Sbjct: 503 HVAEKELNKLKEQVKNAETTKAQALVELERAQRTVDDLTQKLKLITESRESAVKATEA-- 676
Query: 77 RAYRALAERYDHAMGELRHAHKTMPEAFPNSAYYILNDDSPCGSLGP-----DAESHTSA 131
A ++Y + G + + A A + D+ L D+ S
Sbjct: 677 -AKSQAKQKYGESDGVNGAWKEELENAVQRYASIMTELDAAKQDLRKTRQEYDSSSDARV 853
Query: 132 RPTHRSKKNERSSEESNGEVQTLREALAKMQSDKDALFLQYQESLENLSKMETDLNKAQN 191
R+++ E + +E+ V L + ++ ++ + L Y ES + + + T+ + +
Sbjct: 854 SAVKRTEEAENAMKENTERVSELSKEISAVKESIEQTKLAYVESQQQQALVLTEKDALRQ 1033
Query: 192 NARGLDDRASEAEIQVEILKESLMQLKAD--KDAGEVLYNQCLETIARLESMLSQKDNIE 249
+ + +E K+ L+ LK + + + L Q ET+ + ++ ++ +N
Sbjct: 1034SYKAT----------LEQSKKKLLALKKEFNPEITKSLEAQLAETMNEIAALHTELENKR 1183
Query: 250 AKNLKQELTRVVVQKDTVLLQYKQCLEKIPMLENKI-ALAE------ENSRMLNDQIERT 302
+ +L T V + D K+ L+K+ EN + +L E EN + + +++
Sbjct: 1184SSDLDSVKT-VTSELDGA----KESLQKVVDEENTLRSLVETLKVELENVKKEHSELKEK 1348
Query: 303 ELEVET--------LRKNLAEM---NEERDSLSVLYHHCLEKISKMENEILHVQENAEQL 351
E E+E+ LRK+ +E+ + + + + +S++ +E + E +
Sbjct: 1349ESELESTVGNLHVKLRKSKSELEACSADESKVRGASEEMILTLSRLTSETEEARREVEDM 1528
Query: 352 KNKI-----EKEAEKLEISEKHRGMLEKSNQNLQLEAENLVQRIASKDHELLEKHTEIER 406
KNK E EA KL + E + + E + + EA + A + +L + T R
Sbjct: 1529KNKTDELKKEAEATKLALEEAEKKLKEATE---EAEAAKAAEASAIEQITVLTERTSARR 1699
Query: 407 LQTLMHGEHSNFIQIESALQALQKLYSQSQK 437
T + I E ++L++ +S K
Sbjct: 1700ASTSSESGAAMTISTEE-FESLKRKVEESDK 1789
>BI309151 similar to GP|4894628|gb|A NT3 {Nicotiana tabacum}, partial (5%)
Length = 663
Score = 63.2 bits (152), Expect = 8e-10
Identities = 37/78 (47%), Positives = 47/78 (59%), Gaps = 14/78 (17%)
Frame = -2
Query: 1599 EIPRRGSEQIGKLQFEVQNIQYILLKLAEENNNKVKNRISRKTGILLR-----------R 1647
E R+GSE+IG+LQFE+QNI Y+LL L++E NK KN+ S KT + LR
Sbjct: 557 EQARKGSEEIGRLQFEMQNIHYVLLNLSDEKKNKGKNKFSGKTVVFLRDFIHIGNKSSSS 378
Query: 1648 KLRVCGCSG---PSTNED 1662
K R GC G +TNED
Sbjct: 377 KRRSKGCFGGTSKTTNED 324
>BI273318 similar to GP|7269756|emb predicted protein of unknown function
{Arabidopsis thaliana}, partial (5%)
Length = 589
Score = 58.9 bits (141), Expect = 2e-08
Identities = 42/135 (31%), Positives = 60/135 (44%), Gaps = 37/135 (27%)
Frame = +3
Query: 1565 KRHIEEVEEAVMQQVSINDQMAKNVEEGASSLD------------------REIPRRGSE 1606
K +E +EA+ + N ++ KNVEEG SS E RRGSE
Sbjct: 3 KGQLESAQEAITKLFDANRKLMKNVEEGTSSFAGKSTTVSDESGSVSRRRVSEQARRGSE 182
