

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC135162.7 + phase: 0 /pseudo
(2175 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
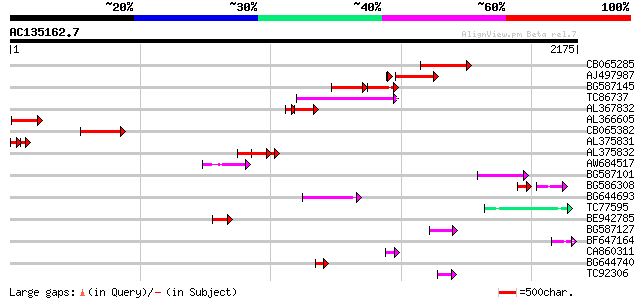
Score E
Sequences producing significant alignments: (bits) Value
CB065285 weakly similar to PIR|A84500|A84 probable retroelement ... 399 e-111
AJ497987 weakly similar to GP|9927273|dbj Similar to Arabidopsis... 300 5e-81
BG587145 similar to PIR|H86337|H8 protein F5M15.26 [imported] - ... 152 9e-59
TC86737 weakly similar to GP|6683624|dbj|BAA89272.1 Pol {Alterna... 179 1e-44
AL367832 weakly similar to GP|14091845|gb Putative retroelement ... 168 2e-41
AL366605 147 6e-35
CB065382 similar to PIR|T09671|T096 RPE15 protein - alfalfa (fra... 144 3e-34
AL375831 90 1e-30
AL375832 128 3e-29
AW684517 similar to GP|9927273|dbj Similar to Arabidopsis thalia... 117 4e-26
BG587101 similar to GP|6691191|gb F7F22.15 {Arabidopsis thaliana... 111 4e-24
BG586308 weakly similar to PIR|F84528|F8 probable retroelement p... 60 6e-18
BG644693 weakly similar to GP|18767374|g Putative 22 kDa kafirin... 85 4e-16
TC77595 weakly similar to PIR|T18350|T18350 probable pol polypro... 77 6e-14
BE942785 72 3e-12
BG587127 weakly similar to PIR|H84506|H84 probable retroelement ... 65 2e-10
BF647164 weakly similar to GP|6691191|gb F7F22.15 {Arabidopsis t... 50 1e-05
CA860311 weakly similar to GP|7289872|gb|A CG17427 gene product ... 49 2e-05
BG644740 similar to PIR|A84460|A84 probable retroelement pol pol... 49 3e-05
TC92306 48 4e-05
>CB065285 weakly similar to PIR|A84500|A84 probable retroelement gag/pol
polyprotein [imported] - Arabidopsis thaliana, partial
(2%)
Length = 592
Score = 399 bits (1024), Expect = e-111
Identities = 187/197 (94%), Positives = 194/197 (97%)
Frame = -2
Query: 1575 KSKDCEEPLINEGPDPDSKWGLVFDGAVNAYGKGIGAVIVSPQGHHIPFTTRILFECTNN 1634
KSKDCEEPLI EGPDP+SKWGLVFDGAVNAYGKGIGAVIVSPQGH+IPFT RILFECTNN
Sbjct: 591 KSKDCEEPLIGEGPDPNSKWGLVFDGAVNAYGKGIGAVIVSPQGHYIPFTARILFECTNN 412
Query: 1635 MAEYEACIFGIEEAIDMRIKHLDIYGDSALIINQIKGEWETHHANLIPYRDYARRLLTYF 1694
