

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC135232.7 + phase: 0
(221 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
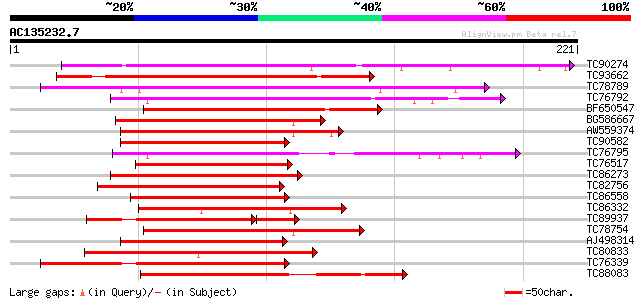
Score E
Sequences producing significant alignments: (bits) Value
TC90274 similar to GP|10177219|dbj|BAB10294. contains similarity... 169 6e-43
TC93662 similar to GP|10177219|dbj|BAB10294. contains similarity... 125 2e-29
TC78789 homologue to PIR|T48580|T48580 hypothetical protein T31B... 122 1e-28
TC76792 similar to GP|3264767|gb|AAC24587.1| AP2 domain containi... 111 2e-25
BF650547 similar to GP|9758388|dbj| AP2 domain transcription fac... 111 3e-25
BG586667 similar to GP|17381278|gb| AT4g34410/F10M10_180 {Arabid... 110 4e-25
AW559374 similar to GP|17381278|gb| AT4g34410/F10M10_180 {Arabid... 110 4e-25
TC90582 similar to GP|17381278|gb|AAL36057.1 AT4g34410/F10M10_18... 107 3e-24
TC76795 similar to GP|18496063|emb|CAD21849. ethylene responsive... 105 1e-23
TC76517 similar to GP|11994632|dbj|BAB02769. AP2 domain transcri... 102 1e-22
TC86273 similar to SP|O80340|ERF4_ARATH Ethylene responsive elem... 101 3e-22
TC82756 similar to GP|17381278|gb|AAL36057.1 AT4g34410/F10M10_18... 100 3e-22
TC86558 similar to GP|20805105|dbj|BAB92777. contains ESTs AU065... 100 4e-22
TC86332 similar to GP|7528276|gb|AAF63205.1| AP2-related transcr... 100 6e-22
TC89937 similar to GP|9758388|dbj|BAB08875.1 AP2 domain transcri... 88 9e-22
TC78754 homologue to GP|4099921|gb|AAD09248.1| EREBP-3 homolog {... 99 1e-21
AJ498314 similar to GP|17381278|gb| AT4g34410/F10M10_180 {Arabid... 99 1e-21
TC80833 weakly similar to PIR|D86175|D86175 hypothetical protein... 98 3e-21
TC76339 homologue to GP|4099914|gb|AAD00708.1| ethylene-responsi... 97 5e-21
TC88083 similar to GP|4587373|dbj|BAA76734.1 ethylene responsive... 97 6e-21
>TC90274 similar to GP|10177219|dbj|BAB10294. contains similarity to AP2
domain transcription factor~gene_id:MPF21.9 {Arabidopsis
thaliana}, partial (32%)
Length = 750
Score = 169 bits (429), Expect = 6e-43
Identities = 105/222 (47%), Positives = 132/222 (59%), Gaps = 22/222 (9%)
Frame = +3
Query: 21 TTTTGRSQVEEQIYEYRTDNNNVKNEDQPKRKYRGVRQRPWGKWAAEIRDPFKATRVWLG 80
TTT ++ +YEY+ N V+ E + KRKYRGVRQRPWGKWAAEIRDPFKA RVWLG
Sbjct: 72 TTTRITREMGNAVYEYKRTENVVR-EGEAKRKYRGVRQRPWGKWAAEIRDPFKAARVWLG 248
Query: 81 TFETAEDAAKAYDQASLRFRGNKAKLNFPENVKLKQ--QQSTPTHLNISHSNSALLSHQP 138
TFETAE AA+AYD+A+LRFRGNKAKLNFPENV L+ +T T N+S+S S+++S
Sbjct: 249 TFETAEAAARAYDEAALRFRGNKAKLNFPENVTLRNPPPATTTTQWNVSNSPSSIVS--I 422