Query: 1607 QIGKLQFEVQNIQYILLKLAEENNNKVKNRI---SRKTGILLR----------------R 1647
+IG+LQ EVQ +Q++LLKL ++ +K K + + +LLR +
Sbjct: 183 KIGRLQLEVQRLQFLLLKLNDDKESKGKGKAMIDDQNPRVLLRDYLYGGTRKSYHKLKKK 362
Query: 1648 KLRVCGCSGPSTNED 1662
K C C P T D
Sbjct: 363 KASFCACVQPPTKGD 407
>TC79642 similar to PIR|T07397|T07397 kinesin heavy chain-like protein
(clone PKCBP) - potato, partial (42%)
Length = 2321
Score = 57.8 bits (138), Expect = 4e-08
Identities = 57/265 (21%), Positives = 119/265 (44%), Gaps = 5/265 (1%)
Frame = +2
Query: 500 EIKEKLEREFHTSTEESNVLQRETHQIKDDIQHLNERYQAMLEQLQSLGLNPNSFAASVR 559
E EK ++ EES QR Q+ D+ E+ M EQL+ L + + +++
Sbjct: 203 ESYEKRIQDLSKLVEES---QRSADQLHKDLHEKQEKEVKMQEQLEGLKESLKANKQNLQ 373
Query: 560 DLQNENFMLKETCKKEHSEKEALREKSK-----DMNEVLMENACMEFSLLGLNDELDGLR 614
+ ++ L+ C K+ +A+ SK N+VL + +++ L ELD
Sbjct: 374 AVTSDCERLRSLCNKKDQALQAIESTSKKDLVETNNQVLQK---LKYELKYCKGELDSAE 544
Query: 615 GTVKEIQQFCQVLQEEKSILADEKSTLLSQLQIITESMQKILENNTVLEKSLSDAKIEFE 674
T+K ++ +L+++ S+L S S L ++K+ + ++ + D + + E
Sbjct: 545 ETIKTLRSEKAILEQKLSVLEKRNSEESSSL------LRKLEQERKAVKSEVYDLERKIE 706
Query: 675 GLRIKSGDLEDCCKLLNDEKNNLQNERSMLISQLEIVEEKLSNLEKKVTNLEEKYADVEK 734
G R + + + + E + LQN +LE + E ++++K E+ A + K
Sbjct: 707 GYRQELMAAKSIISVKDSELSALQNN----FKELEELREMKEDIDRK----NEQTASILK 862
Query: 735 DKESAVNQVEELFASILVQKENHSN 759
+ + + ++E L+ V ++ + N
Sbjct: 863 MQRAQLAEMEGLYKEEQVLRKRYFN 937
Score = 39.3 bits (90), Expect = 0.013
Identities = 51/229 (22%), Positives = 103/229 (44%), Gaps = 33/229 (14%)
Frame = +2
Query: 347 NAEQLKNKIEKEAEKLEISEK-----HRGMLEKSNQNLQLEA--ENLVQRIASKDHELLE 399
N E + +I+ ++ +E S++ H+ + EK + ++++ E L + + + L
Sbjct: 197 NLESYEKRIQDLSKLVEESQRSADQLHKDLHEKQEKEVKMQEQLEGLKESLKANKQNLQA 376
Query: 400 KHTEIERLQTLMHGEHSNFIQIESALQALQKLYSQSQKEQRNLALE-LKYGLLLLK-DLE 457
++ ERL++L + + + ALQA++ + E N L+ LKY L K +L+
Sbjct: 377 VTSDCERLRSLCNKK-------DQALQAIESTSKKDLVETNNQVLQKLKYELKYCKGELD 535
Query: 458 LSKQDFK--EEMQGIVEENKTLHELNFS--STRSLKKQQMEISKLKEIKEKLEREFH--- 510
+++ K + I+E+ ++ E S S+ L+K + E +K LER+
Sbjct: 536 SAEETIKTLRSEKAILEQKLSVLEKRNSEESSSLLRKLEQERKAVKSEVYDLERKIEGYR 715
Query: 511 -----------TSTEESNVLQ------RETHQIKDDIQHLNERYQAMLE 542
E + LQ E ++K+DI NE+ ++L+
Sbjct: 716 QELMAAKSIISVKDSELSALQNNFKELEELREMKEDIDRKNEQTASILK 862
