

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC135318.3 + phase: 1 /pseudo
(726 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
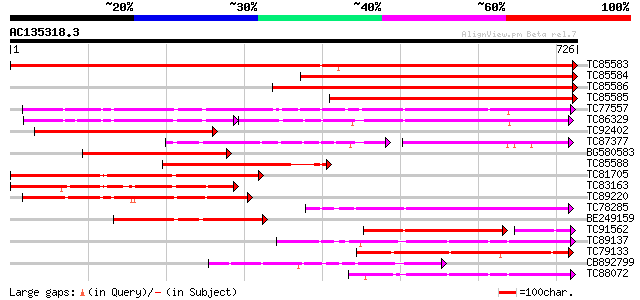
Score E
Sequences producing significant alignments: (bits) Value
TC85583 similar to PIR|T51335|T51335 subtilisin-like proteinase ... 1114 0.0
TC85584 weakly similar to PIR|A84473|A84473 probable serine prot... 704 0.0
TC85586 similar to GP|20521377|dbj|BAB91889. putative subtilisin... 660 0.0
TC85585 similar to GP|22773236|gb|AAN06842.1 Putatvie subtilisin... 519 e-147
TC77557 similar to PIR|JC7519|JC7519 subtilisin-like serine prot... 512 e-145
TC86329 similar to PIR|T07171|T07171 subtilisin-like proteinase ... 273 e-128
TC92402 similar to GP|20198252|gb|AAM15483.1 subtilisin-like ser... 447 e-126
TC87377 weakly similar to PIR|T07171|T07171 subtilisin-like prot... 191 4e-94
BG580583 weakly similar to GP|9757901|dbj serine protease-like p... 337 7e-93
TC85588 similar to GP|20198169|gb|AAM15440.1 subtilisin-like ser... 280 2e-75
TC81705 weakly similar to GP|6721520|dbj|BAA89562.1 EST AU029428... 262 4e-70
TC83163 weakly similar to GP|6721520|dbj|BAA89562.1 EST AU029428... 252 4e-67
TC89220 similar to PIR|T05768|T05768 subtilisin-like proteinase ... 248 4e-66
TC78285 similar to PIR|A84473|A84473 probable serine proteinase ... 244 6e-65
BE249159 weakly similar to GP|20198252|gb subtilisin-like serine... 214 9e-56
TC91562 similar to PIR|T07172|T07172 subtilisin-like proteinase ... 191 1e-55
TC89137 similar to GP|4200338|emb|CAA76726.1 P69C protein {Lycop... 212 4e-55
TC79133 similar to GP|14150446|gb|AAK53065.1 subtilisin-type pro... 212 5e-55
CB892799 weakly similar to PIR|T05768|T05 subtilisin-like protei... 199 2e-51
TC88072 similar to PIR|T06580|T06580 subtilisin-like proteinase ... 198 7e-51
>TC85583 similar to PIR|T51335|T51335 subtilisin-like proteinase AIR3
auxin-induced [imported] - Arabidopsis thaliana
(fragment), partial (40%)
Length = 2691
Score = 1114 bits (2881), Expect = 0.0
Identities = 557/742 (75%), Positives = 618/742 (83%), Gaps = 16/742 (2%)
Frame = +2
Query: 1 DLLGSILGSKENAKEAIIYSYNKQINGFAAMLEEEEAAQIAKNPKVVSVFLSKEHKLHTT 60
DLL S LGS+E AKEAIIYSYNK INGFAA+LE+EEAA IAK VVSVFLSK HKLHTT
Sbjct: 203 DLLSSTLGSREKAKEAIIYSYNKHINGFAALLEDEEAADIAKKRNVVSVFLSKPHKLHTT 382
Query: 61 RSWEFLGLRGNDINSAWQKGRFGENTIIGNIDTGVWPESKSFSDRGIGPIPAKWRGGNIC 120
RSWEFLGLR N N+AWQKG+FGENTII NIDTGVWPESKSF+D+G GP+P+KWRGG C
Sbjct: 383 RSWEFLGLRRNAKNTAWQKGKFGENTIIANIDTGVWPESKSFNDKGYGPVPSKWRGGKAC 562
Query: 121 QLDKLNTSKKVPCNRKLIGARFFNKAYQKRNGKLPRSQQTARDFVGHGTHTLSTAGGNFV 180
++ K + KK PCNRKLIGARFF+ AY+ N KLP Q+TARDF+GHGTHTLSTAGGNFV
Sbjct: 563 EISKFSKYKKNPCNRKLIGARFFSNAYEAYNDKLPSWQRTARDFLGHGTHTLSTAGGNFV 742
Query: 181 PGASIFNIGNGTIKGGSPRARVATYKVCWSLTDATSCFGADVLSAIDQAIDDGVDIISVS 240
P AS+F IGNGT+KGGSPRARVATYKVCWSL D CFGADVL+AIDQAI DGVDIIS+S
Sbjct: 743 PDASVFAIGNGTVKGGSPRARVATYKVCWSLLDLEDCFGADVLAAIDQAISDGVDIISLS 922
Query: 241 AGGPSSTNSEEIFTDEISIGAFHALARNILLVASAGNEGPTPGSVVNVAPWVFTVAASTL 300
G S E+IFTDE+SIGAFHAL+RNILLVASAGNEGPT GSVVNVAPWVFT+AASTL
Sbjct: 923 LAGHSLVYPEDIFTDEVSIGAFHALSRNILLVASAGNEGPTGGSVVNVAPWVFTIAASTL 1102
Query: 301 DRDFSSVMTIGNKTLTGASLFVNLPPNQDFTIVTSTDAKLANATNRDARFCRPRTLDPSK 360
DRDFSS +TIGN+T+ GASLFVNLPPNQ F ++ STD KLANATN DA+FC+P TLDPSK
Sbjct: 1103DRDFSSTITIGNQTIRGASLFVNLPPNQAFPLIVSTDGKLANATNHDAQFCKPGTLDPSK 1282
Query: 361 VNGKIVACDREGKIKSVAEGQEALSAGAKGVILRNQPEINGKTLLSEPHVLSTISYPGN- 419
V GKIV C REG IKSVAEGQEALSAGAKG++L NQP+ GKT L+EPH LS + P +
Sbjct: 1283VKGKIVECIREGNIKSVAEGQEALSAGAKGMLLSNQPK-QGKTTLAEPHTLSCVEVPHHA 1459
Query: 420 ---------------HSRTTGRSLDIIPSDIKSGTKLRMSPAKTLNRRKPAPVMASYSSR 464
S + + S +K+GT ++ S AKTL RKPAPVMAS+SSR
Sbjct: 1460PKPPKPKKSAEQERAGSHAPAFDITSMDSKLKAGTTIKFSGAKTLYGRKPAPVMASFSSR 1639
Query: 465 GPNKVQPSILKPDVTAPGVNILAAYSLFASASNLITDTRRGFPFNVMQGTSMSCPHVAGT 524
GPNK+QPSILKPDVTAPGVNILAAYSL+ASASNL TD R FPFNV+QGTSMSCPHVAG
Sbjct: 1640GPNKIQPSILKPDVTAPGVNILAAYSLYASASNLKTDNRNNFPFNVLQGTSMSCPHVAGI 1819
Query: 525 AGLIKTLHPNWSPAAIKSAIMTTATTRDNTNKPISDAFDKTLANPFAYGSGHIRPNSAMD 584
AGLIKTLHPNWSPAAIKSAIMTTATT DNTN+PI DAF+ LA PF YGSGH++P+ A+D
Sbjct: 1820AGLIKTLHPNWSPAAIKSAIMTTATTLDNTNRPIQDAFENKLAIPFDYGSGHVQPDLAID 1999
Query: 585 PGLVYDLGIKDYLNFLCASGYNQQLISALNFNMTFTCSGTSSIDDLNYPSITLPNLGLNS 644