Query: 139 RTDPDPIVHNETL-HTLQSSNKYYDYFNGQNFP---------------MASLQTSVSPSA 182
T DP+VH ++ Q S +YD F P MAS S S S
Sbjct: 423 TTSTDPVVHTRPFPNSSQHSTNFYDRFQFSGIPPARNVYDDNVIRASSMASHLQSSSSSP 602
Query: 183 SYSSSYTTTTFASSFSSPQTTSS-IPSGLTAWPS---GSSPS 220
+ S S +++ F SS + SS + L W + SSPS
Sbjct: 603 ASSLSSSSSAFVSSMQGNTSVSSYFSTQLPPWSTSDQSSSPS 728
>TC93662 similar to GP|10177219|dbj|BAB10294. contains similarity to AP2
domain transcription factor~gene_id:MPF21.9 {Arabidopsis
thaliana}, partial (31%)
Length = 555
Score = 125 bits (313), Expect = 2e-29
Identities = 65/124 (52%), Positives = 82/124 (65%)
Frame = +3
Query: 19 TVTTTTGRSQVEEQIYEYRTDNNNVKNEDQPKRKYRGVRQRPWGKWAAEIRDPFKATRVW 78
T ++T+ +Q E N + + +R+YRGVRQRPWGKWAAEIRDP KA RVW
Sbjct: 171 TTSSTSNTTQSSEA-----ASTTNEEETGERRRRYRGVRQRPWGKWAAEIRDPHKAARVW 335
Query: 79 LGTFETAEDAAKAYDQASLRFRGNKAKLNFPENVKLKQQQSTPTHLNISHSNSALLSHQP 138
LGTFETAE AA+AYD+A+LRFRGN+AKLNFPENV + T +N S S+ + +H
Sbjct: 336 LGTFETAEAAARAYDEAALRFRGNRAKLNFPENVPASATRPT-FPVNTSTSSVSPATHYS 512
Query: 139 RTDP 142
P
Sbjct: 513 PAPP 524
>TC78789 homologue to PIR|T48580|T48580 hypothetical protein T31B5.150 -
Arabidopsis thaliana, partial (33%)
Length = 1112
Score = 122 bits (305), Expect = 1e-28
Identities = 79/204 (38%), Positives = 105/204 (50%), Gaps = 29/204 (14%)
Frame = +1
Query: 13 SSNWTNTVTTTTGRSQVEEQIYEYRTDNNN-----------VKNEDQP-----------K 50
+SNW +T S + + I T++NN V E QP K
Sbjct: 172 NSNWCPDTDLSTMVSALSQVIGTSSTNHNNPQMAQSISTTMVNEESQPPQPLLDQETKKK 351
Query: 51 RKYRGVRQRPWGKWAAEIRDPFKATRVWLGTFETAEDAAKAYDQASLRFRGNKAKLNFPE 110
YRGVRQRPWGKWAAEIRDP KA RVWLGTF+TAEDAA AYD+A+L+F+G KAKLNFPE
Sbjct: 352 PHYRGVRQRPWGKWAAEIRDPKKAARVWLGTFDTAEDAALAYDKAALKFKGTKAKLNFPE 531
Query: 111 NVKLKQQQSTPTHLNISHSNSALLSHQPRTDPD-----PIVHNETLHTLQSSNKYYDYFN 165
V S+ + I S+ SH + P+ I+ ++ +S+ +
Sbjct: 532 RVVQCNSYSSTANNAIQQSDYVSNSHDQQVFPNLFQYAQILESDAEFPYYASHLFNQQQQ 711
Query: 166 GQNFPMA--SLQTSVSPSASYSSS 187
G P +L+ S + S+S S
Sbjct: 712 GDRHPPGE*TLEISCNQEISHSHS 783
>TC76792 similar to GP|3264767|gb|AAC24587.1| AP2 domain containing protein
{Prunus armeniaca}, partial (49%)
Length = 1729
Score = 111 bits (278), Expect = 2e-25
Identities = 68/171 (39%), Positives = 93/171 (53%), Gaps = 17/171 (9%)
Frame = +3
Query: 40 NNNVKNEDQPKRK----YRGVRQRPWGKWAAEIRDPFKATRVWLGTFETAEDAAKAYDQA 95
N+N + E KRK YRG+RQRPWGKWAAEIRDP K RVWLGTF TAE+AA+AYD
Sbjct: 477 NSNQQAEKDSKRKRKNQYRGIRQRPWGKWAAEIRDPRKGVRVWLGTFNTAEEAARAYDAE 656
Query: 96 SLRFRGNKAKLNFPENVKLKQQQSTPTHLNISHSNSALLSHQPRTDPDPIVHNETLHTLQ 155
