

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
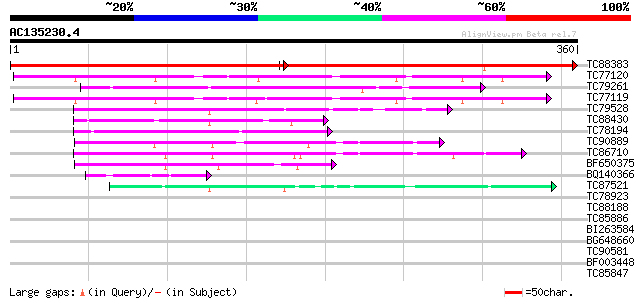
Query= AC135230.4 - phase: 0
(360 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC88383 similar to GP|21592515|gb|AAM64465.1 nuclear receptor bi... 365 0.0
TC77120 similar to PIR|D96533|D96533 ARP protein [imported] - Ar... 81 7e-16
TC79261 similar to PIR|T49047|T49047 quinone reductase-like prot... 79 4e-15
TC77119 similar to PIR|D96533|D96533 ARP protein [imported] - Ar... 72 3e-13
TC79528 similar to GP|15808674|gb|AAL06644.1 putative quinone ox... 69 3e-12
TC88430 weakly similar to PIR|T10824|T10824 auxin-induced protei... 64 1e-10
TC78194 similar to PIR|T05166|T05166 quinone reductase homolog F... 54 9e-08
TC90889 similar to GP|15724256|gb|AAL06521.1 AT4g13010/F25G13_10... 51 6e-07
TC86710 similar to GP|15724256|gb|AAL06521.1 AT4g13010/F25G13_10... 49 4e-06
BF650375 similar to GP|15724256|gb AT4g13010/F25G13_100 {Arabido... 48 5e-06
BQ140366 weakly similar to SP|P41681|ADH2_ Alcohol dehydrogenase... 42 3e-04
TC87521 similar to SP|Q9ZRF1|MTD_FRAAN Probable mannitol dehydro... 42 4e-04
TC78923 similar to PIR|E96680|E96680 hypothetical protein F5I14.... 38 0.005
TC88188 homologue to SP|P80572|ADHX_PEA Alcohol dehydrogenase cl... 37 0.012
TC85886 similar to SP|Q9ZRF1|MTD_FRAAN Probable mannitol dehydro... 37 0.015
BI263584 similar to SP|Q9P6C8|ADH1 Alcohol dehydrogenase I (EC 1... 35 0.045
BG648660 similar to SP|Q9ZRF1|MTD_ Probable mannitol dehydrogena... 33 0.22
TC90581 similar to PIR|E86357|E86357 alcohol dehydrogenase homol... 32 0.50
BF003448 similar to GP|10177732|dbj sorbitol dehydrogenase-like ... 32 0.50
TC85847 homologue to SP|P13603|ADH1_TRIRP Alcohol dehydrogenase ... 29 2.5
>TC88383 similar to GP|21592515|gb|AAM64465.1 nuclear receptor binding
factor-like protein {Arabidopsis thaliana}, partial
(90%)
Length = 1313
Score = 365 bits (938), Expect(2) = 0.0
Identities = 177/177 (100%), Positives = 177/177 (100%)
Frame = +3
Query: 1 MLRSLSHKPSPSSLFNFTSLRSRILNTHTHAFSSAVSPPSKAIIYESHGQPDAVTKLVDI 60
MLRSLSHKPSPSSLFNFTSLRSRILNTHTHAFSSAVSPPSKAIIYESHGQPDAVTKLVDI
Sbjct: 6 MLRSLSHKPSPSSLFNFTSLRSRILNTHTHAFSSAVSPPSKAIIYESHGQPDAVTKLVDI 185
Query: 61 PATEVKENDLCVKMLAAPINPSDINRIQGVYPVRPEPPAVGGYEGVGEVHSVGSAVTCFS 120
PATEVKENDLCVKMLAAPINPSDINRIQGVYPVRPEPPAVGGYEGVGEVHSVGSAVTCFS
Sbjct: 186 PATEVKENDLCVKMLAAPINPSDINRIQGVYPVRPEPPAVGGYEGVGEVHSVGSAVTCFS 365
Query: 121 PGDWVIPSPPSFGTWQTYIVKDQNVWHKVNKGVPMEYAATITVNPLTALLMLEDCVT 177
PGDWVIPSPPSFGTWQTYIVKDQNVWHKVNKGVPMEYAATITVNPLTALLMLEDCVT
Sbjct: 366 PGDWVIPSPPSFGTWQTYIVKDQNVWHKVNKGVPMEYAATITVNPLTALLMLEDCVT 536
Score = 355 bits (912), Expect(2) = 0.0
Identities = 184/199 (92%), Positives = 186/199 (93%), Gaps = 10/199 (5%)
Frame = +1
Query: 172 LEDCVTLNSGDAIVQNGATSMVGQCVIQLAKSRGIHNINIIRDRPGVGEVKERLKDLGAD 231
