

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC135311.8 + phase: 0
(189 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
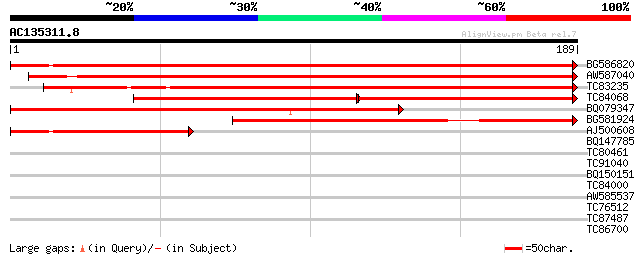
Score E
Sequences producing significant alignments: (bits) Value
BG586820 weakly similar to GP|7406671|emb putative ripening-rela... 356 3e-99
AW587040 weakly similar to GP|7406671|emb putative ripening-rela... 323 2e-89
TC83235 weakly similar to GP|7406671|emb|CAB85629.1 putative rip... 276 3e-75
TC84068 similar to GP|7406671|emb|CAB85629.1 putative ripening-r... 148 3e-71
BQ079347 weakly similar to GP|10140700|gb| hypothetical protein ... 203 4e-53
BG581924 similar to GP|7406671|emb| putative ripening-related pr... 145 1e-35
AJ500608 96 8e-21
BQ147785 weakly similar to GP|19347731|gb| unknown protein {Arab... 29 0.96
TC80461 similar to GP|21592749|gb|AAM64698.1 putative thaumatin-... 27 3.6
TC91040 similar to GP|11994500|dbj|BAB02565. gene_id:F4B12.2~unk... 27 4.7
BQ150151 27 4.7
TC84000 27 4.7
AW585537 SP|P03736|VMTH Minor tail protein precursor H. {Bacteri... 27 6.2
TC76512 similar to GP|4959932|gb|AAD34548.1| cystathionine-gamma... 27 6.2
TC87487 similar to SP|P52780|SYQ_LUPLU Glutaminyl-tRNA synthetas... 26 8.1
TC86700 similar to GP|18377835|gb|AAL67104.1 At1g24120/F3I6_4 {A... 26 8.1
>BG586820 weakly similar to GP|7406671|emb putative ripening-related protein
{Vitis vinifera}, partial (52%)
Length = 613
Score = 356 bits (913), Expect = 3e-99
Identities = 164/189 (86%), Positives = 175/189 (91%)
Frame = +1
Query: 1 MKRFCPKVSFIFLAITLIFTSFVYSEAQNCRPSGRIRGKKAPPGKCNQENDSDCCVQGKM 60
MK FCPKVSF+ L ITL+ TSFV+SEAQ CRP+GRIRGKKAP G+CN+ENDSDCCVQGKM
Sbjct: 28 MKSFCPKVSFLLL-ITLVLTSFVFSEAQKCRPNGRIRGKKAPSGQCNKENDSDCCVQGKM 204
Query: 61 YTTYVCSPSVSTHTKAYLTLNSFQKGGDGGGPSECDKQYHSDDTPVVALSTGWFNHKSRC 120
YTTY CSPSVSTHTKAYLTLNSFQKGGDGGGPS CD QYHSDDTPVVALSTGWFN KSRC
Sbjct: 205 YTTYECSPSVSTHTKAYLTLNSFQKGGDGGGPSACDNQYHSDDTPVVALSTGWFNDKSRC 384
Query: 121 LNNITISANGRNVVAMVVDECDSRKGCDEQHDYQPPCPNNIVDASKAVWKALGVPKEQWG 180
LN ITISANG++VVA VVDECDS GCDE+HDYQPPCPNNIVDASKAVWKALGVP++QWG
Sbjct: 385 LNKITISANGKSVVATVVDECDSTMGCDEEHDYQPPCPNNIVDASKAVWKALGVPQDQWG 564
Query: 181 GLDITWSDA 189
GLDITWSDA
Sbjct: 565 GLDITWSDA 591
>AW587040 weakly similar to GP|7406671|emb putative ripening-related protein
{Vitis vinifera}, partial (52%)
Length = 576
Score = 323 bits (828), Expect = 2e-89
Identities = 149/183 (81%), Positives = 162/183 (88%)
Frame = +3
Query: 7 KVSFIFLAITLIFTSFVYSEAQNCRPSGRIRGKKAPPGKCNQENDSDCCVQGKMYTTYVC 66
K SF+FL +I ++S AQ CRPSGRI+G+K P G+CNQENDSDCCVQGKMYTTY C