+ R RGNKAK+NFPE + + + + N L S +P + +E +
Sbjct: 657 ARRIRGNKAKVNFPEEAPVASSKRLKKNPEMQLLNDNLNSFKPNGN-QMFNFSENMENYY 833
Query: 156 S-----------SNKYYDY--FNGQNFPMASLQTSVSPSASYSSSYTTTTF 193
S +N+Y D F+G + TS +A +SS +++ +F
Sbjct: 834 SPMDQVEQKPLVNNQYADIGAFSGNGVQL----TSADVNAYFSSEHSSNSF 974
>BF650547 similar to GP|9758388|dbj| AP2 domain transcription factor-like
{Arabidopsis thaliana}, partial (24%)
Length = 418
Score = 111 bits (277), Expect = 3e-25
Identities = 55/93 (59%), Positives = 65/93 (69%)
Frame = -1
Query: 53 YRGVRQRPWGKWAAEIRDPFKATRVWLGTFETAEDAAKAYDQASLRFRGNKAKLNFPENV 112
YRGVRQRPWGKWAAEIRDP K RVWLGTFETAE AA AYD+A+LRF+GNKAKLNFP +
Sbjct: 385 YRGVRQRPWGKWAAEIRDPKKGARVWLGTFETAEAAATAYDEAALRFKGNKAKLNFPRLI 206
Query: 113 KLKQQQSTPTHLNISHSNSALLSHQPRTDPDPI 145
P N ++++S+ S P P+
Sbjct: 205 SNLPSDPKPPR-NKNNASSSSSSSSQLAPPKPL 110
>BG586667 similar to GP|17381278|gb| AT4g34410/F10M10_180 {Arabidopsis
thaliana}, partial (28%)
Length = 740
Score = 110 bits (275), Expect = 4e-25
Identities = 52/85 (61%), Positives = 65/85 (76%), Gaps = 3/85 (3%)
Frame = +3
Query: 42 NVKNEDQPKRKYRGVRQRPWGKWAAEIRDPFKATRVWLGTFETAEDAAKAYDQASLRFRG 101
N K + + K+KYRGVRQRPWGKWAAEIRDP +A RVWLGTF TAE+AA+AYD A++ FRG
Sbjct: 96 NKKKQIRAKKKYRGVRQRPWGKWAAEIRDPRRAVRVWLGTFTTAEEAARAYDNAAIEFRG 275
Query: 102 NKAKLNFP---ENVKLKQQQSTPTH 123
+AKLNFP E++K ++ H
Sbjct: 276 PRAKLNFPLVDESLKHVEEPEVIVH 350
>AW559374 similar to GP|17381278|gb| AT4g34410/F10M10_180 {Arabidopsis
thaliana}, partial (27%)
Length = 732
Score = 110 bits (275), Expect = 4e-25
Identities = 54/89 (60%), Positives = 67/89 (74%), Gaps = 2/89 (2%)
Frame = +3
Query: 44 KNEDQPKRKYRGVRQRPWGKWAAEIRDPFKATRVWLGTFETAEDAAKAYDQASLRFRGNK 103
+ + + K+KYRGVRQRPWGKWAAEIRDP +ATRVWLGTF TAE+AA+AYD A++ FRG +
Sbjct: 273 QKQKRAKKKYRGVRQRPWGKWAAEIRDPRRATRVWLGTFSTAEEAARAYDNAAIEFRGPR 452
Query: 104 AKLNFP-ENVKLKQQQSTPTHL-NISHSN 130
AKLNFP + LKQ + L N+ N
Sbjct: 453 AKLNFPMVDDSLKQSEVVAPDLENVKDEN 539
>TC90582 similar to GP|17381278|gb|AAL36057.1 AT4g34410/F10M10_180
{Arabidopsis thaliana}, partial (29%)
Length = 979
Score = 107 bits (268), Expect = 3e-24
Identities = 47/66 (71%), Positives = 57/66 (86%)
Frame = +2
Query: 44 KNEDQPKRKYRGVRQRPWGKWAAEIRDPFKATRVWLGTFETAEDAAKAYDQASLRFRGNK 103
+ + + K+KYRGVRQRPWGKWAAEIRDP +A RVWLGTF TAE+AA+AYD A++ FRG +
Sbjct: 314 QKQKRAKKKYRGVRQRPWGKWAAEIRDPRRAVRVWLGTFTTAEEAARAYDNAAIEFRGPR 493
Query: 104 AKLNFP 109
AKLNFP
Sbjct: 494 AKLNFP 511
>TC76795 similar to GP|18496063|emb|CAD21849. ethylene responsive element
binding protein {Fagus sylvatica}, partial (32%)