L+ + LNSGDAIVQNGATSMVGQCVIQLAKSRGIHNINIIRDRPGVGEVKERLKDLGAD
Sbjct: 520 LKTALLLNSGDAIVQNGATSMVGQCVIQLAKSRGIHNINIIRDRPGVGEVKERLKDLGAD 699
Query: 232 EVFTESELEVKNVKSLLGGIPEPALGFNCVGGNSASLVLKFLRRGGTMVTYGGMSKKPVT 291
EVFTESELEVKNVKSLLGGIPEPALGFNCVGGNSASLVLKFLRRGGTMVTYGGMSKKPVT
Sbjct: 700 EVFTESELEVKNVKSLLGGIPEPALGFNCVGGNSASLVLKFLRRGGTMVTYGGMSKKPVT 879
Query: 292 VSTSSFIFK----------NWLSTDKAEEGRRMIDRLLGLVQDGKLKYKMELTPFNDFNT 341
VSTSSFIFK NWLSTDKAEEGRRMIDRLLGLVQDGKLKYKMELTPFNDFNT
Sbjct: 880 VSTSSFIFKELSLRGFWLQNWLSTDKAEEGRRMIDRLLGLVQDGKLKYKMELTPFNDFNT 1059
Query: 342 ALDKALGKLGSQPKQVIKF 360
ALDKALGKLGSQPKQVIKF
Sbjct: 1060ALDKALGKLGSQPKQVIKF 1116
>TC77120 similar to PIR|D96533|D96533 ARP protein [imported] - Arabidopsis
thaliana, partial (93%)
Length = 2203
Score = 80.9 bits (198), Expect = 7e-16
Identities = 97/369 (26%), Positives = 150/369 (40%), Gaps = 27/369 (7%)
Frame = +3
Query: 3 RSLSHKPSPSSLFNFTSLRSRILNTHTHAFSSAVSPPS--KAIIYESHGQPDAVTKLVDI 60
R L + P+PS + + +R + S P S K +++ T +V
Sbjct: 864 RGLEYWPTPSEEAKYLTRPTRFRRKSEYQAPSIKLPDSFEKTVVHTLTHNFRNATSIVRA 1043
Query: 61 PAT-EVKENDLCVKMLAAPINPSDINRIQGVY------PVRPEPPAVGGYEGVGEVHSVG 113
P +K N + VK++ A +N SD+N G Y P G+E VG + +VG
Sbjct: 1044 PLRLPIKPNHVLVKIIYAGVNASDVNFSSGRYFGGNNKETAARLPFDAGFEAVGIIAAVG 1223
Query: 114 SAVTCFSPGDWVIPSPPSFGTWQTYIVKDQNVWHKVNKGVPME--YAATITVNPLTALLM 171
+VT D + P +F T+ Y + + K VP A + + LTA +
Sbjct: 1224 DSVT-----DLKVGMPCAFMTFGGY-AEFTMIPSKYALPVPRPDPEAVAMLTSGLTASIA 1385
Query: 172 LEDCVTLNSGDAIVQNGATSMVGQCVIQLAKSRGIHNINIIRDRPGVGEVKERLKDLGAD 231
LE + SG ++ A GQ +QLAK G N + G G + LK+LG D
Sbjct: 1386 LEKAGQMESGKVVLVTAAAGGTGQFAVQLAKLAG----NTVVATCGGGAKAKLLKELGVD 1553
Query: 232 EVFTESELEVKNV--KSLLGGIPEPALGFNCVGGNSASLVLKFLRRGGTMVTYGGMS--- 286
V ++K V K GI + + VGG+ L L L G ++ G +S
Sbjct: 1554 RVIDYHSEDIKTVLKKEFPKGID---IIYESVGGDMLKLCLDALAVHGRLIVIGMISQYQ 1724
Query: 287 -KKPVTVSTSSFIFKNWLSTDKAEEG----------RRMIDRLLGLVQDGKLKYKMELTP 335
+K T S + + LS +A G + +DRL L GKLK ++
Sbjct: 1725 GEKGWTPSKYPGLCEKLLSKSQAVAGFFLVQYSHMWQEHLDRLFDLYSQGKLKVAVDPKK 1904
Query: 336 FNDFNTALD 344
F ++ D
Sbjct: 1905 FIGLHSVAD 1931
>TC79261 similar to PIR|T49047|T49047 quinone reductase-like protein -
Arabidopsis thaliana, partial (95%)
Length = 1239
Score = 78.6 bits (192), Expect = 4e-15
Identities = 75/261 (28%), Positives = 119/261 (44%), Gaps = 4/261 (1%)
Frame = +3
Query: 46 ESHGQPDAVTKLVDIPATEVKENDLCVKMLAAPINPSDINRIQGVYPVRPEPPAVGGYEG 105
E P ++K IP + + V++ A +N ++ +I G Y +P P + G +
Sbjct: 75 EDENSPIIISKNHPIPQLD-SPTSVRVRIKATSLNFANYLQILGKYQEKPHLPFIPGSDF 251
Query: 106 VGEVHSVGSAVTCFSPGDWVIPSPPSFGTWQTYIVKDQNVWHKVNKGVPMEYAATITVNP 165
G V SVGS VT F GD V S + G++ Y+V DQ +V +G + A + V
Sbjct: 252 SGIVDSVGSNVTNFRVGDPVC-SFAALGSYAQYLVVDQTQLFRVPEGCDLVAAGALAVAF 428