Sbjct: 6 KASFLFLVFLIIM---LHSAAQQCRPSGRIKGEKTPSGQCNQENDSDCCVQGKMYTTYKC 176
Query: 67 SPSVSTHTKAYLTLNSFQKGGDGGGPSECDKQYHSDDTPVVALSTGWFNHKSRCLNNITI 126
SPSVSTHTKAYLTLNSFQ+GGDGGGPS CD QYHSDDTPVVALSTGWFN+KSRCLN I I
Sbjct: 177 SPSVSTHTKAYLTLNSFQEGGDGGGPSACDNQYHSDDTPVVALSTGWFNNKSRCLNKIKI 356
Query: 127 SANGRNVVAMVVDECDSRKGCDEQHDYQPPCPNNIVDASKAVWKALGVPKEQWGGLDITW 186
SANGR+VVA VVDECDSR GCD++HDYQPPC NNIVDASKAVWKALGVP +QWGGLDITW
Sbjct: 357 SANGRSVVAKVVDECDSRAGCDKEHDYQPPCNNNIVDASKAVWKALGVPHDQWGGLDITW 536
Query: 187 SDA 189
SDA
Sbjct: 537 SDA 545
>TC83235 weakly similar to GP|7406671|emb|CAB85629.1 putative
ripening-related protein {Vitis vinifera}, partial (51%)
Length = 769
Score = 276 bits (706), Expect = 3e-75
Identities = 128/179 (71%), Positives = 150/179 (83%), Gaps = 1/179 (0%)
Frame = +3
Query: 12 FLAITLIF-TSFVYSEAQNCRPSGRIRGKKAPPGKCNQENDSDCCVQGKMYTTYVCSPSV 70
F+A+ ++F T+ +Y EAQ C P+G ++GK +P G+C E+D CCV GK Y TY CSP V
Sbjct: 48 FIALLVLFLTNCLYIEAQKCNPNGIVKGK-SPSGRCKHEDDL-CCVPGKKYQTYKCSPPV 221
Query: 71 STHTKAYLTLNSFQKGGDGGGPSECDKQYHSDDTPVVALSTGWFNHKSRCLNNITISANG 130
S+HTKAYLTLNSF+KGGDG GPS CD +Y+SDDTPVVALSTGWFN K RCL+NITIS NG
Sbjct: 222 SSHTKAYLTLNSFEKGGDGNGPSTCDNKYYSDDTPVVALSTGWFNTKRRCLHNITISGNG 401
Query: 131 RNVVAMVVDECDSRKGCDEQHDYQPPCPNNIVDASKAVWKALGVPKEQWGGLDITWSDA 189
++VVAMVVDECDS KGCD +H+YQPPC NNIVDAS AVWKALGVP+ QWGGLDITWSDA
Sbjct: 402 QSVVAMVVDECDSSKGCDAEHEYQPPCANNIVDASMAVWKALGVPRNQWGGLDITWSDA 578
>TC84068 similar to GP|7406671|emb|CAB85629.1 putative ripening-related
protein {Vitis vinifera}, partial (44%)
Length = 553
Score = 148 bits (373), Expect(2) = 3e-71
Identities = 66/76 (86%), Positives = 69/76 (89%)
Frame = +1
Query: 42 PPGKCNQENDSDCCVQGKMYTTYVCSPSVSTHTKAYLTLNSFQKGGDGGGPSECDKQYHS 101
P G+CNQENDSDCCVQ KMYTTY CSPS STHTKA LTLNSFQKGGDGGGPS CD QYHS
Sbjct: 1 PSGQCNQENDSDCCVQRKMYTTYQCSPSASTHTKANLTLNSFQKGGDGGGPSACDNQYHS 180
Query: 102 DDTPVVALSTGWFNHK 117
D+TPVVALSTGWFN+K
Sbjct: 181 DNTPVVALSTGWFNNK 228
Score = 137 bits (344), Expect(2) = 3e-71
Identities = 62/73 (84%), Positives = 66/73 (89%)
Frame = +2
Query: 117 KSRCLNNITISANGRNVVAMVVDECDSRKGCDEQHDYQPPCPNNIVDASKAVWKALGVPK 176
KSRCLN ITISANGR+VVA VVDECDS GCD++HDYQPPC NNIVDASKAVWKAL VP
Sbjct: 227 KSRCLNKITISANGRSVVATVVDECDSTMGCDKEHDYQPPCNNNIVDASKAVWKALDVPH 406
Query: 177 EQWGGLDITWSDA 189
+QWGGLDITWSDA
Sbjct: 407 DQWGGLDITWSDA 445
>BQ079347 weakly similar to GP|10140700|gb| hypothetical protein {Oryza
sativa}, partial (10%)
Length = 446
Score = 203 bits (516), Expect = 4e-53
Identities = 100/132 (75%), Positives = 106/132 (79%), Gaps = 1/132 (0%)
Frame = +1
Query: 1 MKRFCPKVSFIFLAITLIFTSFVYSEAQNCRPSGRIRGKKAPPGKCNQENDSDCCVQGKM 60
