

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC135317.3 + phase: 0
(701 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
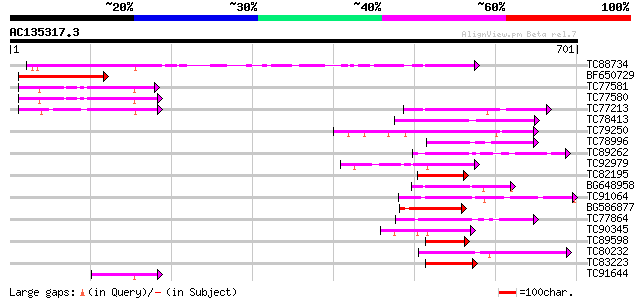
Score E
Sequences producing significant alignments: (bits) Value
TC88734 similar to GP|13346180|gb|AAK19612.1 GHDEL61 {Gossypium ... 246 2e-65
BF650729 homologue to GP|20563691|gb putative basic helix-loop-h... 134 9e-32
TC77581 similar to PIR|T10861|T10861 phaseolin G-box binding pro... 101 1e-21
TC77580 similar to PIR|T10861|T10861 phaseolin G-box binding pro... 101 1e-21
TC77213 similar to PIR|T10861|T10861 phaseolin G-box binding pro... 93 3e-19
TC78413 similar to PIR|T06329|T06329 symbiotic ammonium transpor... 91 2e-18
TC79250 similar to PIR|T06329|T06329 symbiotic ammonium transpor... 80 3e-15
TC78996 similar to GP|20127022|gb|AAM10937.1 putative bHLH trans... 79 5e-15
TC89262 similar to GP|21537346|gb|AAM61687.1 unknown {Arabidopsi... 75 9e-14
TC92979 similar to GP|20127014|gb|AAM10934.1 putative bHLH trans... 74 2e-13
TC82195 homologue to GP|19310475|gb|AAL84972.1 AT3g26744/MLJ15_1... 74 2e-13
BG648958 similar to GP|19310467|gb| At1g01260/F6F3_25 {Arabidops... 74 2e-13
TC91064 weakly similar to GP|20127014|gb|AAM10934.1 putative bHL... 70 2e-12
BG586877 similar to GP|16226919|gb AT4g16430/dl4240w {Arabidopsi... 69 8e-12
TC77864 weakly similar to PIR|T06329|T06329 symbiotic ammonium t... 67 2e-11
TC90345 similar to GP|20127014|gb|AAM10934.1 putative bHLH trans... 67 2e-11
TC89598 weakly similar to GP|20127014|gb|AAM10934.1 putative bHL... 67 2e-11
TC80232 similar to GP|21554896|gb|AAM63723.1 unknown {Arabidopsi... 66 5e-11
TC83223 similar to GP|8777436|dbj|BAA97026.1 gb|AAC78547.1~gene_... 62 7e-10
TC91644 similar to GP|19310467|gb|AAL84968.1 At1g01260/F6F3_25 {... 62 7e-10
>TC88734 similar to GP|13346180|gb|AAK19612.1 GHDEL61 {Gossypium hirsutum},
partial (40%)
Length = 1701
Score = 246 bits (628), Expect = 2e-65
Identities = 191/585 (32%), Positives = 279/585 (47%), Gaps = 24/585 (4%)
Frame = +3
Query: 21 SVQWTY-----SLFWQL--------------CPQQLILVWGDGYYNGSIKTRKTVQPMEV 61
S+QW + +L WQL Q WG+GYYNG IKTRKT Q +E+
Sbjct: 180 SIQWCHKT*RNNLLWQLEAFNGVM*YSGLNLLTNQGC*AWGEGYYNGEIKTRKTSQGVEL 359
Query: 62 SAEEASLQRSQQLRELYESLSAGETNPPTRRPCASLSPEDLTESEWFYLMCVSFSFPPGV 121
S++E LQRS+QLREL+ SL +TNP T+RP A+LSP+DLT++EW+YL+C+SF F G
Sbjct: 360 SSDEIGLQRSEQLRELFRSLKPVDTNPQTKRPSAALSPDDLTDTEWYYLVCMSFVFNIGQ 539
Query: 122 GLPGRAYTKRQHIWLTGANEVDSKIFSRAILAK-----TVVCIPVLDGVVEFGTTDKVQE 176