>TC82817 similar to GP|9294158|dbj|BAB02060.1 contains similarity to
unknown protein~gb|AAD49756.1~gene_id:MKP6.26
{Arabidopsis thaliana}, partial (8%)
Length = 650
Score = 54.3 bits (129), Expect = 4e-07
Identities = 22/48 (45%), Positives = 33/48 (67%)
Frame = +1
Query: 48 EEEADSFARRAEMYYKKRPELMKLVEEFYRAYRALAERYDHAMGELRH 95
EEE D+FA RAE YY+KRP+L+ L+ + Y Y L++RY + + +H
Sbjct: 46 EEEGDTFAERAETYYQKRPQLLSLLHDLYNGYVTLSDRYIQTLAKHKH 189
>TC84423 similar to PIR|F84730|F84730 probable myosin heavy chain [imported]
- Arabidopsis thaliana, partial (10%)
Length = 787
Score = 52.0 bits (123), Expect = 2e-06
Identities = 47/218 (21%), Positives = 96/218 (43%), Gaps = 9/218 (4%)
Frame = +2
Query: 177 ENLSKMETDLNKAQNN-------ARGLDDRASEAEIQVEILKESLMQLKADKDAGEVLYN 229
E+ SK ++ L++A +N + L+D +E+ + ++L+ +A
Sbjct: 143 ESFSKTDSLLSQALSNNSELEQKVKSLEDLHNESGAVAATASQRSLELEGHIEATNAAAE 322
Query: 230 QCLETIARLES--MLSQKDNIEAKNLKQELTRVVVQKDTVLLQYKQCLEKIPMLENKIAL 287
+ + LE+ + +++ N+E L+Q+L V ++ + + EKI L+ K+
Sbjct: 323 EAKSQLRELETRFIAAEQKNVE---LEQQLNLVQLKANDAERDVTEFSEKISHLDAKLKE 493
Query: 288 AEENSRMLNDQIERTELEVETLRKNLAEMNEERDSLSVLYHHCLEKISKMENEILHVQEN 347
AEE +LN ++ ++ L +L + ++ L EK S+ E+ E
Sbjct: 494 AEEEKNLLNSLLQEHMDKLSQLESDLNQSTQKNSQLEEELKIVKEKCSEHEDRATMNNER 673
Query: 348 AEQLKNKIEKEAEKLEISEKHRGMLEKSNQNLQLEAEN 385
+ +L++ I+ K E EK LE + E +N
Sbjct: 674 SRELEDLIQSSHSKSEKCEKRASELELLLEQRNTEFQN 787
Score = 41.2 bits (95), Expect = 0.003
Identities = 57/291 (19%), Positives = 114/291 (38%), Gaps = 5/291 (1%)
Frame = +2
Query: 690 LNDEKNNLQNERSMLISQLEIVEEKLSNLEKKVTNLEEKYADVEKDKESAVNQVEELFAS 749
LN EK L+ L + +E S+LE+K+ +E ++ + A++ EL
Sbjct: 32 LNAEKKGLEETVEDLTVNAKHFKELCSDLEEKLKLSDESFSKTDSLLSQALSNNSELEQK 211
Query: 750 ILVQKENHSNH---KHSSEARLANLENIVRVLQEEQRLGKVEFEQELDRVVNAQIEMFIL 806
+ ++ H+ ++ R LE + K + + R + A+
Sbjct: 212 VKSLEDLHNESGAVAATASQRSLELEGHIEATNAAAEEAKSQLRELETRFIAAE------ 373
Query: 807 QNCIEELELKNFVLLTECEKLVEASKFSDKVISELESENLMQLIEEEFLLHRIRKFKMDI 866
Q +E + N V L + + ++FS+K IS L+++ L + EE+ LL+ +
Sbjct: 374 QKNVELEQQLNLVQLKANDAERDVTEFSEK-ISHLDAK-LKEAEEEKNLLNSL------- 526
Query: 867 HKVCGVLQIDSDGGGDNEIKKEEIPISRILDKIESLESSLVKSQEENQQLLVENSVLLGS 926
+ +DK+ LES L +S ++N QL E ++
Sbjct: 527 -------------------------LQEHMDKLSQLESDLNQSTQKNSQLEEELKIVKEK 631
