

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC135231.7 + phase: 0 /pseudo
(943 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
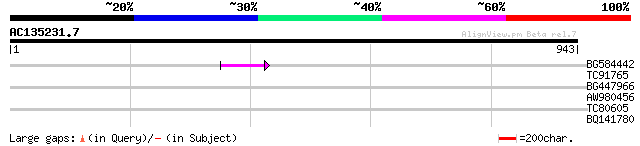
BG584442 56 7e-08
TC91765 weakly similar to GP|19881779|gb|AAM01180.1 Putative ret... 35 0.005
BG447966 weakly similar to PIR|G96509|G96 protein F27F5.21 [impo... 32 0.88
AW980456 30 4.4
TC80605 similar to PIR|S56716|S56716 protein kinase SPK-3 (EC 2.... 30 5.7
BQ141780 similar to SP|P16059|PSBP Oxygen-evolving enhancer prot... 29 9.7
>BG584442
Length = 775
Score = 55.8 bits (133), Expect = 7e-08
Identities = 34/82 (41%), Positives = 47/82 (56%)
Frame = +3
Query: 351 ENQFLEW*VSL*SRSRGYDKICFAGYSFLCDEYLSASHYLTRLN*KDDELLLVGQR*NNT 410
+N FLE + + S RG D +C + FLCDEY S S + +R N*KD E +G +
Sbjct: 381 KN*FLEKQMFVKSYVRGND*VCTPKHIFLCDEYFSPSKFSSR*N*KDYEYFFMGSCWRKS 560
Query: 411 KRDSLDELGEAFSTKDSRRYGF 432
+R++LD LGE T R+GF
Sbjct: 561 QRNALDVLGEIVCT*KLWRHGF 626
>TC91765 weakly similar to GP|19881779|gb|AAM01180.1 Putative retroelement
{Oryza sativa (japonica cultivar-group)}, partial (1%)
Length = 625
Score = 34.7 bits (78), Expect = 0.18
Identities = 27/86 (31%), Positives = 43/86 (49%)
Frame = +2
Query: 266 QS*QGSSTCYEAYSIHL*ISFWSSDQFTKV*NILQS*CS**PQS*HYRYSRRPTCVGHR* 325
QS*+ S YE + + W S +++ +ILQ CS* + H+ Y VG+ *
Sbjct: 311 QS*RASCANYEKHPYSVRGRLW*SY*SSQIIDILQQECS*HFKDFHHIYFGGSIYVGYM* 490
Query: 326 IFGLTFYDRSRSKCYFCLHQGSCLAE 351
+FG + +DR R F + G +A+
Sbjct: 491 VFGTSVHDRQR*NNNFFFY*GWSVAK 568
Score = 32.0 bits (71), Expect(2) = 0.005
Identities = 16/41 (39%), Positives = 25/41 (60%)
Frame = +1
Query: 184 YVRRISGLFCACKWYTGRSYYSRAGCPSRGSSLSVLVYYLC 224
YV I L C+C G SY +++ +RGSSL + +++LC
Sbjct: 100 YVCGIK*LLCSC*Q*CGGSYNTKSRFTTRGSSLPLYLHHLC 222
Score = 26.9 bits (58), Expect(2) = 0.005
Identities = 18/31 (58%), Positives = 20/31 (64%)
Frame = +3
Query: 153 Y*QSL*SYGLELFEGCFE*DGFS*PLDSLDV 183
Y*QSL*S L L G + DG *P+D LDV
Sbjct: 6 Y*QSL*SSRLRLS*GNHDQDGV**PVDLLDV 98
>BG447966 weakly similar to PIR|G96509|G96 protein F27F5.21 [imported] -
Arabidopsis thaliana, partial (7%)
Length = 687
Score = 32.3 bits (72), Expect = 0.88
Identities = 23/83 (27%), Positives = 36/83 (42%)
Frame = +1
Query: 65 HCPYPKRFYAG*HEGLAPYSTL*CVIQNHFESVSKPVEESAA*MHFRQSIRLRPRTLDFR 124
HCPYP R + +GL P L C +NH + S+ + + + R RPR R
Sbjct: 439 HCPYP*RKESEYSKGLPPN*PLQCCHENHHQGYSQQS*ADSPGCN*CGAERFRPRQAHHR 618
Query: 125 *CYGSN*GFTFYESQDARGRQIC 147
C S F + + + ++ C
Sbjct: 619 QCTDSMECFHWLKXKKKGKKESC 687
>AW980456
Length = 779
Score = 30.0 bits (66), Expect = 4.4
Identities = 22/59 (37%), Positives = 30/59 (50%)
Frame = -1
Query: 582 MESGKAQTLLCS*CLQVMCIRTD*FIL*LATGVLVWYLESKSPTESQEFNLVHVSGLFT 640
+E KA +LC CL +C T F L + LV LE+ S + + +L V GLFT
Sbjct: 197 LEG*KAWEILCKECLSFLCRGTLRFFLPSPSR*LVRNLEA*STS*GSKLSLEDV*GLFT 21
>TC80605 similar to PIR|S56716|S56716 protein kinase SPK-3 (EC 2.7.1.-) -
soybean, partial (25%)
Length = 533
Score = 29.6 bits (65), Expect = 5.7
Identities = 14/29 (48%), Positives = 18/29 (61%)
Frame = +1
Query: 701 RKLYFICWRLCRLN*NNICHLYYGVYGST 729
R LYFICW +N* + H+ YG Y S+
Sbjct: 337 RHLYFICW----VN*KSPIHVNYGYYDSS 411
>BQ141780 similar to SP|P16059|PSBP Oxygen-evolving enhancer protein 2
chloroplast precursor (OEE2), partial (43%)
Length = 745
Score = 28.9 bits (63), Expect = 9.7
Identities = 14/34 (41%), Positives = 20/34 (58%)
Frame = +2
Query: 878 TCPNLDALFPHVVLCSVLSFLTLGWSLLGDKPTW 911
TCPN+ P + S FL+ G LLG+KP++
Sbjct: 476 TCPNILTRSPSPTMASPEEFLSNGDYLLGNKPSF 577
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.365 0.165 0.642
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 35,849,043
Number of Sequences: 36976
Number of extensions: 614034
Number of successful extensions: 8264
Number of sequences better than 10.0: 12
Number of HSP's better than 10.0 without gapping: 1811
Number of HSP's successfully gapped in prelim test: 303
Number of HSP's that attempted gapping in prelim test: 6346
Number of HSP's gapped (non-prelim): 2413
length of query: 943
length of database: 9,014,727
effective HSP length: 105
effective length of query: 838
effective length of database: 5,132,247
effective search space: 4300822986
effective search space used: 4300822986
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 14 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 36 (21.6 bits)
S2: 63 (28.9 bits)
Medicago: description of AC135231.7