G+PGRA Q IWL A VD K FSRA+LAK TVVC P + GV+E GTTD V E
Sbjct: 540 GVPGRALENGQPIWLIDAYSVDCKEFSRALLAKSASIQTVVCFPFMKGVIELGTTDHVSE 719
Query: 177 DLNFIKHVKSFFLDGHSLPPKPALSEHSTSNPTSSTDHIPTIMYTMADPPSTNPNQDDMD 236
DL+ I+ +K+ L +SL ++ +T + + + + + + +
Sbjct: 720 DLSLIQQIKTSLL--NSLSVDDPINARATLS-SRNNEGVACVAF---------------- 842
Query: 237 EDEEEEEEEDEEDDEVESESEDETGGRIRRATSMTAIVEPSELMQIEMPDDIRIGSPNDG 296
+D++ + E E G I T+ SPN
Sbjct: 843 -----------DDNDYDVELIPEVGYDIINTTT----------------------SPNGS 923
Query: 297 SNHLDSDFHLLVVSNQGNPLGHVDSYKTDLTQRWGPIEEPVGNLLQVQLPSSDPYIYMWS 356
SN L Q N L + +T + + W +++ + N + + SSD
Sbjct: 924 SNAL-----------QANQLRN----ETFMVESWQVLDDDLSNCVHNSVNSSD------- 1037
Query: 357 ILINSIVYPIFAGSTSMQNRTQFNAQPPKSKNIHTPHVLHHQLEDLTQEDTHYSQTVSTI 416
I+ + N + + P S++ H Y + +S +
Sbjct: 1038-CISQNISSAPKVGEDCNNNQKMSLVDPLSEDWH------------------YQKILSEL 1160
Query: 417 LQNQSTQWTIDSSPSINYITSSNQSSFTNWTNHHNFHPLPPPETTTTTSQCLLKYILFTV 476
L++ + Q T+ +++ +SSF W + P+ TSQ LLK ILF V
Sbjct: 1161LKS-NDQLTM----GMHFQNFHQESSFCVW---NKGVPVNFQRPRQGTSQKLLKKILFEV 1316
Query: 477 PYLHTKNHDETSPQTHDAGVDPSSKLRGKGTPQDELSANHVLAERRRREKLNERFIILRS 536
P +HT E+ + + G DE NHVL+ERRRR KLNERF+ LRS
Sbjct: 1317PRMHTDGLLESQ--------EENDYREGTRLEADE-GMNHVLSERRRRAKLNERFLTLRS 1469
Query: 537 LVPFVTKMDKASILGDTIEYLKQLRRKIQDLETRNRQIETEQQSR 581
+VP +K DK SIL D IEYL++L +KI++LE + E +S+
Sbjct: 1470MVPSNSKDDKVSILDDAIEYLRKLEKKIRELEAQREPTGIESRSK 1604
Score = 28.5 bits (62), Expect = 9.2
Identities = 9/20 (45%), Positives = 15/20 (75%)
Frame = +2
Query: 11 LQNMLQAAVQSVQWTYSLFW 30
++ L AV+S+QW+Y +FW
Sbjct: 203 MKEQLALAVRSIQWSYVIFW 262
>BF650729 homologue to GP|20563691|gb putative basic helix-loop-helix protein
{Medicago sativa}, partial (88%)
Length = 652
Score = 134 bits (338), Expect = 9e-32
Identities = 62/113 (54%), Positives = 86/113 (75%), Gaps = 1/113 (0%)
Frame = +1
Query: 11 LQNMLQAAVQSVQWTYSLFW-QLCPQQLILVWGDGYYNGSIKTRKTVQPMEVSAEEASLQ 69
++ L AV+S+QW+Y +FW + Q +L WG+GYYNG IKTRKT Q +E+S++E LQ
Sbjct: 256 MKEQLALAVRSIQWSYVIFWSESVNQPGVLSWGEGYYNGEIKTRKTSQXVELSSDEIGLQ 435
Query: 70 RSQQLRELYESLSAGETNPPTRRPCASLSPEDLTESEWFYLMCVSFSFPPGVG 122
RS+QLREL+ SL +TNP T+RP A+LSP+DLT++EW+YL+C+SF F G G
Sbjct: 436 RSEQLRELFRSLKPVDTNPQTKRPSAALSPDDLTDTEWYYLVCMSFVFNIGQG 594
>TC77581 similar to PIR|T10861|T10861 phaseolin G-box binding protein PG1 -
kidney bean, partial (27%)
Length = 1033
Score = 101 bits (251), Expect = 1e-21
Identities = 65/185 (35%), Positives = 97/185 (52%), Gaps = 10/185 (5%)
Frame = +2
Query: 11 LQNMLQAAVQSVQ--WTYSLFWQLCPQ---QLILVWGDGYYNGSIKTRKTVQPMEVSAEE 65
