

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC135230.3 + phase: 0 /pseudo
(605 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
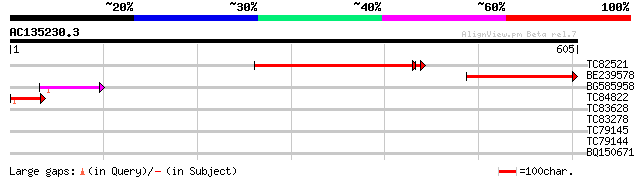
Score E
Sequences producing significant alignments: (bits) Value
TC82521 similar to PIR|T47514|T47514 hypothetical protein T6D9.7... 348 2e-96
BE239578 similar to PIR|T47514|T475 hypothetical protein T6D9.70... 252 3e-67
BG585958 weakly similar to GP|22830897|dbj ORF-B {Oryza sativa (... 64 1e-10
TC84822 similar to GP|21554249|gb|AAM63324.1 unknown {Arabidopsi... 48 1e-05
TC83628 similar to GP|16604352|gb|AAL24182.1 AT5g12010/F14F18_18... 38 0.013
TC83278 similar to GP|9294065|dbj|BAB02022.1 contains similarity... 30 3.5
TC79145 weakly similar to GP|17528994|gb|AAL38707.1 unknown prot... 28 7.8
TC79144 weakly similar to GP|17528994|gb|AAL38707.1 unknown prot... 28 7.8
BQ150671 28 7.8
>TC82521 similar to PIR|T47514|T47514 hypothetical protein T6D9.70 -
Arabidopsis thaliana, partial (41%)
Length = 655
Score = 348 bits (894), Expect(2) = 2e-96
Identities = 173/174 (99%), Positives = 174/174 (99%)
Frame = +3
Query: 262 RPSFGIAFDIDGVILLGNTPVGGSPAALRKLYNYDGTLKFPYVFLTNGGGIPEAKRASEL 321
RPSFGIAFDIDGVILLGNTPVGGSPAALRKLYNYDGTLKFPYVFLTNGGGIPEAKRASEL
Sbjct: 108 RPSFGIAFDIDGVILLGNTPVGGSPAALRKLYNYDGTLKFPYVFLTNGGGIPEAKRASEL 287
Query: 322 SELLGLNVSASQVLQGHSPFRQLVNRFEDKLIVAAGKGEPALVMSEYGFKNVISIDAYAS 381
SELLGLNVSASQVLQGHSPFRQLVNRFEDKLIVAAGKGEPALVMSEYGFKNVISIDAYAS
Sbjct: 288 SELLGLNVSASQVLQGHSPFRQLVNRFEDKLIVAAGKGEPALVMSEYGFKNVISIDAYAS 467
Query: 382 RFENIDPLAPYKKWTTKLATTQNPKFDESGPQIDVFSERVQAAFIVSDPVDWSR 435
RFENIDPLAPYKKWTTKLATTQNPKFDESGPQIDVFSERVQAAFIVSDPVDWS+
Sbjct: 468 RFENIDPLAPYKKWTTKLATTQNPKFDESGPQIDVFSERVQAAFIVSDPVDWSK 629
Score = 23.1 bits (48), Expect(2) = 2e-96
Identities = 9/10 (90%), Positives = 10/10 (100%)
Frame = +1
Query: 434 SRDVQVLCDI 443
+RDVQVLCDI
Sbjct: 625 ARDVQVLCDI 654
>BE239578 similar to PIR|T47514|T475 hypothetical protein T6D9.70 -
Arabidopsis thaliana, partial (26%)
Length = 600
Score = 252 bits (643), Expect = 3e-67
Identities = 118/118 (100%), Positives = 118/118 (100%)
Frame = +3
Query: 488 ESIFNRTHHHSLEYTCFGKPHPSVFKNAEIVLQKHVPRVDEDSYDINHKNAQPFQTLYMI 547
ESIFNRTHHHSLEYTCFGKPHPSVFKNAEIVLQKHVPRVDEDSYDINHKNAQPFQTLYMI
Sbjct: 3 ESIFNRTHHHSLEYTCFGKPHPSVFKNAEIVLQKHVPRVDEDSYDINHKNAQPFQTLYMI 182
Query: 548 GDNPAVDIRGARQTGHPWFSILTRTGVFKGKENHDKFPADLVVDTVEEAVDYILAKEC 605
GDNPAVDIRGARQTGHPWFSILTRTGVFKGKENHDKFPADLVVDTVEEAVDYILAKEC
Sbjct: 183 GDNPAVDIRGARQTGHPWFSILTRTGVFKGKENHDKFPADLVVDTVEEAVDYILAKEC 356
>BG585958 weakly similar to GP|22830897|dbj ORF-B {Oryza sativa (japonica
cultivar-group)}, partial (13%)
Length = 769
Score = 64.3 bits (155), Expect = 1e-10
Identities = 36/73 (49%), Positives = 42/73 (57%), Gaps = 3/73 (4%)
Frame = +2
Query: 32 EEGSASRTR---KYFNRDHVAANQRLIDDYFANEPTYDDAMFRRRYRMQKHVFLRIVGDL 88
E S+SR R + R+ +RL +DYF+ P Y D FRRRYRM KHVFLRIV L
Sbjct: 446 EFASSSRPRSQRRNIERNREEGYKRLFNDYFSEAPVYMDE*FRRRYRMHKHVFLRIVEAL 625
Query: 89 SSSDNYFTQQVDA 101
D YF VDA
Sbjct: 626 GQHDEYFQLTVDA 664