Query: 927 LQQHQSEGEKLKLEKKTVEQEFENMREQNVILQK--DKVELLEENRQLRIE 975
+H+ + +E ++ ++ +K ++ELL E R +
Sbjct: 632 CSEHEDRATMNNERSRELEDLIQSSHSKSEKCEKRASELELLLEQRNTEFQ 784
Score = 40.8 bits (94), Expect = 0.004
Identities = 54/261 (20%), Positives = 110/261 (41%), Gaps = 14/261 (5%)
Frame = +2
Query: 526 IKDDIQHLNERYQAMLEQLQSLGLNPNSFAASVRDLQNENFMLKETCKKEHS-------E 578
+++++ LN + + E ++ L +N F DL+ + + E+ K S
Sbjct: 11 VQEELTKLNAEKKGLEETVEDLTVNAKHFKELCSDLEEKLKLSDESFSKTDSLLSQALSN 190
Query: 579 KEALREKSKDMNEVLMENACMEFSLLGLNDELDGLRGTVKEIQQFCQVLQEEKSILADEK 638
L +K K + ++ E+ + + + EL+G I+ +E KS L + +
Sbjct: 191 NSELEQKVKSLEDLHNESGAVAATASQRSLELEG------HIEATNAAAEEAKSQLRELE 352
Query: 639 STLLSQLQIITESMQKILENNTVLEKSLSDAKIEFEGLRIKSGDLEDCCKLLNDEKNNL- 697
+ ++ Q E Q++ +++ +DA+ + K L+ K +EKN L
Sbjct: 353 TRFIAAEQKNVELEQQL----NLVQLKANDAERDVTEFSEKISHLDAKLKEAEEEKNLLN 520
Query: 698 ------QNERSMLISQLEIVEEKLSNLEKKVTNLEEKYADVEKDKESAVNQVEELFASIL 751
++ S L S L +K S LE+++ ++EK + E + + +N L
Sbjct: 521 SLLQEHMDKLSQLESDLNQSTQKNSQLEEELKIVKEKCS--EHEDRATMNNERSRELEDL 694
Query: 752 VQKENHSNHKHSSEARLANLE 772
+Q + + K E R + LE
Sbjct: 695 IQSSHSKSEK--CEKRASELE 751
Score = 39.7 bits (91), Expect = 0.010
Identities = 61/285 (21%), Positives = 104/285 (36%), Gaps = 34/285 (11%)
Frame = +2
Query: 931 QSEGEKLKLEKKTVEQEFENMR-------------EQNVILQKDKV--------ELLEEN 969
Q E KL EKK +E+ E++ E+ + L + + L N
Sbjct: 14 QEELTKLNAEKKGLEETVEDLTVNAKHFKELCSDLEEKLKLSDESFSKTDSLLSQALSNN 193
Query: 970 RQLRIEVVNGVEKENRSKSTLAALQAEMIELR----QTNQVFQEENGKM---------LD 1016
+L +V + + N S + A +EL TN +E ++ +
Sbjct: 194 SELEQKVKSLEDLHNESGAVAATASQRSLELEGHIEATNAAAEEAKSQLRELETRFIAAE 373
Query: 1017 EKNSLCRNVSDLKDAKSSAEDENSVMFHDVLALSNLNLVYEIFFTENMVEKRALCEHLGN 1076
+KN +L K++ + + F + +S+L+ + E + L EH+
Sbjct: 374 QKNVELEQQLNLVQLKANDAERDVTEFSE--KISHLDAKLKEAEEEKNLLNSLLQEHMDK 547
Query: 1077 LSHLNNDLNQEFGVLRKNFEVKEAENVYLNESIERMDKELLEMDKRLKAAETSNAEFSRH 1136
LS L +DLNQ +KN +++E + + E D+ T N E SR
Sbjct: 548 LSQLESDLNQS---TQKNSQLEEELKIVKEKCSEHEDR------------ATMNNERSRE 682
Query: 1137 IEELKMEQEESTKIKENLDRQILEQSENCMNHKKEIEHLNEANET 1181
+E+L +SE C E+E L E T
Sbjct: 683 LEDLIQSSH--------------SKSEKCEKRASELELLLEQRNT 775
>CA858495
Length = 763
Score = 49.7 bits (117), Expect = 1e-05