LQ LQA ++ V+ WTY++FWQ +L WGDGYY G K + + S E
Sbjct: 251 LQQRLQALIEGVKEIWTYAIFWQPSYDYSGSSLLGWGDGYYKGEEDKTKVKKSIVTSPAE 430
Query: 66 ASLQRSQQLRELYESLSAGETNPPTRRPCASLSPEDLTESEWFYLMCVSFSFPPGVGLPG 125
R + LRELY +S NP T S E++T+ EWF+L+ ++ SF GLPG
Sbjct: 431 QE-HRRKVLRELYSLISG---NPVTEE---SPVDEEVTDMEWFFLVSMTQSFVNDGGLPG 589
Query: 126 RAYTKRQHIWLTGANEVDSKIFSRAILA-----KTVVCIPVLDGVVEFGTTDKVQEDLNF 180
+AY +WL G + RA +T+VC+P +GV+E G+T+ + ++ +F
Sbjct: 590 QAYFNSTPVWLVGGENLVLSHCERARQGQEHGLETLVCVPSANGVLELGSTELIYQNNDF 769
Query: 181 IKHVK 185
+ VK
Sbjct: 770 MDKVK 784
>TC77580 similar to PIR|T10861|T10861 phaseolin G-box binding protein PG1 -
kidney bean, partial (42%)
Length = 1383
Score = 101 bits (251), Expect = 1e-21
Identities = 66/188 (35%), Positives = 96/188 (50%), Gaps = 10/188 (5%)
Frame = +3
Query: 11 LQNMLQAAVQSVQ--WTYSLFWQLCPQ---QLILVWGDGYYNGSIKTRKTVQPMEVSAEE 65
LQ LQA ++ + WTY++FWQ +L WGDGYY G K + S E
Sbjct: 252 LQQRLQALIEGAKEIWTYAIFWQPSYDYSGSSLLGWGDGYYKGEEDKTKAKKSKVTSPAE 431
Query: 66 ASLQRSQQLRELYESLSAGETNPPTRRPCASLSPEDLTESEWFYLMCVSFSFPPGVGLPG 125
R + LREL +S NP T S E++T++EWF+L+ ++ SF G GLPG
Sbjct: 432 QE-HRRKVLRELNSLISG---NPVTDE---SPVDEEVTDTEWFFLVSMTQSFVNGTGLPG 590
Query: 126 RAYTKRQHIWLTGANEVDSKIFSRAILA-----KTVVCIPVLDGVVEFGTTDKVQEDLNF 180
+AY +WLTGA + RA +T+ CI DGV+E G+T+ + ++ +
Sbjct: 591 QAYYNSAPVWLTGAENLALSACERARQGQEHGIQTLACIRSADGVLELGSTELIYQNNDL 770
Query: 181 IKHVKSFF 188
+ VK F
Sbjct: 771 MNKVKMLF 794
>TC77213 similar to PIR|T10861|T10861 phaseolin G-box binding protein PG1 -
kidney bean, partial (70%)
Length = 2391
Score = 93.2 bits (230), Expect = 3e-19
Identities = 59/191 (30%), Positives = 100/191 (51%), Gaps = 9/191 (4%)
Frame = +3
Query: 488 SPQTHDAGVDPSSKLRGKGTPQDELSANHVLAERRRREKLNERFIILRSLVPFVTKMDKA 547
S + + G P + R ++E NHV AER+RREKLN++F LR++VP V+KMDKA
Sbjct: 1467 SSRVVEPGKRPKKRGRKPANGREE-PLNHVEAERQRREKLNQKFYALRAVVPNVSKMDKA 1643
Query: 548 SILGDTIEYLKQLRRKIQDLETRNRQIETEQQSRSGVTVLVG---------PTDKKKVRI 598
S+LGD I Y+ +L+ K+Q LE+ ++E + + LV P DK+K +
Sbjct: 1644 SLLGDAISYINELKSKLQGLESSKGELEKQLGATKKELELVASKNQSQNPIPLDKEKEKT 1823
Query: 599 VEECGATRAKAVETEVVSSVQVSIIESDALLEIECLHREGLLLDVMVMLRELRIEVIGVQ 658
+++ + + V I+ DA++ I+C + +M L+EL ++V
Sbjct: 1824 TSSTSSSKL------IDLDIDVKIMGWDAMIRIQCSKKNHPAAKLMAALKELDLDVNHAS 1985
Query: 659 SSLNNGVFVAE 669
S+ N + + +
Sbjct: 1986 VSVVNDLMIQQ 2018
Score = 89.4 bits (220), Expect = 4e-18
Identities = 60/190 (31%), Positives = 93/190 (48%), Gaps = 12/190 (6%)
Frame = +3