MAEYEACIFGIEEAIDMRIKHLDIYGDSAL+INQIKGEWETHHANLIPYRDYARRLLTYF
Sbjct: 411 MAEYEACIFGIEEAIDMRIKHLDIYGDSALVINQIKGEWETHHANLIPYRDYARRLLTYF 232
Query: 1695 TKVELHHIPRDENQMADALATLSSMFRVNHWNDVPLIKVQRLERPSHVFAIGDVIDQAGE 1754
TKVELHHIPRDENQMADALATLSSMFRVNHWNDVP+IKVQRLERPSHVFAIG+VIDQ GE
Sbjct: 231 TKVELHHIPRDENQMADALATLSSMFRVNHWNDVPIIKVQRLERPSHVFAIGNVIDQTGE 52
Query: 1755 NVVDYRPWYYDIKQFLL 1771
NVVDY+PWYYDIKQFL+
Sbjct: 51 NVVDYKPWYYDIKQFLV 1
>AJ497987 weakly similar to GP|9927273|dbj Similar to Arabidopsis thaliana
chromosome II BAC F26H6; putative retroelement pol
polyprotein, partial (1%)
Length = 636
Score = 300 bits (767), Expect = 5e-81
Identities = 132/167 (79%), Positives = 154/167 (92%)
Frame = -2
Query: 1479 KTCCALAWATKRLRHYLVNHTTWLISRMDPIKYIFEKAAVTGKIARWQMLLSEYDIVFKT 1538
KTCCALAWA KRLRHY++NHTTWL+S+MDPIKYIFEK A+TG+IARWQMLLSEYDI +++
Sbjct: 635 KTCCALAWAAKRLRHYMINHTTWLVSKMDPIKYIFEKPALTGRIARWQMLLSEYDIEYRS 456
Query: 1539 QKAIKGSILADHLAYQPLDDYQPIEFDFPDEEIMYLKSKDCEEPLINEGPDPDSKWGLVF 1598
QKAIKGSILADHLA+QPL+DY+PI+FDFPDEEIMYLK KDC+EPL EGPDPDS WGL+F
Sbjct: 455 QKAIKGSILADHLAHQPLEDYRPIKFDFPDEEIMYLKMKDCDEPLFGEGPDPDSVWGLIF 276
Query: 1599 DGAVNAYGKGIGAVIVSPQGHHIPFTTRILFECTNNMAEYEACIFGI 1645
DGAVN YG GIGAV+++P+G HIPFT R+ F+CTNN+AEYEACI GI
Sbjct: 275 DGAVNVYGNGIGAVLLTPKGAHIPFTARLRFDCTNNIAEYEACIMGI 135
Score = 48.1 bits (113), Expect = 4e-05
Identities = 20/24 (83%), Positives = 23/24 (95%)
Frame = +2
Query: 1445 CVLGQQDETGKKEHAIYYLSKKFT 1468
CVLGQQDETG+KEHAIYYLS+K +
Sbjct: 38 CVLGQQDETGRKEHAIYYLSRKLS 109
>BG587145 similar to PIR|H86337|H8 protein F5M15.26 [imported] - Arabidopsis
thaliana, partial (13%)
Length = 763
Score = 152 bits (383), Expect(2) = 9e-59
Identities = 70/136 (51%), Positives = 101/136 (73%)
Frame = +2
Query: 1236 MSPEDREKTSFITPWGTFCYKVMPFGLINAGATYQRGMTTLFHDMIHKEVEVYVDDMIVK 1295
M P+D EKT+FIT GT+CYKVMPFGL NAG+TYQR + +F D + +EVY+DDM+VK
Sbjct: 2 MHPDDLEKTAFITDRGTYCYKVMPFGLKNAGSTYQRLVNRMFADKLGNTMEVYIDDMLVK 181
Query: 1296 SADEEQHVEYLTKMFERLRKYKLRLNPNKCTFGVRSGKLLGFIVSQKGIEVDPDKVRAIR 1355