Query: 166 LTALLMLEDCVTLNSGDAIVQNGATSMVGQCVIQLAKSRGIHNINIIRDRPGVGEVK--- 222
T+ + L L SG ++ GA VG +Q+ K+ G I + R V +K
Sbjct: 429 GTSHVGLVHRAQLKSGQVLLVLGAAGGVGLAAVQIGKACGAIVIAVARGAEKVQLLKSMG 608
Query: 223 -ERLKDLGADEVFTESELEVKNVKSLLGGIPEPALGFNCVGGNSASLVLKFLRRGGTMVT 281
+ + DLG + V TES E VK L G + ++ VGG L+ L+ G ++
Sbjct: 609 VDHVVDLGNENV-TESVKEFLKVKRLKG----IDVLYDPVGGKLMKESLRLLKWGANILI 773
Query: 282 YGGMSKKPVTVSTSSFIFKNW 302
G S + + + + KNW
Sbjct: 774 IGFASGEIPVIPANIALVKNW 836
>TC77119 similar to PIR|D96533|D96533 ARP protein [imported] - Arabidopsis
thaliana, partial (93%)
Length = 2277
Score = 72.4 bits (176), Expect = 3e-13
Identities = 97/370 (26%), Positives = 149/370 (40%), Gaps = 28/370 (7%)
Frame = +1
Query: 3 RSLSHKPSPSSLFNFTSLRSRILNTHTHAFSSAVSPPS--KAIIYESHGQPDAVTKLVDI 60
R L + P+PS + R+ + S P S K ++ T +V
Sbjct: 838 RGLEYWPTPSEEAKYLVRPRRLRRRAEYKAPSIKLPESFEKIVVQTLTHNFRNATSIVRA 1017
Query: 61 PAT-EVKENDLCVKMLAAPINPSDINRIQGVY------PVRPEPPAVGGYEGVGEVHSVG 113
P VK N + VK++ A +N SD+N G Y P G+E VG + +VG
Sbjct: 1018 PLRLPVKPNLVLVKIIYAGVNASDVNFSSGRYFGGNNKETTARLPFDAGFEAVGIIAAVG 1197
Query: 114 SAVTCFSPGDWVIPSPPSFGTWQTYIVKDQNVWHKVNKGVPM---EYAATITVNPLTALL 170
+VT D + P +F T+ Y + + K VP E A +T + LTA +
Sbjct: 1198 DSVT-----DLKVGMPCAFMTFGGY-AEFTMIPSKYALPVPRPDPEGVAMLT-SGLTASI 1356
Query: 171 MLEDCVTLNSGDAIVQNGATSMVGQCVIQLAKSRGIHNINIIRDRPGVGEVKERLKDLGA 230
LE + SG ++ A GQ +QLAK G N + G G + LK+LG
Sbjct: 1357 ALEKAGQMESGKVVLVTAAAGGTGQFAVQLAKLAG----NTVVATCGGGTKAKLLKELGV 1524
Query: 231 DEVFTESELEVKNV--KSLLGGIPEPALGFNCVGGNSASLVLKFLRRGGTMVTYGGMS-- 286
D V + ++K V K GI + + VGG+ L L L G ++ G +S
Sbjct: 1525 DRVIDYNSEDIKTVLRKEFPKGID---IIYESVGGDMLKLCLDALAVHGRLIVIGMISQY 1695
Query: 287 --KKPVTVSTSSFIFKNWLSTDKAEEG----------RRMIDRLLGLVQDGKLKYKMELT 334
+ T S + + L+ + G + +DRL L GKLK ++
Sbjct: 1696 QGEHGWTPSKYPGLLEKLLAKSQTVAGFFLVQYSHFYQEHLDRLFDLYSKGKLKVAVDPK 1875
Query: 335 PFNDFNTALD 344
F ++ D
Sbjct: 1876 KFIGLHSVAD 1905
>TC79528 similar to GP|15808674|gb|AAL06644.1 putative quinone
oxidoreductase {Fragaria x ananassa}, partial (72%)
Length = 834
Score = 68.9 bits (167), Expect = 3e-12
Identities = 71/252 (28%), Positives = 111/252 (43%), Gaps = 11/252 (4%)
Frame = +2
Query: 41 KAIIYESHGQPDAVTKL-VDIPATEVKENDLCVKMLAAPINPSDINRIQGVYPVRPEP-P 98
KA Y +G V K ++P +VK++ + +K+ AA +NP D R++G + P P
Sbjct: 92 KAWTYSEYGHSVDVLKFDPNVPIPQVKDDQVLIKVAAASLNPIDYKRMEGAFKASDSPLP 271
Query: 99 AVGGYEGVGEVHSVGSAVTCFSPGDWV--------IPSPPSFGTWQTYIVKDQNVWHKVN 150
GY+ G V VGS V F GD V + P G+ Y ++ +
Sbjct: 272 TAPGYDVSGVVVKVGSEVKKFKVGDEVYGDINVKALEYPKVIGSLAEYTAAEEILLAHKP 451
Query: 151 KGVPMEYAATITVNPLTALLMLEDCVTLNSGDAIVQNGATSMVGQCVIQLAKSRGIHNIN 210
K + AA++ + TA LE ++G +I+ G VG VIQLAK ++ +
Sbjct: 452 KNLSFAEAASLPLTIETAYEGLER-TGFSAGKSILVLGGAGGVGTHVIQLAKH--VYGAS 622