Query: 11 LQNMLQAAVQSVQ--WTYSLFWQLCPQQL----ILVWGDGYYNGSIKTRKTVQPMEVSAE 64
LQ+ LQA ++ + WTY++FWQ +L WGDGYY G V+ V+
Sbjct: 321 LQHRLQALIEGAKESWTYAIFWQSSYDYTMATPLLGWGDGYYKGE---DDKVKLKRVTPP 491
Query: 65 EASLQRSQQLRELYESLSAGETNPPTRRPCASLSPEDLTESEWFYLMCVSFSFPPGVGLP 124
E R + LREL +S G + ED+T++EWF+L ++ SF G G
Sbjct: 492 EEQAHRRKILRELNTLISGGSSVSD------DAVEEDVTDTEWFFLTSMTQSFVNGTGSL 653
Query: 125 GRAYTKRQHIWLTGANEVDSKIFSRAILAK-----TVVCIPV-LDGVVEFGTTDKVQEDL 178
+AY +W+TGA + RA A+ T+VCIP GVVE +T+ + +
Sbjct: 654 SQAYFNSTPVWITGAERLSGSPCERAREARVHGFQTLVCIPTSSSGVVELASTEMIPYNA 833
Query: 179 NFIKHVKSFF 188
+ ++ ++ F
Sbjct: 834 DLMEKIRVLF 863
>TC78413 similar to PIR|T06329|T06329 symbiotic ammonium transport protein
SAT1 - soybean, partial (30%)
Length = 1397
Score = 90.5 bits (223), Expect = 2e-18
Identities = 53/181 (29%), Positives = 98/181 (53%), Gaps = 1/181 (0%)
Frame = +2
Query: 476 VPYLHTKNHDETSPQTHDAGVDPSSKLRGKGTPQDEL-SANHVLAERRRREKLNERFIIL 534
VP T+ + + K +G+GT + + +H++AER+RREKL++ I L
Sbjct: 410 VPLSQTELPQNRKGSLQKQNIVETIKPQGQGTKRSVAHNQDHIIAERKRREKLSQCLIAL 589
Query: 535 RSLVPFVTKMDKASILGDTIEYLKQLRRKIQDLETRNRQIETEQQSRSGVTVLVGPTDKK 594
+L+P + KMDKAS+LGD I+Y+K+L+ +++ LE EQ S V +V +++
Sbjct: 590 AALIPGLKKMDKASVLGDAIKYVKELQERLRVLE--------EQNKNSHVQSVVTVDEQQ 745
Query: 595 KVRIVEECGATRAKAVETEVVSSVQVSIIESDALLEIECLHREGLLLDVMVMLRELRIEV 654
+ + E + V+ +++ D L+ I C ++GLLL ++V +++L + V
Sbjct: 746 LSYDSSNSDDSEVASGNNETLPHVEAKVLDKDVLIRIHCQKQKGLLLKILVEIQKLHLFV 925
Query: 655 I 655
+
Sbjct: 926 V 928
>TC79250 similar to PIR|T06329|T06329 symbiotic ammonium transport protein
SAT1 - soybean, partial (34%)
Length = 1145
Score = 79.7 bits (195), Expect = 3e-15
Identities = 69/281 (24%), Positives = 126/281 (44%), Gaps = 27/281 (9%)
Frame = +2
Query: 401 DLTQEDTHYSQTVSTIL---QNQSTQWTIDSSPSINYITS---SNQSSFTNWTNHHN--F 452
DL ED + ++ +N + Q + ++ TS SN S N TN N +
Sbjct: 104 DLEPEDYNIINEINIATLPQENNNLQQSFSGGSHCSHTTSNTMSNSSGDVNKTNPSNIGY 283
Query: 453 HPLPPPETTTTTSQC--------LLKYILFTVPYLHTKNHDET-----SPQTHDAGVDPS 499
LP +++ + C +L P NH ++ S + G +
Sbjct: 284 PSLPKKDSSHSYILCFGNENPESMLNIGSTLKPKGKVSNHGKSLASKGSLENQKKGPKRN 463
Query: 500 SKLRGKGTPQDELSANHVLAERRRREKLNERFIILRSLVPFVTKMDKASILGDTIEYLKQ 559
+ K + +H++AER+RREK++++FI L +L+P + KMDKAS+LGD I ++KQ
Sbjct: 464 IQESKKTDSAARNAQDHIIAERKRREKISQKFIALSALLPDLKKMDKASVLGDAINHVKQ 643
Query: 560 LRRKIQDLETRNRQIETEQQSRSGVTVLVGPTDKKKVRIVE------ECGATRAKAVETE 613
L+ K++ LE +N++ E S V + + V C T + +
Sbjct: 644 LQEKVKLLEEKNQKNNVESVSMVYVEKTKSYSSDEDVSETSSNSGYGNCCHTHT-SKPSR 820