S H+ +L + F+ L +Y ++LNP KCTFGV SG+ LG+IV+Q+GIEV+P ++ AI
Sbjct: 182 SLRATDHLNHLKE*FKTLDEYIMKLNPAKCTFGVTSGEFLGYIVTQQGIEVNPKQITAIL 361
Query: 1356 EMPAPQTEKQVRGFLG 1371
++P+P+ ++V+ G
Sbjct: 362 DLPSPKNSREVQRLTG 409
Score = 95.9 bits (237), Expect(2) = 9e-59
Identities = 48/120 (40%), Positives = 74/120 (61%)
Frame = +3
Query: 1371 GRLNYISRFISHMTATCGPIFKLLRKNQPIVWNDECQEAFDSIKNYLLEPPILVPPVEGR 1430
GR+ ++RFIS T C P +KLL N+ VW+++C+EAF+ +K YL PP+L P G
Sbjct: 408 GRIAALNRFISRSTDKCLPFYKLLCGNKRFVWDEKCEEAFEQLKQYLTTPPVLSKPEAGD 587
Query: 1431 PLIMYLAVFDESMGCVLGQQDETGKKEHAIYYLSKKFTDCETRYTMLEKTCCALAWATKR 1490
L +Y+A+ ++ VL ++D +K I+Y SK+ TD ETRY LEK A+ + ++
Sbjct: 588 TLSLYIAISSTAVSSVLIREDRGEQK--PIFYTSKRMTDPETRYPTLEKMAFAVITSARK 761
>TC86737 weakly similar to GP|6683624|dbj|BAA89272.1 Pol {Alternaria
alternata}, partial (21%)
Length = 1540
Score = 179 bits (454), Expect = 1e-44
Identities = 122/397 (30%), Positives = 198/397 (49%), Gaps = 7/397 (1%)
Frame = +1
Query: 1101 LREYPDIFAWSYEDMPGLDPMIVEHRIPTKPEC----PPVRQ-KLRRTHPDMALKIKNEV 1155
L E+PD+F +++H IP P+ PP+ L L +K +
Sbjct: 346 LEEFPDLFNPEKAYQVPASRGLLDHAIPLIPDKDGNDPPLPWGPLYGMSRQELLVLKKTL 525
Query: 1156 QKQIDAGFLMTVEYPEWVANIVPVPKKDGKVRMCVDFRDLNKASPKDNFPLPHIDVLVDN 1215
+ +D GF+ A ++ V K G +R CVD+R LN + KD +PLP I +
Sbjct: 526 EDLLDKGFIKASGSAAG-APVLFVRKPGGGIRFCVDYRALNAITKKDRYPLPLISETLRR 702
Query: 1216 TAQSKVFSFMDGFSGYNQIKMSPEDREKTSFITPWGTFCYKVMPFGLINAGATYQRGMTT 1275
A ++ F+ +D + ++++++ ED+EKT+F T +G F + V PFGL A AT+QR +
Sbjct: 703 VAGARWFTKLDVVAAFHKMRIKDEDQEKTAFRTRYGLFEWIVCPFGLTGAPATFQRYINK 882
Query: 1276 LFHDMIHKEVEVYVDDMIV-KSADEEQHVEYLTKMFERLRKYKLRLNPNKCTFGVRSGKL 1334
H+ + V Y+DD+++ + ++ H + ++ RL L L+P KC F V + K
Sbjct: 883 TLHEFLDDFVTAYIDDVLIYTTGSKKDHEAQVRRVLRRLADAGLSLDPKKCEFSVTTVKY 1062
Query: 1335 LGFIVSQ-KGIEVDPDKVRAIREMPAPQTEKQVRGFLGRLNYISRFISHMTATCGPIFKL 1393
+GFI++ KG+ DP K+ AIR+ P + K R FLG NY FI + P+ +L
Sbjct: 1063 VGFILTAGKGVSCDPLKLAAIRDWLPPGSVKGARSFLGFCNYYKDFIPGYSEITEPLTRL 1242
Query: 1394 LRKNQPIVWNDECQEAFDSIKNYLLEPPILVPPVEGRPLIMYLAVFDESMGCVLGQQDET 1453