Query: 211 IIRDRPGVGEVKERLKDLGADEVFTESELEVKNVKSLLGGIPEP-ALGFNCVGGNSASLV 269
+ ++ E L +LGAD L + K +PE + F+ VG
Sbjct: 623 KVAATSSTKKL-ELLSNLGAD-------LPIDYTKENFEDLPEKFDVVFDTVG--ETEKA 772
Query: 270 LKFLRRGGTMVT 281
K L+ GG +VT
Sbjct: 773 FKALKEGGKVVT 808
>TC88430 weakly similar to PIR|T10824|T10824 auxin-induced protein (clone
MII-3) - mung bean, partial (41%)
Length = 1450
Score = 63.5 bits (153), Expect = 1e-10
Identities = 57/174 (32%), Positives = 80/174 (45%), Gaps = 12/174 (6%)
Frame = +2
Query: 41 KAIIYESHGQPDAVTKLVDIPATEVKENDLCVKMLAAPINPSDI-NRIQGVYPVRPEPPA 99
KA YE +G P V KL D P EN L V++ AA +NP D R++ ++P + PA
Sbjct: 35 KAWFYEEYG-PKEVLKLGDFPIPSPLENQLLVQVYAAALNPIDSKRRMRPIFP--SDFPA 205
Query: 100 VGGYEGVGEVHSVGSAVTCFSPGDWV---------IPSPPSFGTWQTYIVKDQNVWHKVN 150
V G + G V G V F GD V + P GT +IV ++N+ +
Sbjct: 206 VPGCDMAGVVIGKGVNVKKFDIGDEVYGNIQDFNSMEKPKQLGTLAQFIVVEENLVARKP 385
Query: 151 KGVPMEYAATITVNPLTALLMLEDCVT--LNSGDAIVQNGATSMVGQCVIQLAK 202
K + E AA++ PL +E T G+ + G VG V+QLAK
Sbjct: 386 KRLSFEEAASL---PLAVQTAIEGFKTGDFKKGETMFVVGGAGGVGTLVLQLAK 538
>TC78194 similar to PIR|T05166|T05166 quinone reductase homolog F18E5.200 -
Arabidopsis thaliana, partial (60%)
Length = 709
Score = 53.9 bits (128), Expect = 9e-08
Identities = 42/165 (25%), Positives = 79/165 (47%)
Frame = +1
Query: 41 KAIIYESHGQPDAVTKLVDIPATEVKENDLCVKMLAAPINPSDINRIQGVYPVRPEPPAV 100
KAI+ + G P+ V +L ++ + K+N++ +++ A +N +D + +G YP+
Sbjct: 124 KAIVITTPGGPE-VLQLQEVQDPQPKDNEVLIRVHATALNSADTVQRKGFYPIPQGASPY 300
Query: 101 GGYEGVGEVHSVGSAVTCFSPGDWVIPSPPSFGTWQTYIVKDQNVWHKVNKGVPMEYAAT 160
G E G + VG V+ + GD V G + V + V V G+ ++ AA+
Sbjct: 301 PGLECSGTIVCVGINVSKWKIGDQVCALLAGGGYAEKVAVPEGQVL-PVPPGISLKDAAS 477
Query: 161 ITVNPLTALLMLEDCVTLNSGDAIVQNGATSMVGQCVIQLAKSRG 205
T T + L+ G+ ++ +G +S +G I +AK +G
Sbjct: 478 FTEVACTVWSTIFMTSRLSKGETLLVHGGSSGIGTFAIXIAKYQG 612
>TC90889 similar to GP|15724256|gb|AAL06521.1 AT4g13010/F25G13_100
{Arabidopsis thaliana}, partial (69%)
Length = 779
Score = 51.2 bits (121), Expect = 6e-07
Identities = 66/248 (26%), Positives = 103/248 (40%), Gaps = 13/248 (5%)
Frame = +3
Query: 42 AIIYESHGQPDAVTKLVDIPATEVKENDLCVKMLAAPINPSDINRIQGV--YPVRPEPPA 99
A+ Y+S+ + K V++ K N++ +K+ A+ INP D G+ Y + + P
Sbjct: 66 AVQYDSYDGGSSGLKHVEVHIPTPKANEVLIKVEASSINPVDWKIQDGLLRYLLPRKFPF 245
Query: 100 VGGYEGVGEVHSVGSAVTCFSPGDWVIP--SPPSFGTWQTYIVKDQNVWHKVNKGVPMEY 157
+ GEV GS V F GD VI + G + V ++ + P E
Sbjct: 246 TPCTDVAGEVVDFGSQVKDFKVGDKVIAKLNNQDGGGLAEFAVASES----LTTLRPSEV 413
Query: 158 AATITVNPLTALLMLEDCVTLNSGDAIVQNG---------ATSMVGQCVIQLAKSRGIHN 208
+A A L D +T +G + + G A+ VG +QLAK H
Sbjct: 414 SAAEGAGLPVAGLAAHDAITKMAGIKLDRTGEPKNILVTAASGGVGVYAVQLAKLGNNH- 590
Query: 209 INIIRDRPGVGEVKERLKDLGADEVFTESELEVKNVKSLLGGIPEPALGFNCVGGNSASL 268
+ G + E +K LGADEV E ++KS G + + +C GN L