Query: 614 VVSSVQVSIIESDALLEIECLHREGLLLDVMVMLRELRIEV 654
+ V+ + E + L+ + C +G L++++ + L + V
Sbjct: 821 SLPEVEARVSEKNVLIRVHCEKHKGALMNIIQEIENLHLSV 943
>TC78996 similar to GP|20127022|gb|AAM10937.1 putative bHLH transcription
factor {Arabidopsis thaliana}, partial (27%)
Length = 1081
Score = 79.3 bits (194), Expect = 5e-15
Identities = 48/139 (34%), Positives = 73/139 (51%)
Frame = +3
Query: 516 HVLAERRRREKLNERFIILRSLVPFVTKMDKASILGDTIEYLKQLRRKIQDLETRNRQIE 575
HV+AER+RREKL FI L ++VP + KMDKAS+LGD +Y+KQL+ ++Q LE +
Sbjct: 69 HVMAERKRREKLTRSFIALSAIVPGLKKMDKASVLGDATKYMKQLQARLQTLEE-----Q 233
Query: 576 TEQQSRSGVTVLVGPTDKKKVRIVEECGATRAKAVETEVVSSVQVSIIESDALLEIECLH 635
E ++G TV V K I+ + + ++V + D L++I+C
Sbjct: 234 AEDNKKAGSTVQV------KRSIIFTNNNDDDSNSNNQPLPEIEVRVSSKDVLIKIQCDK 395
Query: 636 REGLLLDVMVMLRELRIEV 654
G V+ L L + V
Sbjct: 396 HSGRAATVLGQLENLNLTV 452
>TC89262 similar to GP|21537346|gb|AAM61687.1 unknown {Arabidopsis
thaliana}, partial (40%)
Length = 1198
Score = 75.1 bits (183), Expect = 9e-14
Identities = 60/195 (30%), Positives = 104/195 (52%)
Frame = +3
Query: 499 SSKLRGKGTPQDELSANHVLAERRRREKLNERFIILRSLVPFVTKMDKASILGDTIEYLK 558
S KL+G+ + +++AERRRR++LN+R +LR++VP ++KMD+ +ILGDTI+Y+K
Sbjct: 528 SKKLQGQ-------PSKNLMAERRRRKRLNDRLSMLRAIVPKISKMDRTAILGDTIDYMK 686
Query: 559 QLRRKIQDLETRNRQIETEQQSRSGVTVLVGPTDKKKVRIVEECGATRAKAVETEVVSSV 618
+L KI++L+ ++IE + S +T +V ++ I + + EV S
Sbjct: 687 ELLEKIKNLQ---QEIELD----SNMTSIVKDVKPNEILI--------RNSPKFEVERS- 818
Query: 619 QVSIIESDALLEIECLHREGLLLDVMVMLRELRIEVIGVQSSLNNGVFVAELRAKVKENG 678
+D +EI C + GLLL + L L +E+ S N ++A E
Sbjct: 819 ------ADTRVEICCAGKPGLLLSTVNTLEALGLEIQQCVISCFNDF---TMQASCSEEL 971
Query: 679 GNGKKVSIVEVKRAL 693
+ +S ++K+AL
Sbjct: 972 EKREMLSSEDIKQAL 1016
>TC92979 similar to GP|20127014|gb|AAM10934.1 putative bHLH transcription
factor {Arabidopsis thaliana}, partial (15%)
Length = 744
Score = 73.9 bits (180), Expect = 2e-13
Identities = 56/186 (30%), Positives = 96/186 (51%), Gaps = 14/186 (7%)
Frame = +2
Query: 410 SQTVSTILQNQSTQW----------TIDSSPSINY-ITSSNQSSFTNWTNHHNFHPLPPP 458
S +S + Q Q+ Q TID+S N ++ S Q ++ N PP
Sbjct: 194 SNDISNVFQEQNKQQCLSLGSTFHETIDNSDKNNESLSPSFQFQVPSFDN--------PP 349
Query: 459 ETTTTTSQCLLKYILFTVPYLHTKNHDETSPQTHDAGVDPSSKLRGKGTPQDELSAN--- 515
++ T S+ ++ I + L NH S +T ++ + KL K + A+
Sbjct: 350 NSSPTNSKENIETIPLSPTDLENMNH---STETSKGSLE-NKKLETKTSKSKRPRAHGRD 517
Query: 516 HVLAERRRREKLNERFIILRSLVPFVTKMDKASILGDTIEYLKQLRRKIQDLETRNRQIE 575
H++AER RREKL + FI L +LVP + KMDK S+L DTI+Y+K+L+ +++D+ N++ +
