

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC135234.5 - phase: 0 /pseudo
(256 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
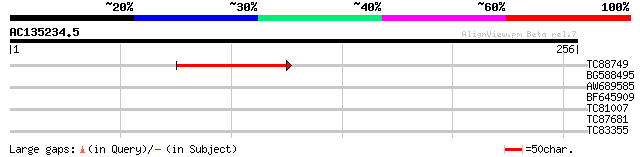
Score E
Sequences producing significant alignments: (bits) Value
TC88749 47 7e-06
BG588495 29 2.0
AW689585 homologue to GP|21166122|gb|A hypothetical protein {Dic... 28 4.5
BF645909 weakly similar to GP|9758286|dbj cytochrome P450-like p... 27 5.9
TC81007 similar to GP|17473902|gb|AAL38369.1 RNA helicase - like... 27 5.9
TC87681 similar to GP|17381090|gb|AAL36357.1 unknown protein {Ar... 27 7.7
TC83355 similar to GP|6016726|gb|AAF01552.1| unknown protein {Ar... 27 7.7
>TC88749
Length = 1221
Score = 47.0 bits (110), Expect = 7e-06
Identities = 22/52 (42%), Positives = 34/52 (65%)
Frame = +2
Query: 76 WREREELLTWVRRQGVKAGFGVCIDKSVLKRPYLTTQSERSGIYKPPKTRKK 127
W+++E++L+ +RRQ +AGF V I +S ++ P L ERSG +K K R K
Sbjct: 551 WKDKEDMLSSIRRQANRAGFTVGIKRSSIQNPMLELMCERSGDHKVSKKRLK 706
>BG588495
Length = 540
Score = 28.9 bits (63), Expect = 2.0
Identities = 11/21 (52%), Positives = 15/21 (71%)
Frame = +3
Query: 77 REREELLTWVRRQGVKAGFGV 97
++R+ELL W R Q KAGF +
Sbjct: 21 KDRDELLEWTRHQANKAGFTI 83
>AW689585 homologue to GP|21166122|gb|A hypothetical protein {Dictyostelium
discoideum}, partial (13%)
Length = 465
Score = 27.7 bits (60), Expect = 4.5
Identities = 21/64 (32%), Positives = 31/64 (47%)
Frame = +1
Query: 153 NDWWLAILCGMHNHDLEEKLQGHLIAGRLSAEEKKKVIDMTKSLTVPRNILTNLKENNKE 212
N W L IL +H+H L+ LQ A + E K ++++T S V + NL K
Sbjct: 25 NQWLLTILSLLHHHLLQSLLQNQ--ACFIMVEVKISMMNLTSSKLV--IFVENLLAKTKT 192
Query: 213 SVTT 216
S+ T
Sbjct: 193 SLCT 204
>BF645909 weakly similar to GP|9758286|dbj cytochrome P450-like protein
{Arabidopsis thaliana}, partial (27%)
Length = 648
Score = 27.3 bits (59), Expect = 5.9
Identities = 30/132 (22%), Positives = 53/132 (39%), Gaps = 11/132 (8%)
Frame = +2
Query: 76 WREREELLTWVRRQGVKAGFGVCIDKSVLKRPYLTTQSERSGIYKPPKTRKKPNLEGIGS 135
++ R LL W + ++ + + + R + T ++ + Y KT K +G
Sbjct: 230 YKNRFRLLDWYTKLLAQSPTNTILVQRLGARRTIVTANQHNVEYVL-KTNFKNFPKGKPF 406
Query: 136 MKCNCPFRLKGFFDKDTNDWWLAILCGMHN-----------HDLEEKLQGHLIAGRLSAE 184
+ F KG F+ D + W + H H LEE+++G LI S
Sbjct: 407 REILSDFLGKGIFNVDDDLWMIQRKLASHEFSLRSINEFIMHTLEEEVKGKLIPLMDSLS 586
Query: 185 EKKKVIDMTKSL 196
+ KV+D + L
Sbjct: 587 IENKVVDFQELL 622
>TC81007 similar to GP|17473902|gb|AAL38369.1 RNA helicase - like protein
{Arabidopsis thaliana}, partial (96%)
Length = 1181
Score = 27.3 bits (59), Expect = 5.9
Identities = 16/53 (30%), Positives = 29/53 (54%), Gaps = 8/53 (15%)
Frame = -3
Query: 100 DKSVLKRPYLTTQSERS--------GIYKPPKTRKKPNLEGIGSMKCNCPFRL 144
++SV+ + YL S+ + GI KP R ++EG GS++C+C ++
Sbjct: 483 NRSVINQTYLHLSSKYTIFECIRIIGIPKPVH*R---SIEGFGSLRCHCDMKI 334
>TC87681 similar to GP|17381090|gb|AAL36357.1 unknown protein {Arabidopsis
thaliana}, partial (29%)
Length = 796
Score = 26.9 bits (58), Expect = 7.7
Identities = 21/65 (32%), Positives = 31/65 (47%)
Frame = +2
Query: 152 TNDWWLAILCGMHNHDLEEKLQGHLIAGRLSAEEKKKVIDMTKSLTVPRNILTNLKENNK 211
T W L IL +H+H L+ LQ A + E K ++++T S V + NL K
Sbjct: 20 TQPWLLTILSLLHHHLLQSLLQNQ--ACFIMVEVKISMMNLTSSKLV--IFVENLLAKTK 187
Query: 212 ESVTT 216
S+ T
Sbjct: 188 TSLCT 202
>TC83355 similar to GP|6016726|gb|AAF01552.1| unknown protein {Arabidopsis
thaliana}, partial (68%)
Length = 1032
Score = 26.9 bits (58), Expect = 7.7
Identities = 12/41 (29%), Positives = 20/41 (48%)
Frame = +2
Query: 156 WLAILCGMHNHDLEEKLQGHLIAGRLSAEEKKKVIDMTKSL 196
W I+ G NH + + H+ RL KKV+ ++K +
Sbjct: 110 WGMIILGTDNHQESQVSRTHIQISRLCTSVSKKVLVLSKKI 232
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.318 0.136 0.410
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 8,046,075
Number of Sequences: 36976
Number of extensions: 107688
Number of successful extensions: 576
Number of sequences better than 10.0: 14
Number of HSP's better than 10.0 without gapping: 563
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 576
length of query: 256
length of database: 9,014,727
effective HSP length: 94
effective length of query: 162
effective length of database: 5,538,983
effective search space: 897315246
effective search space used: 897315246
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 57 (26.6 bits)
Medicago: description of AC135234.5