Sbjct: 591 ---VTATCGARNI-ELIKSLGADEVLDYKTPEGASLKSPSGRKYDAVI--HCTAGNHGQL 752
Query: 269 VLKFLRRG 276
+ +F R G
Sbjct: 753 LNQFDRNG 776
>TC86710 similar to GP|15724256|gb|AAL06521.1 AT4g13010/F25G13_100
{Arabidopsis thaliana}, partial (98%)
Length = 1176
Score = 48.5 bits (114), Expect = 4e-06
Identities = 79/300 (26%), Positives = 122/300 (40%), Gaps = 12/300 (4%)
Frame = +3
Query: 41 KAIIYESHGQPDAVTKLVDIPATEVKENDLCVKMLAAPINPSDINRIQGVYPVRPEP--- 97
+A+ Y S+G + K ++P K N++ +K+ A INP D G P
Sbjct: 60 QALQYSSYGGGVSGLKHAEVPVPIPKTNEVLLKLEATSINPIDWKIQSGALRAIFLPRKF 239
Query: 98 PAVGGYEGVGEVHSVGSAVTCFSPGDWVIP--SPPSFGTWQTYIVKDQNVWHKVNKGVPM 155
P + + GEV VG V F GD VI + G + V +++ V
Sbjct: 240 PHIPCTDVAGEVVEVGPQVKDFKSGDKVIAKLTHQYGGGLAEFAVASESLTATRPPEVSA 419
Query: 156 EYAATITVNPLTALLMLEDCVTLN---SGDA--IVQNGATSMVGQCVIQLAKSRGIHNIN 210
AA + + LTA L + + +G+ ++ A+ VG +QLAK H
Sbjct: 420 AEAAGLPIAGLTARDALTEIGGIKLDGTGEQKNVLVTAASGGVGSFAVQLAKLGNNH--- 590
Query: 211 IIRDRPGVGEVKERLKDLGADEVFTESELEVKNVKSLLGGIPEPALGFNCVGGNSASLVL 270
+ G + + +K LGADEV E ++KS G + + +C G S
Sbjct: 591 -VTATCGARNI-DFVKSLGADEVLDYKTPEGTSLKSPSGKKYDAVI--HCTTGIPWSTFD 758
Query: 271 KFLRRGGTMV--TYGGMSKKPVTVSTSSFIFKNWLSTDKAEEGRRMIDRLLGLVQDGKLK 328
L G +V T G S + ++F K L R ++ L LV+DGKLK
Sbjct: 759 PNLSEKGVVVDLTPGPSSMLTFALKKTTFSKKR-LVPFVVTVKREGLEHLAQLVKDGKLK 935
>BF650375 similar to GP|15724256|gb AT4g13010/F25G13_100 {Arabidopsis
thaliana}, partial (55%)
Length = 564
Score = 48.1 bits (113), Expect = 5e-06
Identities = 52/180 (28%), Positives = 75/180 (40%), Gaps = 14/180 (7%)
Frame = +1
Query: 42 AIIYESHGQPDAVTKLVDIPATEVKENDLCVKMLAAPINPSDINRIQGVYPVRPEP---P 98
A+ Y S+G + K V++P K N++ +K+ A INP D QGV P P
Sbjct: 16 AVQYNSYGGGASGLKHVEVPIPTPKTNEVLIKLEAISINPLDWKIQQGVLRAMFFPRKFP 195
Query: 99 AVGGYEGVGEVHSVGSAVTCFSPGDWVIPSPPS--FGTWQTYIVKDQNVWHKVNKGVPME 156
+ + GEV VG V F GD VI + G + V +++ V
Sbjct: 196 HIPCTDVAGEVVEVGPRVKDFKVGDKVIAKLTNEYGGGLAEFAVASESLTAARPSEVSAA 375
Query: 157 YAATITVNPLTALLMLEDCVTLNSG---------DAIVQNGATSMVGQCVIQLAKSRGIH 207
AA + + +TA D +T N G I+ A+ VG V+QLAK H
Sbjct: 376 EAAGLPIAGITA----HDALTKNGGIKLDGTGEQKNILVTAASGGVGAYVVQLAKLGNNH 543
>BQ140366 weakly similar to SP|P41681|ADH2_ Alcohol dehydrogenase 2 (EC
1.1.1.1). [Deer mouse] {Peromyscus maniculatus}, partial
(11%)
Length = 327
Score = 42.4 bits (98), Expect = 3e-04
Identities = 28/81 (34%), Positives = 47/81 (57%), Gaps = 1/81 (1%)
Frame = +3
Query: 49 GQPDAVTKL-VDIPATEVKENDLCVKMLAAPINPSDINRIQGVYPVRPEPPAVGGYEGVG 107
G+P V ++ VD P K ++ VKML + + +DI+ ++G +P P +G +EG+G
Sbjct: 90 GKPVTVEEIQVDPP----KAKEVRVKMLCSSVCHTDISSLKG-FPHNQFPLXLG-HEGIG 251
Query: 108 EVHSVGSAVTCFSPGDWVIPS 128
V S+G VT GD++ P+
Sbjct: 252 VVESIGDQVTNLKEGDFIXPT 314
>TC87521 similar to SP|Q9ZRF1|MTD_FRAAN Probable mannitol dehydrogenase (EC
1.1.1.255) (NAD-dependent mannitol dehydrogenase).