Length = 966
Score = 105 bits (263), Expect = 1e-23
Identities = 70/171 (40%), Positives = 95/171 (54%), Gaps = 12/171 (7%)
Frame = +3
Query: 41 NNVKNEDQPKRK----YRGVRQRPWGKWAAEIRDPFKATRVWLGTFETAEDAAKAYDQAS 96
+N + E + KRK YRG+RQRPWGKWAAEIRDP K RVWLGTF TAE+AA+AYD +
Sbjct: 318 SNEQGEKELKRKRKNQYRGIRQRPWGKWAAEIRDPRKGVRVWLGTFNTAEEAARAYDAEA 497
Query: 97 LRFRGNKAKLNFPENVKLKQQQSTPTHLNISHSNSALLSHQPRTDPDPIVHNETLHTLQS 156
R RG KAK+NFPE N S S + +T+ + + ++ L++ +
Sbjct: 498 RRIRGKKAKVNFPEEAP-----------NAS-------SKRLKTNSETQLLDKNLNSFKC 623
Query: 157 SN--KYYDYFNG-QNFPMASLQ-TSVSPSA----SYSSSYTTTTFASSFSS 199
N YY N + P+ + Q +V P A +SS T F+S +SS
Sbjct: 624 ENIENYYSPVNQVEQRPLVNAQCANVGPFAGNGVQLASSDVTAYFSSEYSS 776
>TC76517 similar to GP|11994632|dbj|BAB02769. AP2 domain transcription
factor RAP2.3 {Arabidopsis thaliana}, partial (33%)
Length = 1290
Score = 102 bits (254), Expect = 1e-22
Identities = 45/61 (73%), Positives = 52/61 (84%)
Frame = +3
Query: 50 KRKYRGVRQRPWGKWAAEIRDPFKATRVWLGTFETAEDAAKAYDQASLRFRGNKAKLNFP 109
K YRG+RQRPWGKWAAEIRDP + RVWLGTF TAE+AA+AYD A+ R RG+KAKLNFP
Sbjct: 567 KNVYRGIRQRPWGKWAAEIRDPQQGVRVWLGTFSTAEEAARAYDTAAKRIRGDKAKLNFP 746
Query: 110 E 110
+
Sbjct: 747 D 749
>TC86273 similar to SP|O80340|ERF4_ARATH Ethylene responsive element binding
factor 4 (AtERF4). [Mouse-ear cress] {Arabidopsis
thaliana}, partial (43%)
Length = 1171
Score = 101 bits (251), Expect = 3e-22
Identities = 45/75 (60%), Positives = 62/75 (82%)
Frame = +3
Query: 40 NNNVKNEDQPKRKYRGVRQRPWGKWAAEIRDPFKATRVWLGTFETAEDAAKAYDQASLRF 99
N + + + Q + +YRGVR+RPWG++AAEIRDP K TRVWLGTF+TAE+AA+AYD+A+ F
Sbjct: 141 NPSAQAQAQKEIRYRGVRKRPWGRYAAEIRDPGKKTRVWLGTFDTAEEAARAYDKAAREF 320
Query: 100 RGNKAKLNFPENVKL 114
RG+KAK NFP +++
Sbjct: 321 RGSKAKTNFPTPLEI 365
>TC82756 similar to GP|17381278|gb|AAL36057.1 AT4g34410/F10M10_180
{Arabidopsis thaliana}, partial (35%)
Length = 630
Score = 100 bits (250), Expect = 3e-22
Identities = 46/73 (63%), Positives = 54/73 (73%)
Frame = +3
Query: 35 EYRTDNNNVKNEDQPKRKYRGVRQRPWGKWAAEIRDPFKATRVWLGTFETAEDAAKAYDQ 94
E + K + KYRGVRQRPWGKWAAEIRDP +A RVWLGTF TAE+AA+AYD
Sbjct: 210 EEKKQKQKQKQKRAKNNKYRGVRQRPWGKWAAEIRDPRRAVRVWLGTFTTAEEAARAYDN 389
Query: 95 ASLRFRGNKAKLN 107
A++ FRG +AKLN
Sbjct: 390 AAIEFRGPRAKLN 428
>TC86558 similar to GP|20805105|dbj|BAB92777. contains ESTs AU065506(C0784)
AU083422(C0784)~unknown protein, partial (47%)
Length = 1346
Score = 100 bits (249), Expect = 4e-22
Identities = 44/62 (70%), Positives = 51/62 (81%)
Frame = +1