Sbjct: 518 HIMAERNRREKLTQSFIALAALVPNLKKMDKLSVLIDTIKYMKELKNRLEDVGEPNQKTK 697
Query: 576 TEQQSR 581
+ ++
Sbjct: 698 KKSSTK 715
>TC82195 homologue to GP|19310475|gb|AAL84972.1 AT3g26744/MLJ15_15
{Arabidopsis thaliana}, partial (17%)
Length = 1189
Score = 73.6 bits (179), Expect = 2e-13
Identities = 34/63 (53%), Positives = 51/63 (79%)
Frame = +2
Query: 505 KGTPQDELSANHVLAERRRREKLNERFIILRSLVPFVTKMDKASILGDTIEYLKQLRRKI 564
KG + L A +++AERRRR+KLN+R +LRS+VP ++KMD+ASILGD ++YLK+L ++I
Sbjct: 923 KGKKKKGLPAKNLMAERRRRKKLNDRLYMLRSVVPKISKMDRASILGDAVDYLKELLQRI 1102
Query: 565 QDL 567
+L
Sbjct: 1103NNL 1111
>BG648958 similar to GP|19310467|gb| At1g01260/F6F3_25 {Arabidopsis
thaliana}, partial (15%)
Length = 806
Score = 73.6 bits (179), Expect = 2e-13
Identities = 52/145 (35%), Positives = 78/145 (52%), Gaps = 17/145 (11%)
Frame = +3
Query: 498 PSSKLRGKGTPQDELSANHVLAERRRREKLNERFIILRSLVPFVTKMDKASILGDTIEYL 557
P + R +DE NHV AER+RREKLN+RF LR++VP ++KMDKAS+LGD I Y+
Sbjct: 282 PRKRGRKPANGRDE-PLNHVEAERQRREKLNQRFYALRAVVPNISKMDKASLLGDAIAYI 458
Query: 558 KQLRRKIQDLETRNRQIETEQQSRSGV----------TVLVGPTDKKKVRIVEECGATRA 607
+L+ K++ +E+ R+ SR G+ V P D +E +
Sbjct: 459 NELQAKLKIMES-ERESFGSSTSRDGLANSRSENRHQNTRVPPPDVDIQASQDEVIVKVS 635
Query: 608 KAVETEVVSSV-------QVSIIES 625
+++T VS V Q+S++ES
Sbjct: 636 CSLDTHPVSKVIETFKEAQISVVES 710
>TC91064 weakly similar to GP|20127014|gb|AAM10934.1 putative bHLH
transcription factor {Arabidopsis thaliana}, partial
(19%)
Length = 751
Score = 70.5 bits (171), Expect = 2e-12
Identities = 54/228 (23%), Positives = 112/228 (48%), Gaps = 7/228 (3%)
Frame = +2
Query: 481 TKNHDETSPQTHDAGVDPSSKLRGKGTPQDELSANHVLAERRRREKLNERFIILRSLVPF 540
T+ H ++ P + S+ + + T + +H+++ER RR+ L + I L +L+P
Sbjct: 86 TEQHHDSLPLSSSTANQGSNSKKPRNTSD---TLDHIMSERNRRQLLTSKIIELSALIPG 256
Query: 541 VTKMDKASILGDTIEYLKQLRRKIQDLETRNRQIETEQQSRSGVT---VLVGPTDKKKVR 597
+ K+DK ++ + I Y+KQL ++++LE + + + S S +T VL+
Sbjct: 257 LKKIDKVHVVTEAINYMKQLEERLKELEEDIK--KKDAGSLSTITRSRVLIDKDIAIGEM 430
Query: 598 IVEECGATRAKAVETEVVSSVQVSIIESDALLEIECLHREGLLLDVMVMLRELRIEVIGV 657
EEC +E V+ I+E + L++I C +EG+++++M L+ L + +
Sbjct: 431 NTEECYGRNESLLE------VEARILEKEVLIKIYCGMQEGIVVNIMSQLQLLHLSI--- 583
Query: 658 QSSLNNGVFVAELRAKVKENGGNGKKVSIVEVKRALNQI----IPHNI 701
+S+N F L + G+ ++I ++ + L + + HN+
Sbjct: 584 -TSINVLPFGNTLDITIIAKMGDKYNLTIKDLVKKLRVVATLQVSHNV 724
>BG586877 similar to GP|16226919|gb AT4g16430/dl4240w {Arabidopsis thaliana},
partial (35%)
Length = 755
Score = 68.6 bits (166), Expect = 8e-12
Identities = 39/84 (46%), Positives = 54/84 (63%), Gaps = 1/84 (1%)