Identities = 65/270 (24%), Positives = 118/270 (43%), Gaps = 8/270 (2%)
Frame = +3
Query: 892 ISRILDKIESLESSLVKSQEENQQLLVENSVLLGSLQQHQSEGEKLKLEKKTVEQEFENM 951
I+++ ++ SL V E+ LL E + L+ + +E E L+ + +E +
Sbjct: 6 IAQLQNENNSLLQKEVGLAEKTNLLLNEKAGLVEKVNMLLNEKEGLEQKVNILESNLSSF 185
Query: 952 REQNV-------ILQKDKVELLEENRQLRIEVVNGVEKENRSKSTLAALQAEMIELRQTN 1004
E+ +L K+K L ++ L + + EKE + +A LQ+E L Q
Sbjct: 186 SEKEAGSVDTTNLLLKEKEGLEQKLNILESNLSSFSEKETGLEMRIAQLQSETNSLLQKE 365
Query: 1005 QVFQEENGKMLDEKNSLCRNVSDLKDAKSSAEDENSVMFHDVLALSNLNLVYEIFFTENM 1064
F E+ ++L+EKN L V L+ AE + S +
Sbjct: 366 TGFVEKTNQLLNEKNILSLKVDSLERKIIHAESDLS----------------------SF 479
Query: 1065 VEKRALCEHLGNLSHLNNDLNQEFGVLRKNFEVKEAENVYLNESIERMDKELLEMDKRLK 1124
VEK E + +S+LN ++ G + E++E++N L E +++L E
Sbjct: 480 VEKENSTEEV--ISNLNGSISMLQGQVA---ELEESKNNLLLE-----NQQLRENSSSSS 629
Query: 1125 AAETSNA-EFSRHIEELKMEQEESTKIKEN 1153
A++ ++A + + EELK + EE+ + EN
Sbjct: 630 ASQDASAKDRASENEELKSQIEEAYMLVEN 719
Score = 48.5 bits (114), Expect = 2e-05
Identities = 67/236 (28%), Positives = 102/236 (42%), Gaps = 36/236 (15%)
Frame = +3
Query: 657 ENNTVLEKSLSDAKIEFEGLRIKSGDLEDCCKLLN------------------------- 691
ENN++L+K + A+ L K+G +E LLN
Sbjct: 24 ENNSLLQKEVGLAEKTNLLLNEKAGLVEKVNMLLNEKEGLEQKVNILESNLSSFSEKEAG 203
Query: 692 --DEKNNLQNERSMLISQLEIVEEKLSNLEKKVTNLEEKYADVEKDKESAVNQVEELFAS 749
D N L E+ L +L I+E LS+ +K T LE + A ++ + S + Q E F
Sbjct: 204 SVDTTNLLLKEKEGLEQKLNILESNLSSFSEKETGLEMRIAQLQSETNSLL-QKETGF-- 374
Query: 750 ILVQKENH-SNHKHSSEARLANLENIVRVLQEEQRLGK-VEFEQELDRVV-NAQIEMFIL 806
V+K N N K+ ++ +LE +++ E L VE E + V+ N + +L
Sbjct: 375 --VEKTNQLLNEKNILSLKVDSLER--KIIHAESDLSSFVEKENSTEEVISNLNGSISML 542
Query: 807 QNCIEELELKNFVLLTECEKLVEASKFS----DKVISELESEN--LMQLIEEEFLL 856
Q + ELE LL E ++L E S S D + SEN L IEE ++L
Sbjct: 543 QGQVAELEESKNNLLLENQQLRENSSSSSASQDASAKDRASENEELKSQIEEAYML 710
>TC93791 similar to GP|15810373|gb|AAL07074.1 unknown protein {Arabidopsis
thaliana}, partial (40%)
Length = 866
Score = 48.5 bits (114), Expect = 2e-05
Identities = 63/276 (22%), Positives = 119/276 (42%), Gaps = 1/276 (0%)
Frame = +1
Query: 380 QLEAENLVQRIASKDHELLEKHTEIERLQTLMHGEHSNFIQIESALQALQKLYSQSQKEQ 439
+LE+EN ++ ++ H E + E S Q+E ++ +K+ ++ +
Sbjct: 31 ELESENRKMKVELEEFRTEATHLENQEATIRRLEERSR--QLEQQME--EKVKEIAEIKH 198
