

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC135317.6 + phase: 0
(432 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
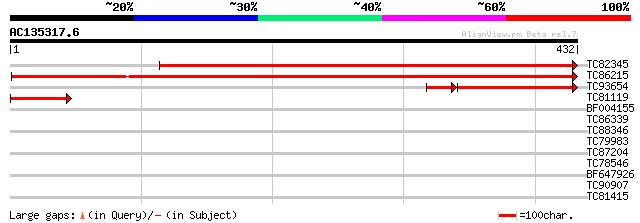
Sequences producing significant alignments: (bits) Value
TC82345 weakly similar to GP|19698867|gb|AAL91169.1 unknown prot... 627 e-180
TC86215 similar to GP|19698867|gb|AAL91169.1 unknown protein {Ar... 427 e-120
TC93654 weakly similar to GP|19571035|dbj|BAB86462. rac-GTP bind... 103 1e-28
TC81119 weakly similar to GP|19571035|dbj|BAB86462. rac-GTP bind... 50 1e-06
BF004155 weakly similar to GP|16648803|gb| AT3g63150/T20O10_250 ... 39 0.004
TC86339 small G-protein ROP6 [Medicago truncatula] 38 0.009
TC88346 homologue to SP|Q40220|RAC2_LOTJA RAC-like GTP binding p... 37 0.015
TC79983 homologue to SP|Q41253|RACD_GOSHI RAC-like GTP binding p... 36 0.025
TC87204 small G-protein ROP9 [Medicago truncatula] 36 0.025
TC78546 small G-protein ROP3 [Medicago truncatula] 35 0.073
BF647926 homologue to PIR|T06679|T066 GTP-binding protein Arac8 ... 34 0.13
TC90907 homologue to PIR|T08950|T08950 GTP binding protein Arac7... 32 0.48
TC81415 weakly similar to PIR|A84509|A84509 probable ABC transpo... 28 5.3
>TC82345 weakly similar to GP|19698867|gb|AAL91169.1 unknown protein
{Arabidopsis thaliana}, partial (27%)
Length = 1237
Score = 627 bits (1618), Expect = e-180
Identities = 317/318 (99%), Positives = 317/318 (99%)
Frame = +1
Query: 115 RLVDTDKDQLLRPAEVDKLFDAAPESPWKDAPYMDAAETTDMGYISLKGFLSRWALMTLL 174
RLVDTDKD LLRPAEVDKLFDAAPESPWKDAPYMDAAETTDMGYISLKGFLSRWALMTLL
Sbjct: 1 RLVDTDKDYLLRPAEVDKLFDAAPESPWKDAPYMDAAETTDMGYISLKGFLSRWALMTLL 180
Query: 175 DPRYSLANLICIGYRDNPSAALRLTSRRSEDRKNQKTERNVFQCYVFGSKTAGKSALLRA 234
DPRYSLANLICIGYRDNPSAALRLTSRRSEDRKNQKTERNVFQCYVFGSKTAGKSALLRA
Sbjct: 181 DPRYSLANLICIGYRDNPSAALRLTSRRSEDRKNQKTERNVFQCYVFGSKTAGKSALLRA 360
Query: 235 LLGRPFSNDYTPTTVERYAANIIELIAGTKKTLILREIPEDEVSNFLSNKDCLAACDVAA 294
LLGRPFSNDYTPTTVERYAANIIELIAGTKKTLILREIPEDEVSNFLSNKDCLAACDVAA
Sbjct: 361 LLGRPFSNDYTPTTVERYAANIIELIAGTKKTLILREIPEDEVSNFLSNKDCLAACDVAA 540
Query: 295 FVHDSSDGYSWKRSIDLLEKVVNQGELTGHRFPSLLIAAKDDLTPFPRAVLDSVKVAQEL 354
FVHDSSDGYSWKRSIDLLEKVVNQGELTGHRFPSLLIAAKDDLTPFPRAVLDSVKVAQEL
Sbjct: 541 FVHDSSDGYSWKRSIDLLEKVVNQGELTGHRFPSLLIAAKDDLTPFPRAVLDSVKVAQEL 720
Query: 355 KIDAPIRLSIKSDDSNNVYSKIINAAEHPHLSIPETEFVRKRKQHQQLLHTFIFALAGAA 414
KIDAPIRLSIKSDDSNNVYSKIINAAEHPHLSIPETEFVRKRKQHQQLLHTFIFALAGAA
Sbjct: 721 KIDAPIRLSIKSDDSNNVYSKIINAAEHPHLSIPETEFVRKRKQHQQLLHTFIFALAGAA 900
Query: 415 MALAGFTARRARANRNSS 432
MALAGFTARRARANRNSS
Sbjct: 901 MALAGFTARRARANRNSS 954
>TC86215 similar to GP|19698867|gb|AAL91169.1 unknown protein {Arabidopsis