PGLVYDLGIKDYLNFLCA GYNQQLISALNFN TF CSG+ SI D NYPSITLPNL LN+
Sbjct: 2000PGLVYDLGIKDYLNFLCAYGYNQQLISALNFNGTFICSGSHSITDFNYPSITLPNLKLNA 2179
Query: 645 VTVTRTVTNVGPPSTYFAKVQLAGYKIAVVPSSLNFKKIGEKKTFQVIVQATSVTPRRKY 704
V VTRTVTNVGPP TY AK QL GYKI V+P+SL FKK GEKKTFQVIVQAT+VTPR KY
Sbjct: 2180VNVTRTVTNVGPPGTYSAKAQLLGYKIVVLPNSLTFKKTGEKKTFQVIVQATNVTPRGKY 2359
Query: 705 QFGELRWTNGKHIVRSPVTVRR 726
QFG L+WT+GKHIVRSP+TVRR
Sbjct: 2360QFGNLQWTDGKHIVRSPITVRR 2425
>TC85584 weakly similar to PIR|A84473|A84473 probable serine proteinase
[imported] - Arabidopsis thaliana, partial (19%)
Length = 1241
Score = 704 bits (1816), Expect = 0.0
Identities = 353/354 (99%), Positives = 354/354 (99%)
Frame = +1
Query: 373 KIKSVAEGQEALSAGAKGVILRNQPEINGKTLLSEPHVLSTISYPGNHSRTTGRSLDIIP 432
+IKSVAEGQEALSAGAKGVILRNQPEINGKTLLSEPHVLSTISYPGNHSRTTGRSLDIIP
Sbjct: 1 EIKSVAEGQEALSAGAKGVILRNQPEINGKTLLSEPHVLSTISYPGNHSRTTGRSLDIIP 180
Query: 433 SDIKSGTKLRMSPAKTLNRRKPAPVMASYSSRGPNKVQPSILKPDVTAPGVNILAAYSLF 492
SDIKSGTKLRMSPAKTLNRRKPAPVMASYSSRGPNKVQPSILKPDVTAPGVNILAAYSLF
Sbjct: 181 SDIKSGTKLRMSPAKTLNRRKPAPVMASYSSRGPNKVQPSILKPDVTAPGVNILAAYSLF 360
Query: 493 ASASNLITDTRRGFPFNVMQGTSMSCPHVAGTAGLIKTLHPNWSPAAIKSAIMTTATTRD 552
ASASNLITDTRRGFPFNVMQGTSMSCPHVAGTAGLIKTLHPNWSPAAIKSAIMTTATTRD
Sbjct: 361 ASASNLITDTRRGFPFNVMQGTSMSCPHVAGTAGLIKTLHPNWSPAAIKSAIMTTATTRD 540
Query: 553 NTNKPISDAFDKTLANPFAYGSGHIRPNSAMDPGLVYDLGIKDYLNFLCASGYNQQLISA 612
NTNKPISDAFDKTLANPFAYGSGHIRPNSAMDPGLVYDLGIKDYLNFLCASGYNQQLISA
Sbjct: 541 NTNKPISDAFDKTLANPFAYGSGHIRPNSAMDPGLVYDLGIKDYLNFLCASGYNQQLISA 720
Query: 613 LNFNMTFTCSGTSSIDDLNYPSITLPNLGLNSVTVTRTVTNVGPPSTYFAKVQLAGYKIA 672
LNFNMTFTCSGTSSIDDLNYPSITLPNLGLNSVTVTRTVTNVGPPSTYFAKVQLAGYKIA
Sbjct: 721 LNFNMTFTCSGTSSIDDLNYPSITLPNLGLNSVTVTRTVTNVGPPSTYFAKVQLAGYKIA 900
Query: 673 VVPSSLNFKKIGEKKTFQVIVQATSVTPRRKYQFGELRWTNGKHIVRSPVTVRR 726
VVPSSLNFKKIGEKKTFQVIVQATSVTPRRKYQFGELRWTNGKHIVRSPVTVRR
Sbjct: 901 VVPSSLNFKKIGEKKTFQVIVQATSVTPRRKYQFGELRWTNGKHIVRSPVTVRR 1062
>TC85586 similar to GP|20521377|dbj|BAB91889. putative subtilisin-like
protease {Oryza sativa (japonica cultivar-group)},
partial (17%)
Length = 1460
Score = 660 bits (1703), Expect = 0.0
Identities = 335/392 (85%), Positives = 356/392 (90%), Gaps = 2/392 (0%)
Frame = +1
Query: 337 DAKLANATNRDARFCRPRTLDPSKVNGKIVACDREGKIKSVAEGQEALSAGAKGVILRNQ 396
DAK +NAT RDARFCRPRTLDPSKV GKIVAC REGKIKSVAEGQEALSAGAKG+ L NQ
Sbjct: 4 DAKFSNATTRDARFCRPRTLDPSKVKGKIVACAREGKIKSVAEGQEALSAGAKGMFLENQ 183
Query: 397 PEINGKTLLSEPHVLSTISYPGNHSRTTGRSLDIIPSD-IKSGTKLRMSPAKTLNRRKPA 455
P+++G TLLSEPHVLST+ G + T L + +D I+SGTK+R S A TL RKPA
Sbjct: 184 PKVSGNTLLSEPHVLSTVGGNGQAAITAPPRLGVTATDTIESGTKIRFSQAITLIGRKPA 363
Query: 456 PVMASYSSRGPNKVQPSILKPDVTAPGVNILAAYSLFASASNLITDTRRGFPFNVMQGTS 515
PVMAS+SSRGPN+VQP ILKPDVTAPGVNILAAYSLFASASNL+TD RRGFPFNVMQGTS
Sbjct: 364 PVMASFSSRGPNQVQPYILKPDVTAPGVNILAAYSLFASASNLLTDNRRGFPFNVMQGTS 543
Query: 516 MSCPHVAGTAGLIKTLHPNWSPAAIKSAIMTTATTRDNTNKPISDAFDKTLANPFAYGSG 575
MSCPHVAGTAGLIKTLHPNWSPAAIKSAIMTTATTRDNTNKPISDAFDKTLA+PFAYGSG
Sbjct: 544 MSCPHVAGTAGLIKTLHPNWSPAAIKSAIMTTATTRDNTNKPISDAFDKTLADPFAYGSG 723
Query: 576 -HIRPNSAMDPGLVYDLGIKDYLNFLCASGYNQQLISALNFNMTFTCSGTSSIDDLNYPS 634
+ P SA+DPGLVYDLGIKDYLNFLCASGYN+QLISALNFNMTFTCSGT SIDDLNYPS
Sbjct: 724 TYSHPTSAIDPGLVYDLGIKDYLNFLCASGYNKQLISALNFNMTFTCSGTHSIDDLNYPS 903
Query: 635 ITLPNLGLNSVTVTRTVTNVGPPSTYFAKVQLAGYKIAVVPSSLNFKKIGEKKTFQVIVQ 694
ITLPNLGLN++TVTRTVTNVGPPSTYFAKVQL GYKIAVVPSSLNFKKIGEKKTFQVIVQ
Sbjct: 904 ITLPNLGLNAITVTRTVTNVGPPSTYFAKVQLPGYKIAVVPSSLNFKKIGEKKTFQVIVQ 1083
Query: 695 ATSVTPRRKYQFGELRWTNGKHIVRSPVTVRR 726
ATS PRRKYQFGELRWTNGKHIVRSPVTV+R
Sbjct: 1084ATSEIPRRKYQFGELRWTNGKHIVRSPVTVQR 1179
>TC85585 similar to GP|22773236|gb|AAN06842.1 Putatvie subtilisin-like
serine protease {Oryza sativa (japonica
cultivar-group)}, partial (21%)
Length = 1353
Score = 519 bits (1337), Expect = e-147
Identities = 255/317 (80%), Positives = 280/317 (87%)
Frame = +2
Query: 410 VLSTISYPGNHSRTTGRSLDIIPSDIKSGTKLRMSPAKTLNRRKPAPVMASYSSRGPNKV 469
V+STI+Y S TT + +I P DIK+ +RMSPA LN RKPAPVMAS+SSRGPNKV
Sbjct: 5 VVSTINYHDPRSITTPKGSEITPEDIKTNATIRMSPANALNGRKPAPVMASFSSRGPNKV 184
Query: 470 QPSILKPDVTAPGVNILAAYSLFASASNLITDTRRGFPFNVMQGTSMSCPHVAGTAGLIK 529
QP ILKPDVTAPGVNILAAYSL AS SNL+TD RRGFPFN+ QGTSMSCPHV GTAGLIK