RK+ P W E + AF +K E P+L + ++G VL Q+D T
Sbjct: 1243 TRKDFPFRWGAEQEAAFTKLKRLFAEEPVLRMFDPEAVTTVETDCSGFALGGVLTQEDGT 1422
Query: 1454 GKKEHAIYYLSKKFTDCETRYTMLEKTCCALAWATKR 1490
G H + + S++ + E Y + +K A+ WA R
Sbjct: 1423 G-AAHPVAFHSQRLSPAEYNYPIHDKELLAV-WACLR 1527
>AL367832 weakly similar to GP|14091845|gb Putative retroelement {Oryza
sativa}, partial (2%)
Length = 384
Score = 168 bits (426), Expect = 2e-41
Identities = 81/94 (86%), Positives = 85/94 (90%)
Frame = -1
Query: 1091 KGLNRRSSNSLREYPDIFAWSYEDMPGLDPMIVEHRIPTKPECPPVRQKLRRTHPDMALK 1150
+G+ R+ LREY DIFA SYEDMPGLDP IVEHRIPTKPECPPVR KLRRTHPDMALK
Sbjct: 282 EGVKRKIFQLLREYLDIFACSYEDMPGLDPKIVEHRIPTKPECPPVR*KLRRTHPDMALK 103
Query: 1151 IKNEVQKQIDAGFLMTVEYPEWVANIVPVPKKDG 1184
IK+EVQKQIDAGFLMTVEYPEWVANIVPVPKKDG
Sbjct: 102 IKSEVQKQIDAGFLMTVEYPEWVANIVPVPKKDG 1
Score = 61.2 bits (147), Expect = 4e-09
Identities = 33/44 (75%), Positives = 38/44 (86%)
Frame = -2
Query: 1057 KKRKPFTLTKKRSSLST*VPRKTSEKSRLVPL*RKGLNRRSSNS 1100
KK KPF+LT+KR SLST*VPR+TSE+SR V L*RKGL RSS+S
Sbjct: 383 KKGKPFSLTRKRLSLST*VPRRTSERSRSVLL*RKGLKGRSSSS 252
>AL366605
Length = 422
Score = 147 bits (370), Expect = 6e-35
Identities = 81/119 (68%), Positives = 93/119 (78%)
Frame = -1
Query: 5 CQKWMFLKSSRFRTSTGTTG*LVLKITSSNTSGRWAITKTMIPL*STVFRIV*WRMPQSG 64
CQ+WM L++S+ + STGTTG*LV K TSSNTS RWAITKTMIPL*ST RIV*W+M QSG
Sbjct: 359 CQRWMSLRNSKSQISTGTTG*LVPKTTSSNTSERWAITKTMIPL*STASRIV*WKMLQSG 180
Query: 65 ILV*VRMTSIPLMS*PPLSRAIMGLTLD*GQIGSFSGPYLRRRRKVSVNMRKGGEGRLP 123
ILV* +M SIP M+*P LSRA MGLT D* + G+ S LRRRRK SVN + G G+LP
Sbjct: 179 ILV*AKMISIPSMN*PLLSRATMGLTPD*SRTGNSSDLSLRRRRKASVNTHRDGGGQLP 3
>CB065382 similar to PIR|T09671|T096 RPE15 protein - alfalfa (fragment),
partial (9%)
Length = 624
Score = 144 bits (364), Expect = 3e-34
Identities = 94/172 (54%), Positives = 108/172 (62%)
Frame = -3
Query: 271 RFNNNNKISKPGQPSLLYQCCMQNFFQLYSREVIVQLDRVSPHLIRCLQGSGLISNVTFI 330
RF NN+ ++ P L YQC M + +LY E Q DR S HLIR L GSGL SNV I
Sbjct: 517 RFTNNDSSNRLDLPFL*YQCYMLSSSRLYFSEGTAQSDRGSLHLIRYLPGSGLTSNVISI 338