thaliana}, partial (91%)
Length = 2293
Score = 427 bits (1099), Expect = e-120
Identities = 212/432 (49%), Positives = 297/432 (68%), Gaps = 1/432 (0%)
Frame = +3
Query: 2 DGTLTDHEVNEFQIRSFGASLQQPDITQLKTMVLRNVPEGVNSLGLTFPGFIEIHNMFLK 61
DG L+D E+N+FQ+R F A LQ +I +K +V + EGVN GLT GF+ +H +F++
Sbjct: 699 DGALSDAELNDFQVRCFNAPLQPAEIVGVKKVVQDKLSEGVNERGLTLTGFLFLHALFIE 878
Query: 62 KGRTETFWAVLRKFGYGNDLKLRDDFLPVPSKKASDQSVELSGAAIEFLKGVFRLVDTDK 121
KGR ET W VLRKFGY +++KL DD +P P K+ DQSVEL+ AI+FLK F D D
Sbjct: 879 KGRLETTWTVLRKFGYNDEIKLADDAVP-PLKRTPDQSVELTHEAIDFLKTTFESFDADL 1055
Query: 122 DQLLRPAEVDKLFDAAPESPWKDAPYMDAAETTDMGYISLKGFLSRWALMTLLDPRYSLA 181
D +LRP E+++LF APESPW Y DA E G +SL FLS WALMTLL+P +SL
Sbjct: 1056 DGMLRPRELEELFSTAPESPWIGNLYEDAVERNAFGGLSLDAFLSEWALMTLLNPSFSLE 1235
Query: 182 NLICIGYRDNPSAALRLTSRRSEDRKNQKTERNVFQCYVFGSKTAGKSALLRALLGRPFS 241
NLI IG+ P +A+R+T +R DRK Q +ERNV QC+VFG + +GKSA+L + +GRP+S
Sbjct: 1236 NLIYIGFPGEPLSAVRVTRKRRVDRKKQHSERNVLQCFVFGPRKSGKSAILNSFIGRPYS 1415
Query: 242 NDYTPTTVERYAANIIELIAGTKKTLILREIPEDEVSNFLSNKDCLAACDVAAFVHDSSD 301
Y PT +RYA N++++ +K L+L+EIPE EV+ +++K+ LA+CD+A FVHD SD
Sbjct: 1416 EAYNPTNEDRYAVNVVDISRENRKYLVLKEIPEGEVARLVADKESLASCDIAVFVHDRSD 1595
Query: 302 GYSWKRSIDLLEKVVNQGELTGHRFPSLLIAAKDDLTPFPRAVLDSVKVAQELKIDAPIR 361
SW+ S DLL ++ GE TG+ P L++AAKDDL PF A+ +S +V+Q++ ++API
Sbjct: 1596 ESSWRVSYDLLVQIAGHGENTGYEVPCLIVAAKDDLDPFALAIQESTRVSQDMGVEAPIP 1775
Query: 362 LSIKSDDSNNVYSKIINAAEHPHLSIPETEFVRKRKQHQQLLHTFIFALA-GAAMALAGF 420
+S+K D N+++ +I+ AAEHPHLSIPETE + RKQ ++L + ++ G A+A+ G
Sbjct: 1776 ISVKLGDFNSLFRRIVTAAEHPHLSIPETEAGKTRKQINKILSRSLLCVSVGTAVAVVGI 1955
Query: 421 TARRARANRNSS 432
A R A R S+
Sbjct: 1956 GAYRIYAARKSA 1991
>TC93654 weakly similar to GP|19571035|dbj|BAB86462. rac-GTP binding protein
-like {Oryza sativa (japonica cultivar-group)}, partial
(17%)
Length = 591
Score = 103 bits (256), Expect(2) = 1e-28
Identities = 55/92 (59%), Positives = 70/92 (75%), Gaps = 1/92 (1%)
Frame = +2
Query: 342 RAVLDSVKVAQELKIDAPIRLSIKSDDSNNVYSKIINAAEHPHLSIPETEFVRKRKQHQQ 401
RAV DSVKV EL I+API +S+K DSNNVY+KI++AAEHPHLSIPETE +KRKQ+ +
Sbjct: 77 RAVQDSVKVTLELGIEAPIHVSMKLGDSNNVYNKIVSAAEHPHLSIPETEIGKKRKQYNR 256
Query: 402 LL-HTFIFALAGAAMALAGFTARRARANRNSS 432
L+ ++ IFA G AMA+ G A RA A + +S
Sbjct: 257 LVQNSLIFASVGTAMAVVGLAACRAYAVKKNS 352
Score = 41.6 bits (96), Expect(2) = 1e-28
Identities = 18/23 (78%), Positives = 21/23 (91%)
Frame = +1
Query: 318 QGELTGHRFPSLLIAAKDDLTPF 340
QG+LTG+R P LLIAAKDDLTP+
Sbjct: 4 QGDLTGYRVPCLLIAAKDDLTPY 72
>TC81119 weakly similar to GP|19571035|dbj|BAB86462. rac-GTP binding protein
-like {Oryza sativa (japonica cultivar-group)}, partial