MKRFCPK SFI L ITLI TSFVYSEAQNCRPSGRIRGKKAPPG+CNQENDSDCCVQGKM
Sbjct: 28 MKRFCPKASFILLVITLILTSFVYSEAQNCRPSGRIRGKKAPPGQCNQENDSDCCVQGKM 207
Query: 61 YTTYVCSPSVSTHTKAYLTLNSFQKGGDGGGP-SECDKQYHSDDTPVVALSTGWFNHKSR 119
YTTYVCSPSVSTHTKAYLTLNSF+KGGDGG DDTPVVALS GW +S+
Sbjct: 208 YTTYVCSPSVSTHTKAYLTLNSFEKGGDGGWSFRNVTSSIIPDDTPVVALSXGWVQPQSQ 387
Query: 120 CLNNITISANGR 131
+ SANG+
Sbjct: 388 VPXPHSXSANGK 423
>BG581924 similar to GP|7406671|emb| putative ripening-related protein {Vitis
vinifera}, partial (31%)
Length = 792
Score = 145 bits (365), Expect = 1e-35
Identities = 73/115 (63%), Positives = 82/115 (70%)
Frame = +2
Query: 75 KAYLTLNSFQKGGDGGGPSECDKQYHSDDTPVVALSTGWFNHKSRCLNNITISANGRNVV 134
KA LTLNSF+KGGDGG PSECD ++HSD T VVALS+ FNHK RCL ITI NG+ V
Sbjct: 140 KAILTLNSFEKGGDGGAPSECDNKFHSDKTLVVALSSALFNHKKRCLKEITIFGNGKRVN 319
Query: 135 AMVVDECDSRKGCDEQHDYQPPCPNNIVDASKAVWKALGVPKEQWGGLDITWSDA 189
A VVDECDS KG C NNIVD S AVWKALGVP+++ G + I WSDA
Sbjct: 320 AKVVDECDSSKG----------CKNNIVDGSAAVWKALGVPEKRRGEMSIFWSDA 454
>AJ500608
Length = 222
Score = 95.9 bits (237), Expect = 8e-21
Identities = 42/61 (68%), Positives = 52/61 (84%)
Frame = -3
Query: 1 MKRFCPKVSFIFLAITLIFTSFVYSEAQNCRPSGRIRGKKAPPGKCNQENDSDCCVQGKM 60
M+ FC K SF+FL + L+ T+ ++SEAQ CRPSGRI+G+KAP G+CNQENDSDCCVQGKM
Sbjct: 181 MQSFCSKASFLFL-VFLVITNLLHSEAQQCRPSGRIKGEKAPSGQCNQENDSDCCVQGKM 5
Query: 61 Y 61
Y
Sbjct: 4 Y 2
>BQ147785 weakly similar to GP|19347731|gb| unknown protein {Arabidopsis
thaliana}, partial (43%)
Length = 449
Score = 29.3 bits (64), Expect = 0.96
Identities = 11/27 (40%), Positives = 18/27 (65%)
Frame = +2
Query: 157 CPNNIVDASKAVWKALGVPKEQWGGLD 183
CP + DAS ++LG+P ++W G+D
Sbjct: 89 CPYSWDDASYLTQRSLGLPDDRWRGID 169
>TC80461 similar to GP|21592749|gb|AAM64698.1 putative thaumatin-like
protein {Arabidopsis thaliana}, partial (63%)
Length = 983
Score = 27.3 bits (59), Expect = 3.6
Identities = 11/27 (40%), Positives = 14/27 (51%)
Frame = -3
Query: 149 EQHDYQPPCPNNIVDASKAVWKALGVP 175
E ++QP P N+ D WK L VP
Sbjct: 429 ESSNWQPVLPQNLPDHPVGAWKVLEVP 349
>TC91040 similar to GP|11994500|dbj|BAB02565. gene_id:F4B12.2~unknown
protein {Arabidopsis thaliana}, partial (64%)
Length = 882
Score = 26.9 bits (58), Expect = 4.7
Identities = 11/22 (50%), Positives = 17/22 (77%)
Frame = +3
Query: 100 HSDDTPVVALSTGWFNHKSRCL 121
H+D++P+VA +T WF +RCL
Sbjct: 747 HNDESPIVATTTCWF--FNRCL 806
>BQ150151
Length = 1052
Score = 26.9 bits (58), Expect = 4.7
Identities = 16/43 (37%), Positives = 19/43 (43%)
Frame = -2
Query: 3 RFCPKVSFIFLAITLIFTSFVYSEAQNCRPSGRIRGKKAPPGK 45
R C FI + I +FT + NC P GR A PGK