Query: 331 KVPWVMTSRGVML*SIS*RSSSIKGS*LLRIMSHMSSTIRYQIMPL*I*SKCVKKLRDLM 390
KV WVM S+ ML*+ *RSS K S*LLR S MSST +QI L* * + ++KL DL
Sbjct: 337 KVLWVMMSKDAML*NTL*RSSLTKES*LLRTTSLMSSTTLFQITLL*T*LRYMRKLPDLT 158
Query: 391 SATSQLLWYLYTSSCAKLLCSIMIMPSVWDVSVILWVATLFKTTFKA**MII 442
S QLLW YTSS AK LC MIMP + S+ILWVA LFK T +A** I
Sbjct: 157 SVMLQLLWCHYTSSYAKPLCLTMIMPIAKNDSIILWVAALFKMTSRA**TTI 2
>AL375831
Length = 467
Score = 89.7 bits (221), Expect(2) = 1e-30
Identities = 44/44 (100%), Positives = 44/44 (100%)
Frame = +3
Query: 1 TFA*CQKWMFLKSSRFRTSTGTTG*LVLKITSSNTSGRWAITKT 44
TFA*CQKWMFLKSSRFRTSTGTTG*LVLKITSSNTSGRWAITKT
Sbjct: 234 TFA*CQKWMFLKSSRFRTSTGTTG*LVLKITSSNTSGRWAITKT 365
Score = 64.3 bits (155), Expect(2) = 1e-30
Identities = 33/37 (89%), Positives = 35/37 (94%)
Frame = +1
Query: 41 ITKTMIPL*STVFRIV*WRMPQSGILV*VRMTSIPLM 77
+ + MIPL*STVFRIV*WRMPQSGILV*VRMTSIPLM
Sbjct: 355 LQRQMIPL*STVFRIV*WRMPQSGILV*VRMTSIPLM 465
>AL375832
Length = 535
Score = 128 bits (321), Expect = 3e-29
Identities = 82/130 (63%), Positives = 86/130 (66%), Gaps = 2/130 (1%)
Frame = +1
Query: 875 GLLMLSLKKPLDLVSDLHLLHQEALRQIGMPLTFLQSCMFLSNVSCCFNALFSK*QSSRS 934
GLLMLSLKK LDLVS LHLLHQEALR IG PLTFLQSCMF SN SCCF+ALFSK
Sbjct: 1 GLLMLSLKKQLDLVSGLHLLHQEALRVIGTPLTFLQSCMFPSNASCCFSALFSKITILSL 180
Query: 935 AQSESDIFCKG--MFVSALPLINKILSFLPRLFFFSAFSLGNNGNARKKNQNVCIFLLIS 992
F G F A+ I ++SF FF FS NGNAR KN NVCIFLLIS
Sbjct: 181 CPKRE*YFL*GRVCFGIAV**IKIVVSF-TTAFFLVPFS-WKNGNAR-KNLNVCIFLLIS 351
Query: 993 KICMKKKKKK 1002
K CMKKKK K
Sbjct: 352 KTCMKKKKYK 381
Score = 105 bits (263), Expect = 1e-22
Identities = 68/109 (62%), Positives = 71/109 (64%), Gaps = 2/109 (1%)
Frame = +2
Query: 926 FSK*QSSRSAQSESDIFCKGMFVSALPLINKILSFLPRLFFFSAFSLGNNGNARKKNQNV 985
F K*QSSRSAQSESDIFCKG+FVSALP LSFL RL FF LG K +V
Sbjct: 155 FQK*QSSRSAQSESDIFCKGVFVSALPFDE*KLSFLSRLLFF*CLFLGKMVMPEKT*TSV 334
Query: 986 CIFLLISKICMKK--KKKKTLSKIIQSLYAG*TIINPLNTVILRFLQVL 1032
+ KK K KKTLSKIIQSLYAG*TIINPLNTVIL Q L
Sbjct: 335 FSY*FQKPA*KKKNTKNKKTLSKIIQSLYAG*TIINPLNTVILCLFQTL 481