Query: 440 RNLALELKYGLLLLKDLELSKQDFKEEMQGIVEENKTLHELNFSSTRSLKKQQMEISKLK 499
RNLA E + L +LK+ E QD + + V K LHEL ++L
Sbjct: 199 RNLAEENQKTLEILKEREQLLQDQLQSAKESVSNMKKLHEL-------------AQNQLF 339
Query: 500 EIKEKLEREFHTSTEESNVLQRETHQIKDDIQHLNERYQAMLEQLQSLGLNPNSFAASVR 559
E++ + E E E+N+L E + + + L + QLQ+ N +S
Sbjct: 340 ELRAQSEEERAAKQAEANLLMDEVERAQTMLLGLEREKGVLRSQLQT--ANEDSETKKSD 513
Query: 560 DLQNENFMLKETCKKEHSEKEALREKSKDMNEVLMENACMEFSLLGLNDELDGLRGTVKE 619
+L + N + + +L K K ++E+ ME +E +L +E + +
Sbjct: 514 NLDSNNAL-----------ENSLIAKEKLISELNMELHNIETTLSNEREE------HIND 642
Query: 620 IQQFCQVLQE-EKSILADEKSTLLSQLQIITESMQK 654
+++F +L E E SI+A +K + I + ++K
Sbjct: 643 VKKFTAMLNEKEASIVAMKKELQTRPTEKIVDDLRK 750
Score = 39.3 bits (90), Expect = 0.013
Identities = 59/294 (20%), Positives = 118/294 (40%), Gaps = 4/294 (1%)
Frame = +1
Query: 835 DKVISELESENLMQLIEEEFLLHRIRKFKMDIHKVCGVLQIDSDGGGDNEIKKEEIPISR 894
D +SELESEN +E + +F+ + ++ +E I R
Sbjct: 16 DLKLSELESENRKMKVE-------LEEFRTEA----------------THLENQEATIRR 126
Query: 895 ILDKIESLESSLVKSQEENQQLLVENSVLLGSLQQHQSEGEKLKLEKKTVEQEFENMREQ 954
+ ++ LE + + +E ++ N +++Q E LK ++ ++ + ++ +E
Sbjct: 127 LEERSRQLEQQMEEKVKEIAEIKHRNLA-----EENQKTLEILKEREQLLQDQLQSAKES 291
Query: 955 NVILQKDKVELLEENRQLRIEVVNGVEKENRSKSTLAALQAEMIELRQTNQVFQEENGKM 1014
+ K+ L +N+ + + E+E +K A L + +E QT + E +
Sbjct: 292 --VSNMKKLHELAQNQLFELRAQS--EEERAAKQAEANLLMDEVERAQTMLLGLEREKGV 459
Query: 1015 LDEKNSLCRNVSDLKDAKSSAEDENSVMFHDVLA----LSNLNLVYEIFFTENMVEKRAL 1070
L + S+ K KS D N+ + + ++A +S LN+ T E+
Sbjct: 460 LRSQLQTANEDSETK--KSDNLDSNNALENSLIAKEKLISELNMELHNIETTLSNERE-- 627
Query: 1071 CEHLGNLSHLNNDLNQEFGVLRKNFEVKEAENVYLNESIERMDKELLEMDKRLK 1124
EH+ ++ LN+ KEA V + + ++ E + D R K
Sbjct: 628 -EHINDVKKFTAMLNE-----------KEASIVAMKKELQTRPTEKIVDDLRKK 753
>BF646001 homologue to GP|20303561|gb| putative kinase {Oryza sativa
(japonica cultivar-group)}, partial (6%)
Length = 667
Score = 47.8 bits (112), Expect = 4e-05
Identities = 36/156 (23%), Positives = 82/156 (52%), Gaps = 2/156 (1%)
Frame = +2
Query: 296 NDQIERTELEVETLRKNLAEMNEERDSLSVLYHHCLEKISKMENEILHVQENAEQLKNKI 355
N++IE+ + E+E LRKN DS + L ++ +++ E++ + E+L+ I
Sbjct: 206 NEEIEKMKTEIEKLRKN-------SDSTAELE----KEAARLRREVVESKVEIEKLRKII 352
Query: 356 EKEAEKLEISEKHRGMLEKSNQNLQLEAENLVQRIASKDHELLEKHTEIERLQTLMHGE- 414