Query: 48 QPKRKYRGVRQRPWGKWAAEIRDPFKATRVWLGTFETAEDAAKAYDQASLRFRGNKAKLN 107
Q K YRG+RQRPWGKWAAEIRDP K RVWLGTF TAE+AA+AYD+ + + RG KAK+N
Sbjct: 388 QRKNLYRGIRQRPWGKWAAEIRDPRKGVRVWLGTFNTAEEAARAYDKEARKIRGKKAKVN 567
Query: 108 FP 109
FP
Sbjct: 568 FP 573
>TC86332 similar to GP|7528276|gb|AAF63205.1| AP2-related transcription
factor {Mesembryanthemum crystallinum}, partial (49%)
Length = 1349
Score = 100 bits (248), Expect = 6e-22
Identities = 52/90 (57%), Positives = 65/90 (71%), Gaps = 9/90 (10%)
Frame = +1
Query: 51 RKYRGVRQRPWGKWAAEIRDPFK-ATRVWLGTFETAEDAAKAYDQASLRFRGNKAKLNF- 108
+ YRGVRQRPWGK+AAEIRDP K RVWLGTFETAEDAA AYD+A+ R RG++A LNF
Sbjct: 535 KHYRGVRQRPWGKFAAEIRDPAKNGARVWLGTFETAEDAALAYDRAAYRMRGSRALLNFP 714
Query: 109 -------PENVKLKQQQSTPTHLNISHSNS 131
P+ V++ ++S+P + S SNS
Sbjct: 715 LRVNSGEPDPVRIASKRSSPERSSSSESNS 804
>TC89937 similar to GP|9758388|dbj|BAB08875.1 AP2 domain transcription
factor-like {Arabidopsis thaliana}, partial (41%)
Length = 578
Score = 88.2 bits (217), Expect(2) = 9e-22
Identities = 44/66 (66%), Positives = 49/66 (73%)
Frame = +2
Query: 31 EQIYEYRTDNNNVKNEDQPKRKYRGVRQRPWGKWAAEIRDPFKATRVWLGTFETAEDAAK 90
EQ + D NV+ +R YRGVRQRPWGKWAAEIRDP KA RVWLGTFETAE AA
Sbjct: 341 EQSQPPQQDQGNVR-----RRHYRGVRQRPWGKWAAEIRDPKKAARVWLGTFETAEAAAV 505
Query: 91 AYDQAS 96
AYD+A+
Sbjct: 506 AYDEAA 523
Score = 32.0 bits (71), Expect(2) = 9e-22
Identities = 13/17 (76%), Positives = 16/17 (93%)
Frame = +3
Query: 97 LRFRGNKAKLNFPENVK 113
LRF+G+KAKLNFPE V+
Sbjct: 525 LRFKGSKAKLNFPERVQ 575
>TC78754 homologue to GP|4099921|gb|AAD09248.1| EREBP-3 homolog
{Stylosanthes hamata}, partial (46%)
Length = 1038
Score = 99.4 bits (246), Expect = 1e-21
Identities = 48/89 (53%), Positives = 62/89 (68%), Gaps = 3/89 (3%)
Frame = +3
Query: 53 YRGVRQRPWGKWAAEIRDPFKATRVWLGTFETAEDAAKAYDQASLRFRGNKAKLNFP--- 109
+RGVR+RPWG++AAEIRDP K +RVWLGTF+TAE+AA+AYD A+ +FRG KAK NFP
Sbjct: 153 FRGVRKRPWGRYAAEIRDPGKKSRVWLGTFDTAEEAARAYDSAARQFRGPKAKTNFPFPI 332
Query: 110 ENVKLKQQQSTPTHLNISHSNSALLSHQP 138
EN+K+ + T + S A P
Sbjct: 333 ENIKIHSPSQSSTVESSSRDRDAAADSSP 419
>AJ498314 similar to GP|17381278|gb| AT4g34410/F10M10_180 {Arabidopsis
thaliana}, partial (22%)
Length = 651
Score = 99.0 bits (245), Expect = 1e-21
Identities = 44/65 (67%), Positives = 57/65 (87%)
Frame = +3
Query: 44 KNEDQPKRKYRGVRQRPWGKWAAEIRDPFKATRVWLGTFETAEDAAKAYDQASLRFRGNK 103
K++ + K+KYRGVRQR WGKW AEIRDP +ATRV LGTF+TAE+AA+AYDQA+++F G +
Sbjct: 417 KDKGENKKKYRGVRQRAWGKWVAEIRDPKRATRV*LGTFQTAENAARAYDQAAIKFHGAR 596
Query: 104 AKLNF 108
AK+NF