Sbjct: 185 QPYILKPDVTAPGVNILAAYSLLASVSNLVTDNRRGFPFNIQQGTSMSCPHVVGTAGLIK 364
Query: 530 TLHPNWSPAAIKSAIMTTATTRDNTNKPISDAFDKTLANPFAYGSGHIRPNSAMDPGLVY 589
TLHPNWSPAAIKSAIMTTATTRDNTN+PI DAF+ T AN FAYGSGHI+PNSA+DPGLVY
Sbjct: 365 TLHPNWSPAAIKSAIMTTATTRDNTNEPIEDAFENTTANAFAYGSGHIQPNSAIDPGLVY 544
Query: 590 DLGIKDYLNFLCASGYNQQLISALNFNMTFTCSGTSSIDDLNYPSITLPNLGLNSVTVTR 649
DLGIKDYLNFLCA+GYNQ+LIS+L FNMTFTC GT SI+DLNYPSITLPNLGLN+VTVTR
Sbjct: 545 DLGIKDYLNFLCAAGYNQKLISSLIFNMTFTCYGTQSINDLNYPSITLPNLGLNAVTVTR 724
Query: 650 TVTNVGPPSTYFAKVQLAGYKIAVVPSSLNFKKIGEKKTFQVIVQATSVTPRRKYQFGEL 709
TVTNVGP STY AK QL GYKI VVPSSL FKKIGEKKTF+V VQATSVTP+ KY+FGEL
Sbjct: 725 TVTNVGPRSTYTAKAQLPGYKIVVVPSSLKFKKIGEKKTFKVTVQATSVTPQGKYEFGEL 904
Query: 710 RWTNGKHIVRSPVTVRR 726
+W+NGKHIVRSP+T+RR
Sbjct: 905 QWSNGKHIVRSPITLRR 955
Score = 35.8 bits (81), Expect = 0.060
Identities = 14/18 (77%), Positives = 16/18 (88%)
Frame = +1
Query: 703 KYQFGELRWTNGKHIVRS 720
KY+FGEL WTNGKH V+S
Sbjct: 958 KYEFGEL*WTNGKHCVKS 1011
>TC77557 similar to PIR|JC7519|JC7519 subtilisin-like serine proteinase (EC
3.4.21.-) - Arabidopsis thaliana, partial (88%)
Length = 2682
Score = 512 bits (1318), Expect = e-145
Identities = 304/719 (42%), Positives = 420/719 (58%), Gaps = 11/719 (1%)
Frame = +3
Query: 17 IIYSYNKQINGFAAMLEEEEAAQIAKNPKVVSVFLSKEHKLHTTRSWEFLGLRGNDINSA 76
++Y+Y I+GF+ L EEA + +++V +++LHTTR+ +FLGL +
Sbjct: 429 MMYTYENAIHGFSTRLTPEEARLLESQTGILAVLPEVKYELHTTRTPQFLGL--DKSADM 602
Query: 77 WQKGRFGENTIIGNIDTGVWPESKSFSDRGIGPIPAKWRGGNICQLDKLNTSKKVPCNRK 136
+ + G ++G +DTGVWPESKSF+D G GPIP W+G C+ T+ CN+K
Sbjct: 603 FPESSSGNEVVVGVLDTGVWPESKSFNDAGFGPIPTTWKGA--CESGTNFTAAN--CNKK 770
Query: 137 LIGARFFNKAYQKRNGKLPRSQQTA--RDFVGHGTHTLSTAGGNFVPGASIFNIGNGTIK 194
LIGARFF+K + G + + ++ RD GHGTHT STA G+ VP AS+F +GT +
Sbjct: 771 LIGARFFSKGVEAMLGPIDETTESKSPRDDDGHGTHTSSTAAGSVVPDASLFGYASGTAR 950
Query: 195 GGSPRARVATYKVCWSLTDATSCFGADVLSAIDQAIDDGVDIISVSAGGPSSTNSEEIFT 254
G + RARVA YKVCW CF +D+L+AID+AI D V+++S+S GG S + F
Sbjct: 951 GMATRARVAVYKVCWK----GGCFSSDILAAIDKAISDNVNVLSLSLGGGMS----DYFR 1106
Query: 255 DEISIGAFHALARNILLVASAGNEGPTPGSVVNVAPWVFTVAASTLDRDFSSVMTIGNK- 313
D ++IGAF A+ + IL+ SAGN GP+ S+ NVAPW+ TV A TLDRDF + +++GN
Sbjct: 1107 DSVAIGAFSAMEKGILVSCSAGNAGPSAYSLSNVAPWITTVGAGTLDRDFPASVSLGNGL 1286
Query: 314 TLTGASLFV-NLPPNQDFTIVTSTDAKLANATNRDARFCRPRTLDPSKVNGKIVACDREG 372
+G SL+ N P ++ + +A NATN C TL P V GKIV CDR G
Sbjct: 1287 NYSGVSLYRGNALPESPLPLIYAGNA--TNATN--GNLCMTGTLSPELVAGKIVLCDR-G 1451
Query: 373 KIKSVAEGQEALSAGAKGVILRNQPEINGKTLLSEPHVLSTISYPGNHSRTTGRSLDIIP 432
V +G +AG G++L N NG+ L+++ H+L + + L
Sbjct: 1452 MNARVQKGAVVKAAGGLGMVLSNTAA-NGEELVADTHLLPATAVGEREGNAIKKYLF--- 1619
Query: 433 SDIKSGTKLRMSPAKTLNRRKPAPVMASYSSRGPNKVQPSILKPDVTAPGVNILAAYSLF 492
S+ K K+ K +P+PV+A++SSRGPN + P ILKPD+ APGVNILA +S
Sbjct: 1620 SEAKPTVKIVFQGTKV--GVEPSPVVAAFSSRGPNSITPQILKPDLIAPGVNILAGWSKA 1793
Query: 493 ASASNLITDTRRGFPFNVMQGTSMSCPHVAGTAGLIKTLHPNWSPAAIKSAIMTTATTRD 552
+ L D RR FN++ GTSMSCPHV+G A LIK+ HP+WSPAA++SA+MTTA
Sbjct: 1794 VGPTGLAVDERR-VDFNIISGTSMSCPHVSGLAALIKSAHPDWSPAAVRSALMTTAYIAY 1970
Query: 553 NTNKPISDAFDKTLANPFAYGSGHIRPNSAMDPGLVYDLGIKDYLNFLCASGYNQQLISA 612
+ D+ + PF +GSGH+ P +A++PGLVYDL DYL FLCA Y I++
Sbjct: 1971 KNGNKLQDSATGKSSTPFDHGSGHVDPVAALNPGLVYDLTADDYLGFLCALNYTATQITS 2150
Query: 613 LNFNMTFTCSGTS--SIDDLNYPSITL---PNLGLNSVTVTRTVTNVGPPSTYFAKV--Q 665
L F C S+ DLNYPS + G N V TR TNVGP TY A V
Sbjct: 2151 L-ARRKFQCDAGKKYSVSDLNYPSFAVVFDTMGGANVVKHTRIFTNVGPAGTYKASVTSD 2327
Query: 666 LAGYKIAVVPSSLNFKKIGEKKTFQVIVQATSVTPRRKYQFGELRWTNGKHIVRSPVTV 724
KI V P L+F K EKK+F V ++ TP++ FG L WTNGK++V SP+++
Sbjct: 2328 SKNVKITVEPEELSF-KANEKKSFTVTFTSSGSTPQKLNGFGRLEWTNGKNVVGSPISI 2501
>TC86329 similar to PIR|T07171|T07171 subtilisin-like proteinase (EC 3.4.21.-)
1 - tomato, partial (91%)
Length = 2671
Score = 273 bits (697), Expect(2) = e-128
Identities = 178/448 (39%), Positives = 243/448 (53%), Gaps = 19/448 (4%)
Frame = +3
Query: 294 TVAASTLDRDFSSVMTIGN-KTLTGASLFVN-LPPNQDFTIVTSTDAKLANATNRDARFC 351
TV A T+DRDF + +T+GN G SL+ LPPN +V + + ++ N C
Sbjct: 1086 TVGARTIDRDFPAYITLGNGNRYNGVSLYNGKLPPNSPLPLVYAANVSQDSSDN----LC 1253
Query: 352 RPRTLDPSKVNGKIVACDREGKIKSVAEGQEALSAGAKGVILRNQPEINGKTLLSEPHVL 411
+L PSKV+GKIV CDR G ++ + AG G+IL N + G+ L+++ +L