(20%)
Length = 737
Score = 50.4 bits (119), Expect = 1e-06
Identities = 23/47 (48%), Positives = 33/47 (69%)
Frame = +1
Query: 1 MDGTLTDHEVNEFQIRSFGASLQQPDITQLKTMVLRNVPEGVNSLGL 47
MD L D E+++FQ+R F A L+ P I +KT+V ++VPE +NS GL
Sbjct: 595 MDDALNDAELSDFQVRCFNAPLELPQIADVKTVVQQDVPERINSHGL 735
>BF004155 weakly similar to GP|16648803|gb| AT3g63150/T20O10_250 {Arabidopsis
thaliana}, partial (12%)
Length = 372
Score = 38.9 bits (89), Expect = 0.004
Identities = 18/25 (72%), Positives = 20/25 (80%)
Frame = +2
Query: 160 SLKGFLSRWALMTLLDPRYSLANLI 184
SL FLS WALMTLL+P +SL NLI
Sbjct: 293 SLDAFLSEWALMTLLNPSFSLENLI 367
>TC86339 small G-protein ROP6 [Medicago truncatula]
Length = 1195
Score = 37.7 bits (86), Expect = 0.009
Identities = 50/205 (24%), Positives = 83/205 (40%), Gaps = 11/205 (5%)
Frame = +2
Query: 205 DRKNQKTERNVF-QCYVFGSKTAGKSALLRALLGRPFSNDYTPTTVERYAANIIELIAGT 263
DR +K + F +C G GK+ LL + F DY PT + ++AN++ + G+
Sbjct: 212 DRV*KKMSASRFIKCVTVGDGAVGKTCLLISYTSNTFPTDYVPTVFDNFSANVV--VNGS 385
Query: 264 KKTLILREIPEDEVSNFLSNKDCLAA-CDVAAFVHDSSDGYSWKRSIDLLEKVVNQGELT 322
L L + E N L A + AF S Y ++ +K + + +
Sbjct: 386 TVNLGLWDTAGQEDYNRLRPLSYRGADVFILAFSLISKASYE-----NVSKKWIPELKHY 550
Query: 323 GHRFPSLLIAAKDDL-------TPFPRAVLDSVKVAQELK--IDAPIRLSIKSDDSNNVY 373
P +L+ K DL P AV + +EL+ I+AP + S NV
Sbjct: 551 APGVPIILVGTKLDLRDDKQFFVDHPGAVPITTAQGEELRKLINAPAYIECSSKSQQNV- 727
Query: 374 SKIINAAEHPHLSIPETEFVRKRKQ 398
+ +AA L P+ + + + Q
Sbjct: 728 KAVFDAAIRVVLQPPKQKKKKSKAQ 802
>TC88346 homologue to SP|Q40220|RAC2_LOTJA RAC-like GTP binding protein
RAC2. {Lotus japonicus}, partial (77%)
Length = 703
Score = 37.0 bits (84), Expect = 0.015
Identities = 14/57 (24%), Positives = 27/57 (46%)
Frame = +2
Query: 201 RRSEDRKNQKTERNVFQCYVFGSKTAGKSALLRALLGRPFSNDYTPTTVERYAANII 257
++ E K + + +C G GK+ +L + F DY PT + ++AN++
Sbjct: 221 KKKEGVKQKMSTARFIKCVTVGDGAVGKTCMLISYTSNTFPTDYVPTVFDNFSANVV 391
>TC79983 homologue to SP|Q41253|RACD_GOSHI RAC-like GTP binding protein
RAC13. [Upland cotton] {Gossypium hirsutum}, complete
Length = 902
Score = 36.2 bits (82), Expect = 0.025
Identities = 17/57 (29%), Positives = 27/57 (46%), Gaps = 4/57 (7%)
Frame = +3
Query: 205 DRKNQKTER----NVFQCYVFGSKTAGKSALLRALLGRPFSNDYTPTTVERYAANII 257
D K +KT R +C G GK+ +L + F DY PT + ++AN++
Sbjct: 105 DLKLRKTRRMSTARFIKCVTVGDGAVGKTCMLISYTSNTFPTDYVPTVFDNFSANVV 275
>TC87204 small G-protein ROP9 [Medicago truncatula]
Length = 995
Score = 36.2 bits (82), Expect = 0.025
Identities = 46/192 (23%), Positives = 77/192 (39%), Gaps = 10/192 (5%)
Frame = +3
Query: 217 QCYVFGSKTAGKSALLRALLGRPFSNDYTPTTVERYAANIIELIAGTKKTLILREIPEDE 276
+C G GK+ LL + F DY PT + ++AN++ + G+ L L + E