Sbjct: 597 AKINF 611
>TC80833 weakly similar to PIR|D86175|D86175 hypothetical protein [imported]
- Arabidopsis thaliana, partial (60%)
Length = 709
Score = 97.8 bits (242), Expect = 3e-21
Identities = 46/92 (50%), Positives = 63/92 (68%), Gaps = 1/92 (1%)
Frame = +1
Query: 30 EEQIYEYRTDNNNVKNEDQPKRKYRGVRQRPWGKWAAEIRDPF-KATRVWLGTFETAEDA 88
+ ++ T+ N N + + KYRG+R+RPWGK+AAEIRDP K TR+WLGTF+TAE A
Sbjct: 109 DHKLVSNSTNGNGNGNGNSDQIKYRGIRRRPWGKFAAEIRDPTRKGTRIWLGTFDTAEQA 288
Query: 89 AKAYDQASLRFRGNKAKLNFPENVKLKQQQST 120
A+AYD A+ FRG++A LNFP + S+
Sbjct: 289 ARAYDAAAFHFRGHRAILNFPNEYQAPNSSSS 384
>TC76339 homologue to GP|4099914|gb|AAD00708.1| ethylene-responsive element
binding protein homolog {Stylosanthes hamata}, partial
(37%)
Length = 1110
Score = 97.1 bits (240), Expect = 5e-21
Identities = 49/97 (50%), Positives = 68/97 (69%)
Frame = +2
Query: 13 SSNWTNTVTTTTGRSQVEEQIYEYRTDNNNVKNEDQPKRKYRGVRQRPWGKWAAEIRDPF 72
SS+ T++ T ++ +I + +NNN + +RGVR+RPWG++AAEIRDP
Sbjct: 38 SSSPTSSSMVTKTKNAANLKIKKPSNNNNNAT-----EIHFRGVRKRPWGRYAAEIRDPG 202
Query: 73 KATRVWLGTFETAEDAAKAYDQASLRFRGNKAKLNFP 109
K +RVWLGTF+TAE+AA+AYD A+ +FRG KAK NFP
Sbjct: 203 KKSRVWLGTFDTAEEAARAYDNAARQFRGPKAKTNFP 313
>TC88083 similar to GP|4587373|dbj|BAA76734.1 ethylene responsive element
binding factor {Nicotiana tabacum}, partial (46%)
Length = 1214
Score = 96.7 bits (239), Expect = 6e-21
Identities = 49/104 (47%), Positives = 67/104 (64%)
Frame = +1
Query: 52 KYRGVRQRPWGKWAAEIRDPFKATRVWLGTFETAEDAAKAYDQASLRFRGNKAKLNFPEN 111
+YRGVR+RPWG++AAEIRDP+K TRVWLGT++TAE AA AYD A+++FR +KAK NFP
Sbjct: 154 RYRGVRKRPWGRFAAEIRDPWKKTRVWLGTYDTAEQAALAYDSAAIKFRASKAKTNFP-- 327
Query: 112 VKLKQQQSTPTHLNISHSNSALLSHQPRTDPDPIVHNETLHTLQ 155
P H+ + + +A ++ Q P V N L+
Sbjct: 328 --------IPEHILAAAAETAAVAAQ-NPQPPAAVDNRPAEVLK 432
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.310 0.124 0.366
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 7,172,016
Number of Sequences: 36976
Number of extensions: 102859
Number of successful extensions: 1104
Number of sequences better than 10.0: 189
Number of HSP's better than 10.0 without gapping: 1007
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1062
length of query: 221
length of database: 9,014,727
effective HSP length: 92
effective length of query: 129
effective length of database: 5,612,935
effective search space: 724068615
effective search space used: 724068615
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.8 bits)
S2: 56 (26.2 bits)
Medicago: description of AC135232.7