Sbjct: 1254 STDSLIPSKVSGKIVICDRGGNPRA-EKSLVVKRAGGIGMILANNQDY-GEELVADSFLL 1427
Query: 412 STISYPGNHSRTTGRSLDIIPSDIKS----GTKLRMSPAKTLNRRKPAPVMASYSSRGPN 467
+ S + P+ GT+ + +P+PV+A++SSRGPN
Sbjct: 1428 PAAALGEKASNEIKKYASSAPNPTAKIAFGGTRFGV---------QPSPVVAAFSSRGPN 1580
Query: 468 KVQPSILKPDVTAPGVNILAAYSLFASASNLITDTRRGFPFNVMQGTSMSCPHVAGTAGL 527
+ P ILKPD+ APGVNILA +S + L DTR FN++ GTSMSCPHV+G A L
Sbjct: 1581 ILTPKILKPDLIAPGVNILAGWSGKVGPTGLSVDTRH-VSFNIISGTSMSCPHVSGLAAL 1757
Query: 528 IKTLHPNWSPAAIKSAIMTTATTRDNTNKPISDAFDKTLANPFAYGSGHIRPNSAMDPGL 587
+K HP WSPAAI+SA+MTT+ + I D A P YGSGH+ P +A+DPGL
Sbjct: 1758 LKGAHPEWSPAAIRSALMTTSYGTYKNGQTIKDVATGIPATPLDYGSGHVDPVAALDPGL 1937
Query: 588 VYDLGIKDYLNFLCASGYNQQLISALNFNMTFTCSGTSS--IDDLNYPSITLP------- 638
VYD DYLNFLCA YN I L FTC ++DLNYPS ++P
Sbjct: 1938 VYDATTDDYLNFLCALNYNSFQIK-LVARREFTCDKRIKYRVEDLNYPSFSVPFDTASGR 2114
Query: 639 --NLGLNSVTVTRTVTNVGPPSTYFAKV--QLAGYKIAVVPSSLNFKKIGEKKTFQVIVQ 694
+ + V R +TNVG PSTY V Q KI V P +L+FK++ EKK++ V
Sbjct: 2115 GSSHNPSIVQYKRILTNVGAPSTYKVSVSSQSPLDKIVVEPQTLSFKELNEKKSYTVTFT 2294
Query: 695 ATSVTPRRKYQFGELRWTNGKHIVRSPV 722
+ S+ P F L W++GKH V SP+
Sbjct: 2295 SHSM-PSGTTSFAHLEWSDGKHKVTSPI 2375
Score = 206 bits (524), Expect(2) = e-128
Identities = 118/279 (42%), Positives = 165/279 (58%), Gaps = 4/279 (1%)
Frame = +2
Query: 18 IYSYNKQINGFAAMLEEEEAAQIAKNPKVVSVFLSKEHKLHTTRSWEFLGLRGNDINSAW 77
+Y+Y +GF+ L +EA + K P ++SV ++LHTTR+ EFLGL I
Sbjct: 284 MYTYKHVAHGFSTRLTTQEADLLTKQPGILSVIPDVRYELHTTRTPEFLGLE-KTITLLP 460
Query: 78 QKGRFGENTIIGNIDTGVWPESKSFSDRGIGPIPAKWRGGNICQLDKLNTSKKVPCNRKL 137
G+ E I+G IDTGVWPE KSF D G+GP+P W+G C+ K S CN+KL
Sbjct: 461 SSGKQSE-VIVGVIDTGVWPELKSFDDTGLGPVPKSWKGE--CETGKTFNSSN--CNKKL 625
Query: 138 IGARFFNKAYQKRNGKLPRSQQTA--RDFVGHGTHTLSTAGGNFVPGASIFNIGNGTIKG 195
+GARFF K Y+ G + + ++ RD GHG+HT +TA G+ V GAS+F +GT KG
Sbjct: 626 VGARFFAKGYEAAFGPIDENTESKSPRDDDGHGSHTSTTAAGSAVAGASLFGFASGTAKG 805
Query: 196 GSPRARVATYKVCWSLTDATSCFGADVLSAIDQAIDDGVDIIS--VSAGGPSSTNSEEIF 253
+ +ARVA YKVCW CF +D+ +AID+AID + VS G T + +
Sbjct: 806 MATQARVAAYKVCW----LGGCFTSDIAAAIDKAIDTQARTAAYKVSRLGGGLT---DYY 964
Query: 254 TDEISIGAFHALARNILLVASAGNEGPTPGSVVNVAPWV 292
D +++G F A+ IL+ +SAGN GP+ S+ NVAPW+
Sbjct: 965 KDTVAMGTFAAIEHGILVSSSAGNGGPSKASLANVAPWI 1081
>TC92402 similar to GP|20198252|gb|AAM15483.1 subtilisin-like serine
protease AIR3 {Arabidopsis thaliana}, partial (16%)
Length = 704
Score = 447 bits (1150), Expect = e-126
Identities = 215/234 (91%), Positives = 225/234 (95%)
Frame = +1
Query: 33 EEEEAAQIAKNPKVVSVFLSKEHKLHTTRSWEFLGLRGNDINSAWQKGRFGENTIIGNID 92
EEEEAAQ+AKNPKVVSVFLSKEHKLHTTRSWEFLGL GNDINSAWQKGRFGENTII NID
Sbjct: 1 EEEEAAQLAKNPKVVSVFLSKEHKLHTTRSWEFLGLHGNDINSAWQKGRFGENTIIANID 180
Query: 93 TGVWPESKSFSDRGIGPIPAKWRGGNICQLDKLNTSKKVPCNRKLIGARFFNKAYQKRNG 152
TGVWPES+SFSDRGIGPIPAKWRGGN+CQ++KL SKKVPCNRKLIGARFF+ AY++ NG
Sbjct: 181 TGVWPESRSFSDRGIGPIPAKWRGGNVCQINKLRGSKKVPCNRKLIGARFFSDAYERYNG 360
Query: 153 KLPRSQQTARDFVGHGTHTLSTAGGNFVPGASIFNIGNGTIKGGSPRARVATYKVCWSLT 212
KLP SQ+TARDFVGHGTHTLSTAGGNFVPGASIFNIGNGTIKGGSPRARVATYKVCWSLT
Sbjct: 361 KLPTSQRTARDFVGHGTHTLSTAGGNFVPGASIFNIGNGTIKGGSPRARVATYKVCWSLT 540
Query: 213 DATSCFGADVLSAIDQAIDDGVDIISVSAGGPSSTNSEEIFTDEISIGAFHALA 266
DA SCFGADVLSAIDQAIDDGVDII VSAGGPSSTNSEEIFTDE+SIGAFHALA
Sbjct: 541 DAASCFGADVLSAIDQAIDDGVDIIFVSAGGPSSTNSEEIFTDEVSIGAFHALA 702
>TC87377 weakly similar to PIR|T07171|T07171 subtilisin-like proteinase (EC
3.4.21.-) 1 - tomato, partial (54%)
Length = 2071
Score = 191 bits (484), Expect(2) = 4e-94
Identities = 122/294 (41%), Positives = 177/294 (59%), Gaps = 6/294 (2%)
Frame = +2
Query: 200 ARVATYKVCWSLTDATSCFGADVLSAIDQAIDDGVDIISVSAGGPSSTNSEEIFTDEISI 259
ARVA YKVCW + CF +D+ + +D+AI+DGV+I+S+S GG + + + D I+I
Sbjct: 11 ARVAAYKVCW----LSGCFTSDIAAGMDKAIEDGVNILSMSIGG----SIMDYYRDIIAI 166
Query: 260 GAFHALARNILLVASAGNEGPTPGSVVNVAPWVFTVAASTLDRDFSSVMTIGN-KTLTGA 318
GAF A++ IL+ +SAGN GP+ S+ NVAPW+ TV A T+DRDF S +T+GN KT TGA
Sbjct: 167 GAFTAMSHGILVSSSAGNGGPSAESLSNVAPWITTVGAGTIDRDFPSYITLGNGKTYTGA 346
Query: 319 SLFVNLPPNQDFTIVTSTDAKLANATNRDARFCRPRTLDPSKVNGKIVACDREGKIKSVA 378
SL+ N P+ D + ++ ++ C P +L SKV GKIV C+R G + V
Sbjct: 347 SLY-NGKPSSDSLLPVVYAGNVSESS--VGYLCIPDSLTSSKVLGKIVICERGGNSR-VE 514
Query: 379 EGQEALSAGAKGVILRNQPEINGKTLLSEPHVLSTISYPGNHSRTTGRSLDIIPSD---- 434