Sbjct: 174 KCVTVGDGAVGKTCLLISYTSNTFPTDYVPTVFDNFSANVV--VNGSIVNLGLWDTAGQE 347
Query: 277 VSNFLSNKDCLAA-CDVAAFVHDSSDGYSWKRSIDLLEKVVNQGELTGHRFPSLLIAAKD 335
N L A + AF S Y ++ +K + + + P +L+ K
Sbjct: 348 DYNRLRPLSYRGADVFILAFSLISKASYE-----NVSKKWIPELKHYAPGVPIILVGTKL 512
Query: 336 DL-------TPFPRAVLDSVKVAQELK--IDAPIRLSIKSDDSNNVYSKIINAAEHPHLS 386
DL P AV + +EL+ I+AP + S NV + +AA L
Sbjct: 513 DLRDDKQFCIDHPGAVPITTAQGEELRKLINAPAYIECSSKSQENV-KAVFDAAIRVVLQ 689
Query: 387 IPETEFVRKRKQ 398
P+ + + + Q
Sbjct: 690 PPKQKKKKNKAQ 725
>TC78546 small G-protein ROP3 [Medicago truncatula]
Length = 1177
Score = 34.7 bits (78), Expect = 0.073
Identities = 13/41 (31%), Positives = 21/41 (50%)
Frame = +1
Query: 217 QCYVFGSKTAGKSALLRALLGRPFSNDYTPTTVERYAANII 257
+C G GK+ LL + F DY PT + ++AN++
Sbjct: 160 KCVTVGDGAVGKTCLLISYTSNTFPTDYVPTVFDNFSANVV 282
>BF647926 homologue to PIR|T06679|T066 GTP-binding protein Arac8 -
Arabidopsis thaliana, partial (31%)
Length = 646
Score = 33.9 bits (76), Expect = 0.13
Identities = 13/47 (27%), Positives = 21/47 (44%)
Frame = +1
Query: 211 TERNVFQCYVFGSKTAGKSALLRALLGRPFSNDYTPTTVERYAANII 257
T +C G GK+ +L F DY PT + ++AN++
Sbjct: 88 TASRFIKCVTVGDGAVGKTCMLICYTSNKFPTDYIPTVFDNFSANVV 228
>TC90907 homologue to PIR|T08950|T08950 GTP binding protein Arac7 [imported]
- Arabidopsis thaliana, partial (55%)
Length = 474
Score = 32.0 bits (71), Expect = 0.48
Identities = 12/40 (30%), Positives = 19/40 (47%)
Frame = +3
Query: 217 QCYVFGSKTAGKSALLRALLGRPFSNDYTPTTVERYAANI 256
+C G GK+ +L F DY PT + ++AN+
Sbjct: 144 KCVTVGDGAVGKTCMLICYTSNKFPTDYIPTVFDNFSANV 263
>TC81415 weakly similar to PIR|A84509|A84509 probable ABC transporter
[imported] - Arabidopsis thaliana, partial (13%)
Length = 702
Score = 28.5 bits (62), Expect = 5.3
Identities = 13/30 (43%), Positives = 16/30 (53%)
Frame = +2
Query: 220 VFGSKTAGKSALLRALLGRPFSNDYTPTTV 249
V G GKS LLR + GR D+ P T+
Sbjct: 284 VVGPSGTGKSTLLRIIAGRIKDKDFNPKTI 373
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.319 0.135 0.390
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 11,362,522
Number of Sequences: 36976
Number of extensions: 139351
Number of successful extensions: 632
Number of sequences better than 10.0: 26
Number of HSP's better than 10.0 without gapping: 629
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 630
length of query: 432
length of database: 9,014,727
effective HSP length: 99
effective length of query: 333
effective length of database: 5,354,103
effective search space: 1782916299
effective search space used: 1782916299
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 60 (27.7 bits)
Medicago: description of AC135317.6


