

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC135233.1 + phase: 0
(87 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
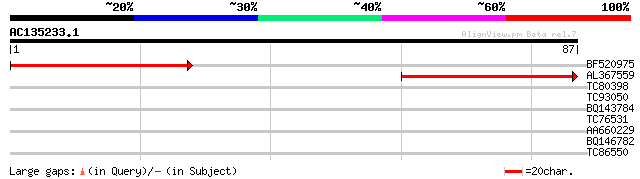
BF520975 similar to GP|9757942|dbj GAMM1 protein-like {Arabidops... 60 1e-10
AL367559 similar to GP|12324441|gb| unknown protein; 11418-9170 ... 57 1e-09
TC80398 similar to GP|20521392|dbj|BAB91903. putative FtsH prote... 28 0.62
TC93050 similar to GP|8886934|gb|AAF80620.1| F2D10.25 {Arabidops... 28 0.81
BQ143784 weakly similar to SP|P33485|VNUA Probable nuclear antig... 27 1.4
TC76531 similar to SP|P13915|CVCA_PEA Convicilin precursor. [Gar... 26 2.3
AA660229 26 3.1
BQ146782 25 6.8
TC86550 similar to GP|15810601|gb|AAL07188.1 unknown protein {Ar... 24 8.9
>BF520975 similar to GP|9757942|dbj GAMM1 protein-like {Arabidopsis
thaliana}, partial (45%)
Length = 547
Score = 60.5 bits (145), Expect = 1e-10
Identities = 28/28 (100%), Positives = 28/28 (100%)
Frame = +1
Query: 1 MKIDPPIKYVLYEDERSKQWRVQAVSVS 28
MKIDPPIKYVLYEDERSKQWRVQAVSVS
Sbjct: 454 MKIDPPIKYVLYEDERSKQWRVQAVSVS 537
>AL367559 similar to GP|12324441|gb| unknown protein; 11418-9170
{Arabidopsis thaliana}, partial (7%)
Length = 328
Score = 57.0 bits (136), Expect = 1e-09
Identities = 27/27 (100%), Positives = 27/27 (100%)
Frame = +3
Query: 61 FVHMSGFIGGNQTFEGALAMAKAALKM 87
FVHMSGFIGGNQTFEGALAMAKAALKM
Sbjct: 3 FVHMSGFIGGNQTFEGALAMAKAALKM 83
>TC80398 similar to GP|20521392|dbj|BAB91903. putative FtsH protease {Oryza
sativa (japonica cultivar-group)}, partial (26%)
Length = 981
Score = 28.1 bits (61), Expect = 0.62
Identities = 24/90 (26%), Positives = 37/90 (40%), Gaps = 6/90 (6%)
Frame = +2
Query: 4 DPPIKYVLYEDERSKQWRVQAVS------VSPDRFESRKALPSQWRGLRDDILSKESGIP 57
DP +E + S A+S V DR + + L + RG+ + + +ESGI
Sbjct: 395 DPEAVIRAFESQPSLHTNSAALSEYVKALVKVDRLDESELLQTLRRGISNSVREEESGIG 574
Query: 58 GCVFVHMSGFIGGNQTFEGALAMAKAALKM 87
G + G T +G L A A + M
Sbjct: 575 GLAVLRNV----GKSTKDGILGTAGAPIHM 652
>TC93050 similar to GP|8886934|gb|AAF80620.1| F2D10.25 {Arabidopsis
thaliana}, partial (4%)
Length = 800
Score = 27.7 bits (60), Expect = 0.81
Identities = 12/39 (30%), Positives = 21/39 (53%)
Frame = -2
Query: 27 VSPDRFESRKALPSQWRGLRDDILSKESGIPGCVFVHMS 65
++ E K +Q+ GL ++L +SG+ G F+H S
Sbjct: 757 LNESNLEKSKVFSNQFLGLCGELLELKSGVLGTEFIHWS 641
>BQ143784 weakly similar to SP|P33485|VNUA Probable nuclear antigen. [strain
Kaplan PRV] {Pseudorabies virus}, partial (2%)
Length = 1022
Score = 26.9 bits (58), Expect = 1.4
Identities = 11/22 (50%), Positives = 17/22 (77%), Gaps = 1/22 (4%)
Frame = -3
Query: 41 QWRGLRDDILSKE-SGIPGCVF 61
+W G+R D +S+ +G+PGCVF
Sbjct: 345 RWAGVRRDGVSRGVAGVPGCVF 280
>TC76531 similar to SP|P13915|CVCA_PEA Convicilin precursor. [Garden pea]
{Pisum sativum}, partial (86%)
Length = 2217
Score = 26.2 bits (56), Expect = 2.3
Identities = 13/46 (28%), Positives = 19/46 (41%)
Frame = +2
Query: 13 EDERSKQWRVQAVSVSPDRFESRKALPSQWRGLRDDILSKESGIPG 58
+++R QWR + E +K P QWR R + PG
Sbjct: 221 QEQRPGQWRPSHEREEDEEDEHQKRRPGQWRPTRGGRQEGQEQRPG 358
>AA660229
Length = 374
Score = 25.8 bits (55), Expect = 3.1
Identities = 9/14 (64%), Positives = 11/14 (78%)
Frame = +2
Query: 9 YVLYEDERSKQWRV 22
YVLY+ RS+ WRV
Sbjct: 242 YVLYQSNRSRSWRV 283
>BQ146782
Length = 649
Score = 24.6 bits (52), Expect = 6.8
Identities = 17/37 (45%), Positives = 22/37 (58%), Gaps = 1/37 (2%)
Frame = +3
Query: 21 RVQAVSVSPDRFESRKALPSQWRGLRDDILSKE-SGI 56
R+Q S + + ALP++ R L D ILSKE SGI
Sbjct: 222 RLQESGSSQQKRQKLSALPTEERILYDLILSKENSGI 332
>TC86550 similar to GP|15810601|gb|AAL07188.1 unknown protein {Arabidopsis
thaliana}, partial (67%)
Length = 1240
Score = 24.3 bits (51), Expect = 8.9
Identities = 10/14 (71%), Positives = 13/14 (92%)
Frame = -1
Query: 25 VSVSPDRFESRKAL 38
VS+SP+RF+SR AL
Sbjct: 832 VSLSPERFDSRAAL 791
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.320 0.136 0.409
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 2,554,102
Number of Sequences: 36976
Number of extensions: 27250
Number of successful extensions: 107
Number of sequences better than 10.0: 18
Number of HSP's better than 10.0 without gapping: 106
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 107
length of query: 87
length of database: 9,014,727
effective HSP length: 63
effective length of query: 24
effective length of database: 6,685,239
effective search space: 160445736
effective search space used: 160445736
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 51 (24.3 bits)
Medicago: description of AC135233.1