+G +AG G+IL N E G+ L+++ H+L + G S T + +
Sbjct: 515 KGLVVKNAGGVGMILVNN-EAYGEELIADSHLLPAAAL-GQKSSTVLKDYVFTTKNPRAK 688
Query: 435 -IKSGTKLRMSPAKTLNRRKPAPVMASYSSRGPNKVQPSILKPDVTAPGVNILA 487
+ GT L++ +P+PV+A++SSRGPN + P ILKPD+ APGVNILA
Sbjct: 689 LVFGGTHLQV---------QPSPVVAAFSSRGPNSLTPKILKPDLIAPGVNILA 823
Score = 173 bits (438), Expect(2) = 4e-94
Identities = 100/237 (42%), Positives = 131/237 (55%), Gaps = 17/237 (7%)
Frame = +1
Query: 503 RRGFPFNVMQGTSMSCPHVAGTAGLIKTLHPNWSPAAIKSAIMTTATTRDNTNKPISDAF 562
+R FN++ GTSMSCPH +G A ++K +P WSPAAI+SA+MTTA T + I D
Sbjct: 868 KRHVNFNIISGTSMSCPHASGLAAIVKGAYPEWSPAAIRSALMTTAYTSYKNGQTIVDVA 1047
Query: 563 DKTLANPFAYGSGHIRPNSAMDPGLVYDLGIKDYLNFLCASGYNQQLISALNFNMTFTCS 622
A PF +GSGH+ P SA+DPGLVYD+ + DYL F CA Y I L FTC
Sbjct: 1048 TGKPATPFDFGSGHVDPVSALDPGLVYDINVDDYLGFFCALNYTSYQIK-LAARREFTCD 1224
Query: 623 GTSS--IDDLNYPSI-----TLPNLGLNS-----VTVTRTVTNVGPPSTYFAKVQL---- 666
++D NYPS T +G S V R +TNVG P TY A V L
Sbjct: 1225 ARKKYRVEDFNYPSFAVALETASGIGGGSNKPIIVEYNRVLTNVGAPGTYNATVVLSSVD 1404
Query: 667 -AGYKIAVVPSSLNFKKIGEKKTFQVIVQATSVTPRRKYQFGELRWTNGKHIVRSPV 722
+ K+ V P +++FK++ EKK ++V S+ P FG L W +GKH V SP+
Sbjct: 1405 SSSVKVVVEPETISFKEVYEKKGYKVRFICGSM-PSGTKSFGYLEWNDGKHKVGSPI 1572
>BG580583 weakly similar to GP|9757901|dbj serine protease-like protein
{Arabidopsis thaliana}, partial (13%)
Length = 592
Score = 337 bits (865), Expect = 7e-93
Identities = 162/190 (85%), Positives = 173/190 (90%)
Frame = +3
Query: 94 GVWPESKSFSDRGIGPIPAKWRGGNICQLDKLNTSKKVPCNRKLIGARFFNKAYQKRNGK 153
GVWPES+SF+DRGIGPIP +WRGGNICQLDKLNTSKKVPCNRKLIGARFFNKAY+ +GK
Sbjct: 21 GVWPESESFNDRGIGPIPLRWRGGNICQLDKLNTSKKVPCNRKLIGARFFNKAYEAFHGK 200
Query: 154 LPRSQQTARDFVGHGTHTLSTAGGNFVPGASIFNIGNGTIKGGSPRARVATYKVCWSLTD 213
LP SQQTARDFVGHGTHTLSTAGGNFV A+IF IGNGTIKGGSPR+RVATYK CWSLTD
Sbjct: 201 LPSSQQTARDFVGHGTHTLSTAGGNFVQNATIFGIGNGTIKGGSPRSRVATYKACWSLTD 380
Query: 214 ATSCFGADVLSAIDQAIDDGVDIISVSAGGPSSTNSEEIFTDEISIGAFHALARNILLVA 273
CFGADVL+AIDQ+I DG D+ISVSAGG +TN E IFTDEISIGAFHALARNILLVA
Sbjct: 381 VVDCFGADVLAAIDQSIYDGADLISVSAGGKPNTNPEVIFTDEISIGAFHALARNILLVA 560
Query: 274 SAGNEGPTPG 283
SAGNEGPTPG
Sbjct: 561 SAGNEGPTPG 590
>TC85588 similar to GP|20198169|gb|AAM15440.1 subtilisin-like serine
protease AIR3 {Arabidopsis thaliana}, partial (7%)
Length = 572
Score = 280 bits (715), Expect = 2e-75
Identities = 146/217 (67%), Positives = 163/217 (74%)
Frame = +1
Query: 196 GSPRARVATYKVCWSLTDATSCFGADVLSAIDQAIDDGVDIISVSAGGPSSTNSEEIFTD 255
GSPRARVATYKVCWSL D CFGADVL+AIDQAI DGVDIIS+S G S E+IFTD
Sbjct: 1 GSPRARVATYKVCWSLLDLEDCFGADVLAAIDQAISDGVDIISLSLAGHSLVYPEDIFTD 180
Query: 256 EISIGAFHALARNILLVASAGNEGPTPGSVVNVAPWVFTVAASTLDRDFSSVMTIGNKTL 315
E+SIGAFHAL+RNILLVASAGNEGPT GSVVNV PWVFT+AASTLDRDFSS +TIGN+T+
Sbjct: 181 EVSIGAFHALSRNILLVASAGNEGPTGGSVVNVTPWVFTIAASTLDRDFSSTITIGNQTI 360
Query: 316 TGASLFVNLPPNQDFTIVTSTDAKLANATNRDARFCRPRTLDPSKVNGKIVACDREGKIK 375
GASLFVNLPPNQ F ++ STD KLANATN DA+FC+P TLDPSKV
Sbjct: 361 RGASLFVNLPPNQAFPLIVSTDGKLANATNHDAQFCKPGTLDPSKV-------------- 498
Query: 376 SVAEGQEALSAGAKGVILRNQPEINGKTLLSEPHVLS 412
KG++L NQP+ GKT L+EPH LS
Sbjct: 499 -------------KGMLLSNQPK-QGKTTLAEPHTLS 567
>TC81705 weakly similar to GP|6721520|dbj|BAA89562.1 EST AU029428(E30359)
corresponds to a region of the predicted gene.~Similar
to antifreeze-like, partial (33%)
Length = 1443
Score = 262 bits (669), Expect = 4e-70
Identities = 150/330 (45%), Positives = 202/330 (60%), Gaps = 6/330 (1%)
Frame = +1
Query: 1 DLLGSILGSKENAKEAIIYSYNKQINGFAAMLEEEEAAQIAKNPKVVSVFLSKEHKLHTT 60
+LL SI+ S+E+ + +I+ YN +GF+AML + EA+ ++ + VVSVF +LHTT
Sbjct: 151 ELLSSIIPSEESERIKLIHHYNHAFSGFSAMLTQSEASALSGHDGVVSVFPDPILELHTT 330
Query: 61 RSWEFLG--LRGNDINSAWQKGRFGENTIIGNIDTGVWPESKSFSDRGIGPIPAKWRGGN 118
RSW+FL L + + II IDTG+WPES SF+D GIG IP+ W+G
Sbjct: 331 RSWDFLDSDLGMKPSTNVLTHQHSSNDIIIALIDTGIWPESPSFTDEGIGKIPSVWKG-- 504
Query: 119 ICQLDKLNTSKKVPCNRKLIGARFFNK--AYQKRNGKLPRSQQTARDFVGHGTHTLSTAG 176
IC + + KK CNRKLIGAR++N + + ++ + RD VGHGTHT STA
Sbjct: 505 ICM--EGHDFKKSNCNRKLIGARYYNTQDTFGSNKTHIGGAKGSPRDTVGHGTHTASTAA 678
Query: 177 GNFVPGASIFNIGNGTIKGGSPRARVATYKVCWSLTDATSCFGADVLSAIDQAIDDGVDI 236
G V A+ + + GT +GGSP R+A YK C C G+ +L A+D AI DGVDI
Sbjct: 679 GVNVNNANYYGLAKGTARGGSPSTRIAAYKTC----SEEGCSGSTILKAMDDAIKDGVDI 846