>AW684517 similar to GP|9927273|dbj Similar to Arabidopsis thaliana
chromosome II BAC F26H6; putative retroelement pol
polyprotein, partial (1%)
Length = 488
Score = 117 bits (294), Expect = 4e-26
Identities = 83/181 (45%), Positives = 98/181 (53%)
Frame = +2
Query: 741 WTLMHPTVACWEDRGYTMQVRLLPPYIRS*NL*RMENSLQSMVKKHT*SANCLLFLVLKQ 800
W M WED GY R PY RS*NL*RM+ +L++
Sbjct: 17 WISMLRIAVSWEDLGYMTLAR*PLPYTRS*NL*RMD--------------------LLRE 136
Query: 801 GLLKGLLSKV*PLKVQSLRKLGLLWLH*RMLREPSKKVRLPAGAR*YSSVRTSVKKVSGS 860
L KG L KVQS R+L L WL *R R ++ RLP GA * +SVR S +KVS S
Sbjct: 137 LLFKGCL-----WKVQSPRRLELQWLL*RTPRRLFRRDRLPTGAS*SNSVRISERKVSDS 301
Query: 861 PRHQGFPRESSTVQGLLMLSLKKPLDLVSDLHLLHQEALRQIGMPLTFLQSCMFLSNVSC 920
P+H+ PRE V GL + LK+ LDL L +QEALR+IGMPL F QSCM NVSC
Sbjct: 302 PQHRVSPRELFIVPGLSIHLLKRWLDLFLGPCL*YQEALRKIGMPLMFPQSCMCPXNVSC 481
Query: 921 C 921
C
Sbjct: 482 C 484
>BG587101 similar to GP|6691191|gb F7F22.15 {Arabidopsis thaliana}, partial
(10%)
Length = 624
Score = 111 bits (277), Expect = 4e-24
Identities = 62/196 (31%), Positives = 96/196 (48%)
Frame = +2
Query: 1793 RFLLDGNILYKRNYDMVLLRCVDEHEAEQLMHDVHDGTFGTHATGHTMSRKLLRAGYYWM 1852
R+L D LYK+ D + +RCV E E ++ H + H K+ +AG++W
Sbjct: 17 RYLWDEPFLYKQCADNIYIRCVAEEEIPGILFHCHGSNYAGHFAVSKTVSKIQQAGFWWP 196
Query: 1853 AMEHDCYQHARKCHKCQIYADKIHVPPHALNVMSSPWPFSMWGIDMIGRIEPKASNGHRF 1912
M D + KC CQ + N + F +WGID +G +N ++
Sbjct: 197 TMFKDAHSFISKCDPCQRQGNIS*RNEMPQNFILEVEVFDVWGIDFMGPFPSSYNN--KY 370
Query: 1913 ILVAIDYFTKWVEAASYTNVTKQVVAKFIKNNIICRYGVPSKIITDNGTNLNNNVVQALC 1972
ILVA+DY +KWVEA + VV K K+ I R+GVP +I+D G++ N V + L
Sbjct: 371 ILVAVDYVSKWVEAIASPTNDATVVVKMFKSVIFPRFGVPRVVISDGGSHFINKVFEKLL 550
Query: 1973 EEFKIEHHNSSPYRPQ 1988
++ + H ++ Y PQ
Sbjct: 551 KKNGVRHKVATAYHPQ 598
>BG586308 weakly similar to PIR|F84528|F8 probable retroelement pol polyprotein
[imported] - Arabidopsis thaliana, partial (7%)
Length = 686
Score = 60.5 bits (145), Expect(2) = 6e-18
Identities = 39/122 (31%), Positives = 68/122 (54%), Gaps = 4/122 (3%)
Frame = -3