[Strawberry], partial (98%)
Length = 1433
Score = 42.0 bits (97), Expect = 4e-04
Identities = 72/324 (22%), Positives = 126/324 (38%), Gaps = 40/324 (12%)
Frame = +2
Query: 64 EVKENDLCVKMLAAPINPSDINRIQGVYPVRPEPPAVGGYEGVGEVHSVGSAVTCFSPGD 123
E E D+ K+L I SD++ I+ + + P V G+E G V VGS V F GD
Sbjct: 167 ETGEKDVAFKVLYCGICHSDLHMIKNEWGMSTYP-LVPGHEIAGIVTEVGSKVEKFKIGD 343
Query: 124 WV----------------------------------IPSPPSFGTWQTYIVKDQNVWHKV 149
V ++G + +V D++ +
Sbjct: 344 KVGVGCLVDSCRACQNCEENLENYCPKQTNTYSAKYSDGSITYGGYSDSMVADEHFIVHI 523
Query: 150 NKGVPMEYAATITVNPLTALLMLE----DCVTLNSGDAIVQNGATSMVGQCVIQLAKSRG 205
G+P+E AA + +T L D +N G IV G +G ++ AK+ G
Sbjct: 524 PDGLPLESAAPLLCAGITVYSPLRYFGLDKPGMNIG--IVGLGGLGHLG---VKFAKAFG 688
Query: 206 IHNINIIRDRPGVGEVKERLKDLGADE-VFTESELEVKNVKSLLGGIPEPALGFNCVGGN 264
N+ +I P + KE +++LGAD + + + +++ L GI + V +
Sbjct: 689 A-NVTVISTSP--NKEKEAIENLGADSFLISHDQDKMQAAMGTLDGI------IDTVSAD 841
Query: 265 SASL-VLKFLRRGGTMVTYGGMSKKPVTVSTSSFIFKNWLSTDKAEEGRRMIDRLLGLVQ 323
L ++ L+ G +V G K P + + +S G + ++
Sbjct: 842 HPLLPLVGLLKYHGKLVMVGAPDKPPELPHIPLIMGRKTIS-GSGIGGMKETQEMIDFAA 1018
Query: 324 DGKLKYKMELTPFNDFNTALDKAL 347
+K +E+ P + NTA+ + L
Sbjct: 1019KHNIKPDIEVIPVDYVNTAMKRLL 1090
>TC78923 similar to PIR|E96680|E96680 hypothetical protein F5I14.9
[imported] - Arabidopsis thaliana, partial (66%)
Length = 1367
Score = 38.1 bits (87), Expect = 0.005
Identities = 47/192 (24%), Positives = 81/192 (41%), Gaps = 5/192 (2%)
Frame = +2
Query: 104 EGVGEVHSVGSAVTCFSPGDWVIPSPPSFGTWQTYIV---KDQNVWHKVNKGVPMEYAAT 160
EG G + S F G ++ F W+ Y + DQ + + +P+ +
Sbjct: 311 EGFGVSKVIQSDNPNFKAGHYI----SGFTGWEEYSIITKTDQLRKIEPDDHIPLSFHLG 478
Query: 161 ITVNP-LTALLMLEDCVTLNSGDAIVQNGATSMVGQCVIQLAKSRGIHNINIIRDRPGVG 219
+ P TA + + SG+ + + A+ VGQ V QLAK G + + + V
Sbjct: 479 LLGMPGFTAYAGFYEVCSPKSGEYVFVSAASGAVGQLVGQLAKLHGCYVVGSAGSKEKVE 658
Query: 220 EVKERLKDLGADEVFTESELEVKNVKSLLGGIPEPA-LGFNCVGGNSASLVLKFLRRGGT 278
+K + LG DE F E E+ +L PE + F+ VGG+ L ++ G
Sbjct: 659 LLKSK---LGFDEAFNYKE-ELDLDAALKRYFPEGIDIYFDNVGGDMLDAALLNMKIHGR 826
Query: 279 MVTYGGMSKKPV 290
+ G +S++ +
Sbjct: 827 IAVCGMISQQSI 862