Query: 237 ISVSAGGPSSTNSEEIFTDEISIGAFHALARNILLVASAGNEGPTPGSVVNVAPWVFTVA 296
IS+S G SS + D I+IGAFHA R + V SAGN+GP P +VVN APW+FTVA
Sbjct: 847 ISISIG-LSSLMQSDYLNDPIAIGAFHAEQRGVTAVCSAGNDGPDPNTVVNTAPWIFTVA 1023
Query: 297 ASTLDRDFSSVMTIGN-KTLTGASL-FVNL 324
AS +DR+F S + +GN K+ GA + F NL
Sbjct: 1024ASNIDRNFQSTIVLGNGKSFQGAGINFSNL 1113
>TC83163 weakly similar to GP|6721520|dbj|BAA89562.1 EST AU029428(E30359)
corresponds to a region of the predicted gene.~Similar
to antifreeze-like, partial (29%)
Length = 990
Score = 252 bits (643), Expect = 4e-67
Identities = 145/297 (48%), Positives = 187/297 (62%), Gaps = 4/297 (1%)
Frame = +2
Query: 1 DLLGSILGSKENAKEAIIYSYNKQINGFAAMLEEEEAAQIAKNPKVVSVFLSKEHKLHTT 60
DLL SI+ S+E+ + A+I+ Y+ NGF+AML + EA+ +A N VVSVF +LHTT
Sbjct: 161 DLLSSIIPSEESERIALIHHYSHAFNGFSAMLTQSEASALAGNDGVVSVFEDPFLELHTT 340
Query: 61 RSWEF----LGLRGNDINSAWQKGRFGENTIIGNIDTGVWPESKSFSDRGIGPIPAKWRG 116
RSW+F LG+R + I + + IIG IDTG+WPES SF D GIG IP++W+G
Sbjct: 341 RSWDFLESDLGMRPHGI---LKHQHSSNDIIIGVIDTGIWPESPSFKDEGIGKIPSRWKG 511
Query: 117 GNICQLDKLNTSKKVPCNRKLIGARFFNKAYQKRNGKLPRSQQTARDFVGHGTHTLSTAG 176
+C + + KK CNRKLIGAR++NK K P+ + RDF GHGTHT STA
Sbjct: 512 --VCM--EAHDFKKSNCNRKLIGARYYNK-------KDPKG--SPRDFNGHGTHTASTAA 652
Query: 177 GNFVPGASIFNIGNGTIKGGSPRARVATYKVCWSLTDATSCFGADVLSAIDQAIDDGVDI 236
G V AS + + GT +GGSP AR+A YK C C G +L AID AI DGVDI
Sbjct: 653 GVIVNNASYYGLAKGTARGGSPSARIAAYKAC----SGEGCSGGTLLKAIDDAIKDGVDI 820
Query: 237 ISVSAGGPSSTNSEEIFTDEISIGAFHALARNILLVASAGNEGPTPGSVVNVAPWVF 293
IS+S G S SE + +D I+IGAFHA R +++V SAGNEGP +VVN PW+F
Sbjct: 821 ISISIGFSSEFLSEYL-SDPIAIGAFHAEQRGVMVVCSAGNEGPDHYTVVNTTPWIF 988
>TC89220 similar to PIR|T05768|T05768 subtilisin-like proteinase (EC
3.4.21.-) - Arabidopsis thaliana, partial (39%)
Length = 1149
Score = 248 bits (634), Expect = 4e-66
Identities = 126/302 (41%), Positives = 195/302 (63%), Gaps = 8/302 (2%)
Frame = +3
Query: 17 IIYSYNKQINGFAAMLEEEEAAQIAKNPKVVSVFLSKEHKLHTTRSWEFLGLRGNDINSA 76
I+++Y+ +GF+A+L ++ A I+ +P +++VF + +LHTTRS +FLGLR
Sbjct: 228 ILHTYDTAFHGFSAVLTRQQVASISNHPSILAVFEDRRRQLHTTRSPQFLGLRNQ--RGL 401
Query: 77 WQKGRFGENTIIGNIDTGVWPESKSFSDRGIGPIPAKWRGGNICQLDKLNTSKKVPCNRK 136
W + +G + I+G DTG+WPE +SFSD +GPIP +W+G +C+ + + + CNRK
Sbjct: 402 WSESDYGSDVIVGVFDTGIWPERRSFSDMNLGPIPRRWKG--VCESGEKFSPRN--CNRK 569
Query: 137 LIGARFFNKAYQKRNGKL----PRSQ----QTARDFVGHGTHTLSTAGGNFVPGASIFNI 188
LIGAR+F+K ++ G P ++ ++ RD GHGTHT STA G + A++
Sbjct: 570 LIGARYFSKGHEVGAGSAGPLNPINETVEFRSPRDADGHGTHTASTAAGRYAFQANMSGY 749
Query: 189 GNGTIKGGSPRARVATYKVCWSLTDATSCFGADVLSAIDQAIDDGVDIISVSAGGPSSTN 248
+G KG +P+AR+A YKVCW + CF +D+L+A D A++DGVD+IS+S GG
Sbjct: 750 ASGIAKGVAPKARLAVYKVCWK---NSGCFDSDILAAFDAAVNDGVDVISISIGGGDGIA 920
Query: 249 SEEIFTDEISIGAFHALARNILLVASAGNEGPTPGSVVNVAPWVFTVAASTLDRDFSSVM 308
S + D I+IG++ A++R + + +SAGN+GP+ SV N+APW+ TV A T+DRDF S +
Sbjct: 921 SP-YYLDPIAIGSYGAVSRGVFVSSSAGNDGPSGMSVTNLAPWLTTVGAGTIDRDFPSQI 1097
Query: 309 TI 310
I
Sbjct: 1098II 1103
>TC78285 similar to PIR|A84473|A84473 probable serine proteinase [imported]
- Arabidopsis thaliana, partial (41%)
Length = 1353
Score = 244 bits (624), Expect = 6e-65
Identities = 143/350 (40%), Positives = 202/350 (56%), Gaps = 6/350 (1%)
Frame = +1
Query: 379 EGQEALSAGAKGVILRNQPEINGKTLLSEPHVLSTISYPGNHSRTTGRSL-DIIPSDIKS 437
+G+ AG G+IL N +G+ L+++ H+L ++ R G + + SD+
Sbjct: 13 KGRVVKEAGGIGMILANTAA-SGEELVADSHLLPAVAV----GRIIGDQIRKYVSSDLNP 177
Query: 438 GTKLRMSPAKTLNRRKPAPVMASYSSRGPNKVQPSILKPDVTAPGVNILAAYSLFASASN 497
T L S T+ +P+PV+A++SSRGPN + ILKPDV PGVNILA +S S
Sbjct: 178 TTVL--SFGGTVLNVRPSPVVAAFSSRGPNMITKEILKPDVIGPGVNILAGWSEAVGPSG 351
Query: 498 LITDTRRGFPFNVMQGTSMSCPHVAGTAGLIKTLHPNWSPAAIKSAIMTTATTRDNTNKP 557
L DTR+ FN+M GTSMSCPH++G A L+K HP WSP+AIKSA+MTTA DN+ P
Sbjct: 352 LAEDTRKT-KFNIMSGTSMSCPHISGLAALLKAAHPTWSPSAIKSALMTTAYNHDNSKSP 528
Query: 558 ISDAFDKTLANPFAYGSGHIRPNSAMDPGLVYDLGIKDYLNFLCASGYNQQLISALNFNM 617
+ DA D + + P A+G+GH+ P A+ PGLVYD KDY+ FLC+ YN + I +
Sbjct: 529 LRDAADGSFSTPLAHGAGHVNPQKALSPGLVYDASTKDYITFLCSLNYNSEQIQLIVKRP 708
Query: 618 TFTCSGT-SSIDDLNYPSITLPNLGLNSVTVTRTVTNVGPP-STYFAKVQL-AGYKIAVV 674
+ C+ ++ LNYPS ++ V TR VTNVG S Y V + + I V
Sbjct: 709 SVNCTKKFANPGQLNYPSFSVVFSSKRVVRYTRIVTNVGEAGSVYNVVVDVPSSVGITVK 888
Query: 675 PSSLNFKKIGEKKTFQVIVQATSVTPRRKYQ--FGELRWTNGKHIVRSPV 722