Frame = +1
Query: 483 NHDETSPQTHDAGVDPSSKLRGKGTPQD-ELSANHVLAERRRREKLNERFIILRSLVPFV 541
N D +SP G + + RG+ E NHV AER+RREKLN+RF LR+ VP +
Sbjct: 508 NEDSSSP----LGEERKPRKRGRKPANGREEPLNHVEAERQRREKLNQRFYALRAGVPNI 675
Query: 542 TKMDKASILGDTIEYLKQLRRKIQ 565
+KMDKAS+LGD I ++ L++KI+
Sbjct: 676 SKMDKASLLGDAITHITDLQKKIK 747
>TC77864 weakly similar to PIR|T06329|T06329 symbiotic ammonium transport
protein SAT1 - soybean, partial (21%)
Length = 1411
Score = 67.4 bits (163), Expect = 2e-11
Identities = 44/178 (24%), Positives = 86/178 (47%)
Frame = +3
Query: 477 PYLHTKNHDETSPQTHDAGVDPSSKLRGKGTPQDELSANHVLAERRRREKLNERFIILRS 536
P + +H GV ++K K E +H++AER+RR +L E I L +
Sbjct: 453 PIIEPMSHKTKRRTDESRGVKEATK---KVRRSCETVQDHLMAERKRRRELTENIIALSA 623
Query: 537 LVPFVTKMDKASILGDTIEYLKQLRRKIQDLETRNRQIETEQQSRSGVTVLVGPTDKKKV 596
++P + KMDK +L + + Y KQL+++I++LE +N+ S+ + K K
Sbjct: 624 MIPGLKKMDKCYVLSEAVNYTKQLQKRIKELENQNK------DSKPNPAIF-----KWKS 770
Query: 597 RIVEECGATRAKAVETEVVSSVQVSIIESDALLEIECLHREGLLLDVMVMLRELRIEV 654
++ + K +E + V+ + E + L+ I C ++ ++L + +L + I +
Sbjct: 771 QV------SSNKKKSSESLLEVEARVKEKEVLIRIHCEKQKDIVLKIHELLEKFNITI 926
>TC90345 similar to GP|20127014|gb|AAM10934.1 putative bHLH transcription
factor {Arabidopsis thaliana}, partial (21%)
Length = 906
Score = 67.0 bits (162), Expect = 2e-11
Identities = 42/140 (30%), Positives = 75/140 (53%), Gaps = 23/140 (16%)
Frame = +1
Query: 459 ETTTTTSQCLLKYILF---TVPYLHTKNHDETSPQTHDAGVDPSSKL--------RGKGT 507
+TTTT S +K T Y+ + ++ P+ + + P +K+ R K
Sbjct: 337 KTTTTCSSSYMKTEHLSQKTASYVFSFENENPPPRKVEPALKPKTKVVNSKNGPRRVKNN 516
Query: 508 PQDELSAN------------HVLAERRRREKLNERFIILRSLVPFVTKMDKASILGDTIE 555
++ + + H++AER RREK+++ FI L +L+P + KMDKAS+LGD I+
Sbjct: 517 ESNKKNGSFSKSTTTHHTPDHIIAERIRREKISQLFIALSALIPNLKKMDKASVLGDAIK 696
Query: 556 YLKQLRRKIQDLETRNRQIE 575
Y+K+L+ ++ LE ++ +E
Sbjct: 697 YVKELKGAVKMLEEPSKSVE 756
>TC89598 weakly similar to GP|20127014|gb|AAM10934.1 putative bHLH
transcription factor {Arabidopsis thaliana}, partial
(24%)
Length = 849
Score = 67.0 bits (162), Expect = 2e-11
Identities = 30/54 (55%), Positives = 46/54 (84%)
Frame = +3
Query: 515 NHVLAERRRREKLNERFIILRSLVPFVTKMDKASILGDTIEYLKQLRRKIQDLE 568
+HV+AER+RREKL++RFI L S++P + KMDKA+IL D I+++KQL+ +++ LE
Sbjct: 537 DHVIAERKRREKLSQRFIALSSILPGLKKMDKATILEDAIKHMKQLQERVKTLE 698
>TC80232 similar to GP|21554896|gb|AAM63723.1 unknown {Arabidopsis
thaliana}, partial (62%)
Length = 952
Score = 65.9 bits (159), Expect = 5e-11