>TC88188 homologue to SP|P80572|ADHX_PEA Alcohol dehydrogenase class III (EC
1.1.1.1) (Glutathione-dependent formaldehyde
dehydrogenase), complete
Length = 1597
Score = 37.0 bits (84), Expect = 0.012
Identities = 33/121 (27%), Positives = 52/121 (42%)
Frame = +1
Query: 7 HKPSPSSLFNFTSLRSRILNTHTHAFSSAVSPPSKAIIYESHGQPDAVTKLVDIPATEVK 66
H PS S F+F +L +T + A+ +E P+ + D+ +
Sbjct: 175 HFPSVVSDFSFLNLI*MASSTEGQVITCKA-----AVAWE----PNKPLTIEDVEVAPPQ 327
Query: 67 ENDLCVKMLAAPINPSDINRIQGVYPVRPEPPAVGGYEGVGEVHSVGSAVTCFSPGDWVI 126
N++ +++L + +D G P P + G+E G V SVG VT PGD VI
Sbjct: 328 ANEVRIQILFTALCHTDAYTWSGKDP-EGLFPCILGHEAAGIVESVGEGVTDVKPGDHVI 504
Query: 127 P 127
P
Sbjct: 505 P 507
>TC85886 similar to SP|Q9ZRF1|MTD_FRAAN Probable mannitol dehydrogenase (EC
1.1.1.255) (NAD-dependent mannitol dehydrogenase).
[Strawberry], partial (94%)
Length = 1395
Score = 36.6 bits (83), Expect = 0.015
Identities = 66/321 (20%), Positives = 125/321 (38%), Gaps = 39/321 (12%)
Frame = +2
Query: 64 EVKENDLCVKMLAAPINPSDINRIQGVYPVRPEPPAVGGYEGVGEVHSVGSAVTCFSPGD 123
E E D+ K+L I SD++ +Q + + P V G+E G V VGS V F GD
Sbjct: 170 ETGEKDVAFKVLYCGICHSDLHIMQNEWG-KTTYPLVPGHELTGVVTEVGSKVKKFKVGD 346
Query: 124 -----WVIPSPPS-----------------------------FGTWQTYIVKDQNVWHKV 149
+++ S S +G + +V D++ ++
Sbjct: 347 KVGVGYMVDSCRSCENCADDIENYCTKYTQTFNGKSRDGTITYGGFSDSMVADEHFVIRI 526
Query: 150 NKGVPMEYAATITVNPLTALLMLE----DCVTLNSGDAIVQNGATSMVGQCVIQLAKSRG 205
+P++ A + +T L D +N G +V G +G ++ AK+ G
Sbjct: 527 PDSLPLDGAGPLLCAGVTVYSPLRHFGLDKPGMNIG--VVGLGG---LGHMAVKFAKAFG 691
Query: 206 IHNINIIRDRPGVGEVKERLKDLGADEVFTESELE-VKNVKSLLGGIPEPALGFNCVGGN 264
+ +I P + KE ++ LGAD + E ++ S L GI + +
Sbjct: 692 A-KVTVISTSP--KKEKEAIEHLGADSFLVSRDPEQMQAATSTLNGIID-----TVSASH 847
Query: 265 SASLVLKFLRRGGTMVTYGGMSKKPVTVSTSSFIFKNWLSTDKAEEGRRMIDRLLGLVQD 324
++ L+ G +V G ++ KP+ + S + G + ++
Sbjct: 848 PVVPLIGLLKSNGKLVMVGAVA-KPLELPIFSLLGGRKSIAGSLIGGIKETQEMIDFAAK 1024
Query: 325 GKLKYKMELTPFNDFNTALDK 345
+ ++E+ P + NTA+++
Sbjct: 1025HNVTPEIEVVPIDYVNTAMER 1087
>BI263584 similar to SP|Q9P6C8|ADH1 Alcohol dehydrogenase I (EC 1.1.1.1).