PS L F+K+GE+K + V + K + FG + W+N +H VRSP+
Sbjct: 889 PSRLVFEKVGERKRYTVTFVSKKGADASKVRSGFGSILWSNAQHQVRSPI 1038
>BE249159 weakly similar to GP|20198252|gb subtilisin-like serine protease
AIR3 {Arabidopsis thaliana}, partial (24%)
Length = 661
Score = 214 bits (545), Expect = 9e-56
Identities = 114/198 (57%), Positives = 137/198 (68%), Gaps = 1/198 (0%)
Frame = +1
Query: 134 NRKLIGARFFNKAYQKRNGKLPRSQQTARDFVGHGTHTLSTAGGNFVPGASIFNIGNGTI 193
+RKLIGAR F K Y+ GKL S TARD +GHG+HTLSTAGGNFV G S++ GNGT
Sbjct: 82 SRKLIGARAFYKGYEAYVGKLDASFYTARDTIGHGSHTLSTAGGNFVQGVSVYGNGNGTA 261
Query: 194 KGGSPRARVATYKVCWSLTDATSCFGADVLSAIDQAIDDGVDIISVSAGGPSSTNSEEIF 253
KGGSP+A VA YKVCW C ADVL+ + AI DGVD++SVS G + +F
Sbjct: 262 KGGSPKAHVAAYKVCWK----GGCSDADVLAGFEAAISDGVDVLSVSLG----MKTHNLF 417
Query: 254 TDEISIGAFHALARNILLVASAGNEGPTPGSVVNVAPWVFTVAASTLDRDFSSVMTIG-N 312
TD ISIG+FHA+A I++VASAGN GP G+V NVAPW+FTVAAST+DRDF+S +T+G N
Sbjct: 418 TDSISIGSFHAVANGIVVVASAGNSGPYFGTVSNVAPWLFTVAASTIDRDFASYVTLGDN 597
Query: 313 KTLTGASLFVNLPPNQDF 330
K G SL P F
Sbjct: 598 KHFKGTSLSSKDLPTHKF 651
>TC91562 similar to PIR|T07172|T07172 subtilisin-like proteinase (EC
3.4.21.-) 2 - tomato, partial (36%)
Length = 1062
Score = 191 bits (484), Expect(2) = 1e-55
Identities = 94/186 (50%), Positives = 125/186 (66%), Gaps = 1/186 (0%)
Frame = +2
Query: 453 KPAPVMASYSSRGPNKVQPSILKPDVTAPGVNILAAYSLFASASNLITDTRRGFPFNVMQ 512
+P+P++A++SSRGP+ + ILKPD+ APGVNILAA+S S+L D RR FN++
Sbjct: 44 RPSPIVAAFSSRGPSLLTLEILKPDIVAPGVNILAAWSGLTGPSSLPIDHRR-VKFNILS 220
Query: 513 GTSMSCPHVAGTAGLIKTLHPNWSPAAIKSAIMTTATTRDNTNKPISDAFDKTLANPFAY 572
GTSMSCPHV+G A +IK HP WSPAAIKSAIMTTA DNT KP+ DA + P+ +
Sbjct: 221 GTSMSCPHVSGIAAMIKAKHPEWSPAAIKSAIMTTAYVHDNTIKPLRDASSAEFSTPYDH 400
Query: 573 GSGHIRPNSAMDPGLVYDLGIKDYLNFLCASGYNQQLISALNFNMTFTCSGT-SSIDDLN 631
G+GHI P A+DPGL+YD+ +DY FLC + + + N C T +S DLN
Sbjct: 401 GAGHINPRKALDPGLLYDIEPQDYFEFLCTKKLSPSELVVFSKNSNRNCKHTLASASDLN 580
Query: 632 YPSITL 637
YP+I++
Sbjct: 581 YPAISV 598
Score = 45.1 bits (105), Expect(2) = 1e-55
Identities = 26/80 (32%), Positives = 45/80 (55%), Gaps = 2/80 (2%)
Frame = +1
Query: 647 VTRTVTNVGPPSTYFAKV--QLAGYKIAVVPSSLNFKKIGEKKTFQVIVQATSVTPRRKY 704
+ RTVTNVGP + + + G + V P +LNF + +K ++++ + TS + +
Sbjct: 634 IHRTVTNVGPAVSKYHVIGTPFKGAVVKVEPDTLNFTRKYQKLSYKISFKVTS--RQSEP 807
Query: 705 QFGELRWTNGKHIVRSPVTV 724
+FG L W + H VRSP+ +
Sbjct: 808 EFGGLVWKDRLHKVRSPIVI 867
>TC89137 similar to GP|4200338|emb|CAA76726.1 P69C protein {Lycopersicon
esculentum}, partial (37%)
Length = 1230
Score = 212 bits (540), Expect = 4e-55
Identities = 144/393 (36%), Positives = 206/393 (51%), Gaps = 10/393 (2%)
Frame = +3
Query: 342 NATNRDARFCRPRTLDPSKVNGKIVACDREGKIKSVAEGQEALSAGAKGVILRNQPEING 401
N ++ FC P + V GKIV C++ G + VA+GQ AG +IL N E
Sbjct: 21 NTSDDSIAFCGPIAMKKVDVKGKIVVCEQGGFVGRVAKGQAVKDAGGAAMILLNS-EGED 197
Query: 402 KTLLSEPHVLSTISYPGNHSRTTGRSLDIIPSDIKSGTKLRMSPAK---TLNRRKPAPVM 458
+++ HVL + S + G ++ D + T M+ T+ AP +
Sbjct: 198 FNPIADVHVLPAVHV----SYSAGLNIQ----DYINSTSTPMATILFKGTVIGNPNAPQV 353
Query: 459 ASYSSRGPNKVQPSILKPDVTAPGVNILAAYSLFASASNLITDTRRGFPFNVMQGTSMSC 518
AS+SSRGP+K P ILKPD+ PG+NILA + + S FN++ GTSMSC
Sbjct: 354 ASFSSRGPSKASPGILKPDILGPGLNILAGWPISLDNSTS--------SFNIIAGTSMSC 509
Query: 519 PHVAGTAGLIKTLHPNWSPAAIKSAIMTTATTRDNTNKPISDAFDKTL-ANPFAYGSGHI 577
PH++G A L+K HP+WSPAAIKSAIMTTA + KPI D + L A+ FA G+GH+
Sbjct: 510 PHLSGIAALLKNSHPDWSPAAIKSAIMTTANHVNLHGKPILD--QRLLPADVFATGAGHV 683
Query: 578 RPNSAMDPGLVYDLGIKDYLNFLCASGYNQQLISALNFNMTFTCSGTSSID--DLNYPSI 635
P+ A DPGLVYD+ DY+ +LC Y + + CS SI LNYPSI
Sbjct: 684 NPSKANDPGLVYDIETNDYVPYLCGLNYTDIQVGII-LQQKVKCSDVKSIPQAQLNYPSI 860
Query: 636 TLPNLGLNSVTVTRTVTNVGPPSTYFAKV--QLAGYKIAVVPSSLNFKKIGEKKTFQV-- 691
++ LG S +RT+TNVGP +T + V +++V PS + F ++ +K T+ V
Sbjct: 861 SI-RLGNTSQFYSRTLTNVGPVNTTYNVVIDVPVAVRMSVRPSQITFTEVKQKVTYWVDF 1037
Query: 692 IVQATSVTPRRKYQFGELRWTNGKHIVRSPVTV 724
I + G ++W + K+ V P+ V
Sbjct: 1038IPEDKENRGDNFIAQGSIKWISAKYSVSIPIAV 1136
>TC79133 similar to GP|14150446|gb|AAK53065.1 subtilisin-type protease
precursor {Glycine max}, partial (42%)
Length = 1101
Score = 212 bits (539), Expect = 5e-55
Identities = 121/286 (42%), Positives = 176/286 (61%), Gaps = 9/286 (3%)
Frame = +1
Query: 445 PAKTLNRRKPAPVMASYSSRGPNKVQPSILKPDVTAPGVNILAAYSLFASASNLITDTRR 504