+++ K+EI EK L++ N ++++ L +RI + + +E++++ R++ M +
Sbjct: 353 DEKENKIEIVEKEGKELKQENVEMEMKVRELERRIGVIEMKEVEENSKRVRIEEEMKEKI 532
Query: 415 HSNFIQIESALQALQKLYSQSQKEQRN-LALELKYG 449
+ +++E L + +K ++ LE K+G
Sbjct: 533 YEKEMEVEELKSMLMGKKVEEEKWLKDKRVLEKKFG 640
Score = 36.6 bits (83), Expect = 0.084
Identities = 39/174 (22%), Positives = 75/174 (42%)
Frame = +2
Query: 349 EQLKNKIEKEAEKLEISEKHRGMLEKSNQNLQLEAENLVQRIASKDHELLEKHTEIERLQ 408
E +KN E E K EI EK +N AE L + A E++E EIE+L+
Sbjct: 191 ENIKNNEEIEKMKTEI--------EKLRKNSDSTAE-LEKEAARLRREVVESKVEIEKLR 343
Query: 409 TLMHGEHSNFIQIESALQALQKLYSQSQKEQRNLALELKYGLLLLKDLELSKQDFKEEMQ 468
++ + + +E + L++ + E + LE + G++ +K+
Sbjct: 344 KIIDEKENKIEIVEKEGKELKQ--ENVEMEMKVRELERRIGVIEMKE------------- 478
Query: 469 GIVEENKTLHELNFSSTRSLKKQQMEISKLKEIKEKLEREFHTSTEESNVLQRE 522
VEEN + + +++ME+ +LK + + E ++ VL+++
Sbjct: 479 --VEENSKRVRIEEEMKEKIYEKEMEVEELKSMLMGKKVEEEKWLKDKRVLEKK 634
Score = 36.2 bits (82), Expect = 0.11
Identities = 30/159 (18%), Positives = 72/159 (44%)
Frame = +2
Query: 256 ELTRVVVQKDTVLLQYKQCLEKIPMLENKIALAEENSRMLNDQIERTELEVETLRKNLAE 315
EL ++ + + + K +EK+ + A E+ + L ++ +++E+E LRK + E
Sbjct: 179 ELETENIKNNEEIEKMKTEIEKLRKNSDSTAELEKEAARLRREVVESKVEIEKLRKIIDE 358
Query: 316 MNEERDSLSVLYHHCLEKISKMENEILHVQENAEQLKNKIEKEAEKLEISEKHRGMLEKS 375
+ +E + K E+ E ++E+ +E+ E +E++
Sbjct: 359 KENK-----------IEIVEKEGKELKQENVEMEMKVRELERRIGVIEMKE-----VEEN 490
Query: 376 NQNLQLEAENLVQRIASKDHELLEKHTEIERLQTLMHGE 414
++ +++E E ++ EK E+E L++++ G+
Sbjct: 491 SKRVRIEEE--------MKEKIYEKEMEVEELKSMLMGK 583
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.310 0.128 0.336
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 39,496,703
Number of Sequences: 36976
Number of extensions: 459744
Number of successful extensions: 2773
Number of sequences better than 10.0: 138
Number of HSP's better than 10.0 without gapping: 2629
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 2749
length of query: 1662
length of database: 9,014,727
effective HSP length: 109
effective length of query: 1553
effective length of database: 4,984,343
effective search space: 7740684679
effective search space used: 7740684679
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.8 bits)
S2: 65 (29.6 bits)
Medicago: description of AC135320.2