Sbjct: 349 RTCSTHVFIVI-IVFVFTLLIIKTENNC*PRGRAVTHAA*PGK 224
>TC84000
Length = 709
Score = 26.9 bits (58), Expect = 4.7
Identities = 14/37 (37%), Positives = 18/37 (47%)
Frame = +3
Query: 33 SGRIRGKKAPPGKCNQENDSDCCVQGKMYTTYVCSPS 69
S RIR ++ K + N S C K TT+VC S
Sbjct: 3 SCRIRHEERKNNKV*ESNPSHCSTMTKFITTFVCLSS 113
>AW585537 SP|P03736|VMTH Minor tail protein precursor H. {Bacteriophage
lambda}, partial (9%)
Length = 254
Score = 26.6 bits (57), Expect = 6.2
Identities = 16/64 (25%), Positives = 25/64 (39%), Gaps = 1/64 (1%)
Frame = +1
Query: 126 ISANGRNVVAMVVDECDSRKGCDEQHDYQPPCPNNIVDASKAVWKALGVPKEQW-GGLDI 184
I A R + + + + EQ+ P NN++ K W A + W GL
Sbjct: 31 IDAKSRGLTDRQAEREATEQRLKEQYGDNPLALNNVMSEQKKTWAAEDQLRGNWMAGLKS 210
Query: 185 TWSD 188
WS+
Sbjct: 211 GWSE 222
>TC76512 similar to GP|4959932|gb|AAD34548.1| cystathionine-gamma-synthase
precursor {Glycine max}, partial (90%)
Length = 2103
Score = 26.6 bits (57), Expect = 6.2
Identities = 16/48 (33%), Positives = 23/48 (47%), Gaps = 3/48 (6%)
Frame = -2
Query: 69 SVSTHTKAYLTLNSFQKGGDGGGPSECDKQ---YHSDDTPVVALSTGW 113
+V+T A LTL ++K DGGG C + + VV + GW
Sbjct: 275 AVTTRFGAELTLEIWRKTEDGGGDGSCRGKGGCWEMTTAVVVVVVGGW 132
>TC87487 similar to SP|P52780|SYQ_LUPLU Glutaminyl-tRNA synthetase (EC
6.1.1.18) (Glutamine--tRNA ligase) (GLNRS). [Yellow
lupine], partial (97%)
Length = 2844
Score = 26.2 bits (56), Expect = 8.1
Identities = 11/21 (52%), Positives = 14/21 (66%)
Frame = +2
Query: 109 LSTGWFNHKSRCLNNITISAN 129
LS+GW + SR L +TIS N
Sbjct: 1070 LSSGWDGNHSRLLTQVTISKN 1132
>TC86700 similar to GP|18377835|gb|AAL67104.1 At1g24120/F3I6_4 {Arabidopsis
thaliana}, partial (83%)
Length = 1929
Score = 26.2 bits (56), Expect = 8.1
Identities = 15/40 (37%), Positives = 18/40 (44%), Gaps = 2/40 (5%)
Frame = +2
Query: 6 PKVSFIFLAIT--LIFTSFVYSEAQNCRPSGRIRGKKAPP 43
PK SF F + + F Y CR GR R +K PP
Sbjct: 119 PKKSFCFHSDS*RRFLGFFDYDVVSRCRDDGRSRRRKTPP 238
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.318 0.134 0.435
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 7,861,501
Number of Sequences: 36976
Number of extensions: 120094
Number of successful extensions: 583
Number of sequences better than 10.0: 32
Number of HSP's better than 10.0 without gapping: 576
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 577
length of query: 189
length of database: 9,014,727
effective HSP length: 91
effective length of query: 98
effective length of database: 5,649,911
effective search space: 553691278
effective search space used: 553691278
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 55 (25.8 bits)
Medicago: description of AC135311.8