P T+ KPAP++A +SSRGP+ + +ILKPD+ APGV ILAA+ + + ++
Sbjct: 121 PTVTVIDYKPAPMVAIFSSRGPSALSKNILKPDIAAPGVTILAAW--IGNDDENVPKGKK 294
Query: 505 GFPFNVMQGTSMSCPHVAGTAGLIKTLHPNWSPAAIKSAIMTTATTRDNTNKPISDAFDK 564
P+ + GTSMSCPHV+G AG IK+ +P WS +AI+SAIMT+AT +N PI+
Sbjct: 295 PLPYKLETGTSMSCPHVSGLAGSIKSRNPTWSASAIRSAIMTSATQINNMKAPITTDLG- 471
Query: 565 TLANPFAYGSGHIRPNSAMDPGLVYDLGIKDYLNFLCASGYNQQLISALNFNM--TFTCS 622
++A P+ YG+G I + PGLVY+ DYLN+LC GYN I ++ + TF C
Sbjct: 472 SVATPYDYGAGDITTIESFQPGLVYETSTIDYLNYLCYIGYNTTTIKVISKTVPDTFNCP 651
Query: 623 GTSS---IDDLNYPSITLPNL-GLNSVTVTRTVTNVGPPS--TYFAKVQL-AGYKIAVVP 675
S+ I ++NYPSI + N G +V V+RTVTNVG Y A V +G K+ ++P
Sbjct: 652 KESTPDHISNINYPSIAISNFTGKETVNVSRTVTNVGEEDEVAYSAIVNAPSGVKVQLIP 831
Query: 676 SSLNFKKIGEKKTFQVIVQATSVTPRRKYQFGELRWTNGKHIVRSP 721
L F K +K+++Q I +T++T ++ FG + W+NGK+ VRSP
Sbjct: 832 EKLQFTKSNKKQSYQAIF-STTLTSLKEDLFGSITWSNGKYSVRSP 966
>CB892799 weakly similar to PIR|T05768|T05 subtilisin-like proteinase (EC
3.4.21.-) - Arabidopsis thaliana, partial (20%)
Length = 875
Score = 199 bits (507), Expect = 2e-51
Identities = 126/309 (40%), Positives = 186/309 (59%), Gaps = 4/309 (1%)
Frame = +2
Query: 255 DEISIGAFHALARNILLVASAGNEGPTPGSVVNVAPWVFTVAASTLDRDFSSVMTIGNKT 314
D ISIG+FHA R + +V+SAGNEG GS N+APW+ TVAA + DRDF+S + +GN
Sbjct: 2 DAISIGSFHAANRGLFVVSSAGNEGNL-GSATNLAPWMLTVAAGSTDRDFTSDIILGNGA 178
Query: 315 -LTGASLFVNLPPNQDFTIVTSTDAKLANATNRDARFCRPRTLDPSKVNGKIVAC---DR 370
+TG SL + N I+++++A T + +C +L+ +K GK++ C +R
Sbjct: 179 KITGESLSL-FEMNASTRIISASEAFAGYFTPYQSSYCLESSLNKTKTKGKVLVCRHVER 355
Query: 371 EGKIKSVAEGQEALSAGAKGVILRNQPEINGKTLLSEPHVLSTISYPGNHSRTTGRSLDI 430
+ K VA+ + AG G+IL ++ + + ++ P V+ + + + L
Sbjct: 356 STESK-VAKSKIVKEAGGVGMILIDETDQD----VAIPFVIPSAIVGKKKGQ---KILSY 511
Query: 431 IPSDIKSGTKLRMSPAKTLNRRKPAPVMASYSSRGPNKVQPSILKPDVTAPGVNILAAYS 490
+ + K +K+ AKT+ + AP +A++SSRGPN + P ILKPD+TAPG+NILAA+S
Sbjct: 512 LKTTRKPMSKILR--AKTVIGAQSAPRVAAFSSRGPNALNPEILKPDITAPGLNILAAWS 685
Query: 491 LFASASNLITDTRRGFPFNVMQGTSMSCPHVAGTAGLIKTLHPNWSPAAIKSAIMTTATT 550
A G FN++ GTSM+CPHV G A L+K +HP+WSP+AIKSAIMTTAT
Sbjct: 686 PVA-----------GNMFNILSGTSMACPHVTGIATLVKAVHPSWSPSAIKSAIMTTATI 832
Query: 551 RDNTNKPIS 559
D +KPIS
Sbjct: 833 LDKRHKPIS 859
>TC88072 similar to PIR|T06580|T06580 subtilisin-like proteinase (EC
3.4.21.-) p69f - tomato, partial (27%)
Length = 1097
Score = 198 bits (503), Expect = 7e-51
Identities = 130/303 (42%), Positives = 178/303 (57%), Gaps = 13/303 (4%)
Frame = +2
Query: 435 IKSGTKLRMSPAKTLNRRKP------APVMASYSSRGPNKVQPSILKPDVTAPGVNILAA 488
IKS K +P TL + AP + +SSRGP++ P ILKPD+ PGVNILAA
Sbjct: 59 IKSYIKSTYNPTATLIFKGTIIGDSLAPSVVYFSSRGPSQESPGILKPDIIGPGVNILAA 238
Query: 489 YSLFASASNLITDTRRGFPFNVMQGTSMSCPHVAGTAGLIKTLHPNWSPAAIKSAIMTTA 548
+++ S N I F+++ GTSMSCPH++G A LIK+ HP+WSPAAIKSAIMTTA
Sbjct: 239 WAV--SVDNKIP------AFDIVSGTSMSCPHLSGIAALIKSSHPDWSPAAIKSAIMTTA 394
Query: 549 TTRDNTNKPISDAFDKTL-ANPFAYGSGHIRPNSAMDPGLVYDLGIKDYLNFLCASGYNQ 607
T + PI D + L A+ FA G+GH+ P A DPGLVYD+ +DY+ +LC GY+
Sbjct: 395 NTLNLGGIPILD--QRLLPADIFATGAGHVNPFKANDPGLVYDIEPEDYVPYLCGLGYSD 568
Query: 608 QLISALNFNMTFTCSGTSSIDD--LNYPSITLPNLGLNSVTVTRTVTNVG-PPSTYFAKV 664
+ I + CS SI + LNYPS ++ LG +S TRT+TNVG STY ++
Sbjct: 569 KEIEVI-VQWKVKCSNVKSIPEAQLNYPSFSI-LLGSDSQYYTRTLTNVGFANSTYKVEL 742
Query: 665 QL-AGYKIAVVPSSLNFKKIGEKKTFQVIVQATSVTPRRKYQFGE--LRWTNGKHIVRSP 721
++ ++V PS + F ++ EK +F V RR FG+ L W + KH VR P
Sbjct: 743 EVPLALGMSVNPSEITFTEVNEKVSFSVEFIPQIKENRRNQTFGQGSLTWVSDKHAVRVP 922
Query: 722 VTV 724
++V
Sbjct: 923 ISV 931
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.316 0.132 0.387
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 20,152,596
Number of Sequences: 36976
Number of extensions: 265830
Number of successful extensions: 1466
Number of sequences better than 10.0: 104
Number of HSP's better than 10.0 without gapping: 1302
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1315
length of query: 726
length of database: 9,014,727
effective HSP length: 103
effective length of query: 623
effective length of database: 5,206,199
effective search space: 3243461977
effective search space used: 3243461977
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 62 (28.5 bits)
Medicago: description of AC135318.3