{Neurospora crassa}, partial (57%)
Length = 682
Score = 35.0 bits (79), Expect = 0.045
Identities = 39/177 (22%), Positives = 72/177 (40%), Gaps = 27/177 (15%)
Frame = +2
Query: 60 IPATEVKENDLCVKMLAAPINPSDINRIQGVYPVRPEPPAVGGYEGVGEVHSVGSAVTCF 119
IP + +++ V + + + +D++ QG +P+ + P VGG+EG G V + G V
Sbjct: 140 IPVQKPGPDEVLVNVKFSGVCHTDLHAWQGDWPLDTKLPLVGGHEGAGVVVARGDLVKDV 319
Query: 120 SPGD-----WVIPSPPSF----------------------GTWQTYIVKDQNVWHKVNKG 152
G+ W+ S S G++Q Y + ++ +
Sbjct: 320 KIGEKVGIKWLNGSCLSCSYCQNADESLCAEALLSGYTVDGSFQQYAIAKAIHVARIPEE 499
Query: 153 VPMEYAATITVNPLTALLMLEDCVTLNSGDAIVQNGATSMVGQCVIQLAKSRGIHNI 209
+E + I +T L++ + +G +I GA +G Q K+ GIH I
Sbjct: 500 CDLESISPILCAGITVYKGLKES-GVKAGQSIAIVGAGGGLGSIAXQYCKAMGIHAI 667
>BG648660 similar to SP|Q9ZRF1|MTD_ Probable mannitol dehydrogenase (EC
1.1.1.255) (NAD-dependent mannitol dehydrogenase).
[Strawberry], partial (64%)
Length = 721
Score = 32.7 bits (73), Expect = 0.22
Identities = 22/62 (35%), Positives = 33/62 (52%)
Frame = +1
Query: 64 EVKENDLCVKMLAAPINPSDINRIQGVYPVRPEPPAVGGYEGVGEVHSVGSAVTCFSPGD 123
E+ E D+ +++L I SD+++ + + P V G+E VG V VGS V F GD
Sbjct: 115 EICEKDVALRVLYCGICHSDLHKAKNEWGTT-NYPLVPGHEIVGIVTEVGSKVKKFKVGD 291
Query: 124 WV 125
V
Sbjct: 292 RV 297
>TC90581 similar to PIR|E86357|E86357 alcohol dehydrogenase homolog
[imported] - Arabidopsis thaliana, partial (52%)
Length = 1289
Score = 31.6 bits (70), Expect = 0.50
Identities = 27/90 (30%), Positives = 44/90 (48%), Gaps = 3/90 (3%)
Frame = +1
Query: 41 KAIIYESHGQPDAVTKL-VDIPATEVKENDLCVKMLAAPINPSDIN--RIQGVYPVRPEP 97
KA I + G+P + ++ +D P K ++ +K+L + SD+ ++ P
Sbjct: 295 KAAICKKSGEPLVIEEVELDPP----KSWEVRIKILCTSLCHSDVTFWKMNSSAPTA-RF 459
Query: 98 PAVGGYEGVGEVHSVGSAVTCFSPGDWVIP 127
P + G+E VG V SVG V GD V+P
Sbjct: 460 PRILGHEAVGLVESVGENVEEVKEGDLVVP 549
>BF003448 similar to GP|10177732|dbj sorbitol dehydrogenase-like protein
{Arabidopsis thaliana}, partial (21%)
Length = 750
Score = 31.6 bits (70), Expect = 0.50
Identities = 23/77 (29%), Positives = 36/77 (45%), Gaps = 2/77 (2%)
Frame = -3
Query: 55 TKLVDIPATEVKENDLCVKMLAAPINPSDINRIQGVYPVR--PEPPAVGGYEGVGEVHSV 112
TK+ + +D+ +KM A I SD++ ++ + + P V G+E G + V
Sbjct: 253 TKIQPFNLPSLGPHDVRIKMKAVGICGSDVHYLKTLRCADFIVKEPMVIGHECAGIIEEV 74
Query: 113 GSAVTCFSPGDWVIPSP 129
GS V PGD V P
Sbjct: 73 GSQVKTLVPGDRVAIEP 23
>TC85847 homologue to SP|P13603|ADH1_TRIRP Alcohol dehydrogenase 1 (EC
1.1.1.1). [Creeping white clover] {Trifolium repens},
partial (60%)
Length = 751
Score = 29.3 bits (64), Expect = 2.5
Identities = 20/78 (25%), Positives = 37/78 (46%), Gaps = 2/78 (2%)
Frame = +1
Query: 48 HGQPDAVTKLVDIPATEVKENDLCVKMLAAPINPSDIN--RIQGVYPVRPEPPAVGGYEG 105
HG+ + ++ + ++ +K+L + +D+ +G P+ P + G+E
Sbjct: 94 HGRXGKPLVMEEVEVAPPQAGEVRLKILFTSLCHTDVYFWEAKGQTPLFPR---IFGHEA 264
Query: 106 VGEVHSVGSAVTCFSPGD 123
G V SVG VT +PGD
Sbjct: 265 GGIVESVGEGVTHLTPGD 318
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.316 0.135 0.397
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 10,956,876
Number of Sequences: 36976
Number of extensions: 159861
Number of successful extensions: 790
Number of sequences better than 10.0: 65
Number of HSP's better than 10.0 without gapping: 778
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 783
length of query: 360
length of database: 9,014,727
effective HSP length: 97
effective length of query: 263
effective length of database: 5,428,055
effective search space: 1427578465
effective search space used: 1427578465
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 59 (27.3 bits)
Medicago: description of AC135230.4