>TC84822 similar to GP|21554249|gb|AAM63324.1 unknown {Arabidopsis
thaliana}, partial (47%)
Length = 636
Score = 47.8 bits (112), Expect = 1e-05
Identities = 23/42 (54%), Positives = 31/42 (73%), Gaps = 4/42 (9%)
Frame = +1
Query: 1 MDP----FDIEAYFQKRDVEDTYIVNRFIQRRKKLEEGSASR 38
MDP FDIEAY +K ++E+ YI+NRF +RRK++EEG R
Sbjct: 40 MDPSKFRFDIEAYKRKSEIEERYILNRFRERRKQIEEGYTPR 165
>TC83628 similar to GP|16604352|gb|AAL24182.1 AT5g12010/F14F18_180
{Arabidopsis thaliana}, partial (23%)
Length = 773
Score = 37.7 bits (86), Expect = 0.013
Identities = 28/106 (26%), Positives = 44/106 (41%)
Frame = +2
Query: 61 NEPTYDDAMFRRRYRMQKHVFLRIVGDLSSSDNYFTQQVDAANKKGISPLAKCTTAMRML 120
N + D FRR +RM K F I +L SS + + + I + + L
Sbjct: 431 NREDFPDDEFRRCFRMSKQTFNMICNELDSS----VTKKNTTLRDAIPVRQRVAVCIYRL 598
Query: 121 AYGVAADAVDEYIKIGSSTTLECLLRFCKGIIRLYEQVYLRAPTQD 166
A G V + +G ST + +L C I + Q +LR P ++
Sbjct: 599 ATGEPLRLVSKKFGLGISTCHKLVLEVCAAIKSVLMQKFLRWPDEE 736
>TC83278 similar to GP|9294065|dbj|BAB02022.1 contains similarity to myb
proteins~gene_id:MRC8.8 {Arabidopsis thaliana}, partial
(11%)
Length = 920
Score = 29.6 bits (65), Expect = 3.5
Identities = 18/51 (35%), Positives = 24/51 (46%), Gaps = 2/51 (3%)
Frame = -2
Query: 181 PWMIGSIDCMHW--EWKNYPKAWEGKLAFANHVRARSRVASRITSRSSEAH 229
P++ G ID H W ++ G+L ANH R ASR + SS H
Sbjct: 244 PFLTGQIDFFHLFEHWVPVRRSK*GRLQTANHTDQRHATASRS*ASSSPVH 92
>TC79145 weakly similar to GP|17528994|gb|AAL38707.1 unknown protein
{Arabidopsis thaliana}, partial (13%)
Length = 733
Score = 28.5 bits (62), Expect = 7.8
Identities = 11/30 (36%), Positives = 16/30 (52%)
Frame = +1
Query: 483 FRIALESIFNRTHHHSLEYTCFGKPHPSVF 512
F + L + +N HHH +T P+PS F
Sbjct: 130 FGVVLANSYNHVHHHYFFFTPLLSPYPSQF 219
>TC79144 weakly similar to GP|17528994|gb|AAL38707.1 unknown protein
{Arabidopsis thaliana}, partial (41%)
Length = 941
Score = 28.5 bits (62), Expect = 7.8
Identities = 11/30 (36%), Positives = 16/30 (52%)
Frame = +1
Query: 483 FRIALESIFNRTHHHSLEYTCFGKPHPSVF 512
F + L + +N HHH +T P+PS F
Sbjct: 136 FGVVLANSYNHVHHHYFFFTPLLSPYPSQF 225
>BQ150671
Length = 948
Score = 28.5 bits (62), Expect = 7.8
Identities = 18/47 (38%), Positives = 24/47 (50%), Gaps = 3/47 (6%)
Frame = +1
Query: 100 DAANKKGISPLAKCT-TAMRMLA--YGVAADAVDEYIKIGSSTTLEC 143
D A ++ + PLA C T + LA + V A + DEY S T EC
Sbjct: 526 DVAGRR*MEPLAPCVGTRIDALASPFSVTAPSSDEYASRPQSATAEC 666
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.322 0.139 0.418
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 18,122,088
Number of Sequences: 36976
Number of extensions: 249532
Number of successful extensions: 1118
Number of sequences better than 10.0: 18
Number of HSP's better than 10.0 without gapping: 1114
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1118
length of query: 605
length of database: 9,014,727
effective HSP length: 102
effective length of query: 503
effective length of database: 5,243,175
effective search space: 2637317025
effective search space used: 2637317025
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 61 (28.1 bits)
Medicago: description of AC135230.3