Identities = 47/200 (23%), Positives = 96/200 (47%), Gaps = 11/200 (5%)
Frame = +1
Query: 506 GTPQDELSANHVLAERRRREKLNERFIILRSLVPFVTKMDKASILGDTIEYLKQLRRKIQ 565
G +S+ ++++ER RR+KLNER LR++VP ++KMDKASI+ D IEY++ L + +
Sbjct: 196 GAASSGVSSKNIVSERNRRKKLNERLFALRAVVPNISKMDKASIIKDAIEYIQLLHEQEK 375
Query: 566 DLETRNRQIETEQQSRSGVTVLVGPT-----------DKKKVRIVEECGATRAKAVETEV 614
++ ++E SG+ + P+ KK R + + ++ EV
Sbjct: 376 VIQAEIMELE------SGMPNNINPSYDFDQELPMLLRSKKKRTDQLYDSVSSRNFPIEV 537
Query: 615 VSSVQVSIIESDALLEIECLHREGLLLDVMVMLRELRIEVIGVQSSLNNGVFVAELRAKV 674
+ + E+ ++ + C R ++ + + L++++I + +G + + +
Sbjct: 538 LELRVTYMGENTMVVSLTCNKRADTMVKLCEVFESLKLKIITANITSFSGRLLKTVFIEA 717
Query: 675 KENGGNGKKVSIVEVKRALN 694
E + + +I ALN
Sbjct: 718 NEEDKDQLQTNIQTAIAALN 777
>TC83223 similar to GP|8777436|dbj|BAA97026.1
gb|AAC78547.1~gene_id:MHM17.7~similar to unknown protein
{Arabidopsis thaliana}, partial (13%)
Length = 514
Score = 62.0 bits (149), Expect = 7e-10
Identities = 32/64 (50%), Positives = 45/64 (70%)
Frame = +1
Query: 515 NHVLAERRRREKLNERFIILRSLVPFVTKMDKASILGDTIEYLKQLRRKIQDLETRNRQI 574
+H+++ERRRREKLNE F LR+L+P TK DKASIL E L+ L +I L RN+++
Sbjct: 199 HHMISERRRREKLNENFQALRALLPQGTKKDKASILITAKETLRSLMAEIDKLSKRNQEL 378
Query: 575 ETEQ 578
++Q
Sbjct: 379 MSQQ 390
>TC91644 similar to GP|19310467|gb|AAL84968.1 At1g01260/F6F3_25 {Arabidopsis
thaliana}, partial (20%)
Length = 1021
Score = 62.0 bits (149), Expect = 7e-10
Identities = 35/93 (37%), Positives = 53/93 (56%), Gaps = 6/93 (6%)
Frame = +1
Query: 102 LTESEWFYLMCVSFSFPPGVGLPGRAYTKRQHIWLTGANEVDSKIFS-RAILA-----KT 155
+T++E F+L + FSFP G G PG+ + +H+WL A + S + RA LA KT
Sbjct: 130 VTDTEMFFLASMYFSFPNGHGGPGKCFASGKHLWLCDALKSSSSDYCVRAFLAKSVGFKT 309
Query: 156 VVCIPVLDGVVEFGTTDKVQEDLNFIKHVKSFF 188
+V +P GVVE G+ V E + ++ +KS F
Sbjct: 310 IVLVPTDLGVVELGSVTTVGESVELLQTIKSVF 408
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.313 0.131 0.377
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 21,703,765
Number of Sequences: 36976
Number of extensions: 334225
Number of successful extensions: 4984
Number of sequences better than 10.0: 302
Number of HSP's better than 10.0 without gapping: 2891
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 3941
length of query: 701
length of database: 9,014,727
effective HSP length: 103
effective length of query: 598
effective length of database: 5,206,199
effective search space: 3113307002
effective search space used: 3113307002
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 62 (28.5 bits)
Medicago: description of AC135317.3