Query: 2022 LHGYRTTVRSSTGATPFSLVYGMEAVLPLEVEIPSL---RVIMEAKLSEAEWCQSRYDQL 2078
L +RT R +T +TPFS+ + +EA+ P EV + SL R+ +L+ ++ L
Sbjct: 462 LWSHRTNPRGATKSTPFSMAHRVEAMAPAEVNVTSL*RSRMPQNIELNN----DRLFNAL 295
Query: 2079 NLIEEKRMDAMARGQSYQARMKTAFDKKVHPREFKVGELVLKRRISQQPD-PRGKWTPNY 2137
IEE+R A+ R Q+YQ ++++ ++K V + K+G++VL + + GK N+
Sbjct: 294 ETIEERRDQALLRIQNYQHQIESYYNKTVKSQPLKLGDIVLCKVFENTKELNAGKLGTNW 115
Query: 2138 EG 2139
EG
Sbjct: 114 EG 109
Score = 50.4 bits (119), Expect(2) = 6e-18
Identities = 22/52 (42%), Positives = 35/52 (67%)
Frame = -2
Query: 1949 YGVPSKIITDNGTNLNNNVVQALCEEFKIEHHNSSPYRPQMNGAVEAANKNI 2000
+G+P +I+TDNG++ +N + CE ++I + +SP PQ NG EA+NK I
Sbjct: 685 HGLPYEIVTDNGSHFISNKFREFCERWRIRLNTASPRYPQSNGQAEASNKII 530
>BG644693 weakly similar to GP|18767374|g Putative 22 kDa kafirin cluster;
Ty3-Gypsy type {Oryza sativa}, partial (15%)
Length = 716
Score = 84.7 bits (208), Expect = 4e-16
Identities = 65/230 (28%), Positives = 109/230 (47%), Gaps = 5/230 (2%)
Frame = +2
Query: 1123 VEHRIPTKPECPPVRQKLRRTHPDMALKIKNEVQKQIDAGFLMTVEYPEWVANIVPVPKK 1182
++ I P P+ R +P +K +++ ++ GF+ YP V ++ + KK
Sbjct: 35 IDFGIDLLPNMNPI*IPSYRINPLKLKVLKLQLKDLLEKGFIQPSIYP*GVV-VLFLKKK 211
Query: 1183 DGKVRMCVDFRDLNKASPKDNFPLPHIDVLVDNTAQSKVFSFMDGFSGYNQIKMSPEDRE 1242
DG +RM +D+ LN + K +PLP ID L DN SK F +D G +Q ++ ED
Sbjct: 212 DGFLRMSIDYPQLNNVNIKIKYPLPLIDELFDNLQGSKWFFKIDLRLG*HQHRVIGEDVP 391
Query: 1243 KTSFITPWGTFCYKVMPFGLINAGATYQRGMTTLFHDMIHKEVEVYVDDMIVKSADEEQH 1302
KT+F +G + VM FG N + M +F D + V V+ +D+++ S +E +H
Sbjct: 392 KTAFRIRYGHYEILVMSFG*TNPPMAFMELMNRVFQDYLDSLVIVFSNDILIYSKNENEH 571
Query: 1303 VEYLTKMFERLRKYKLRLNPNKCTFG-----VRSGKLLGFIVSQKGIEVD 1347
+L + L+ L C V G ++S +G++VD
Sbjct: 572 ENHLRLALKVLKDIGL------CQISYV*ILVEVGFFSLHVISGEGLKVD 703
>TC77595 weakly similar to PIR|T18350|T18350 probable pol polyprotein - rice
blast fungus gypsy retroelement (fragment), partial (14%)
Length = 1708
Score = 77.4 bits (189), Expect = 6e-14
Identities = 75/353 (21%), Positives = 140/353 (39%), Gaps = 15/353 (4%)
Frame = +2
Query: 1821 QLMHDVHDGTFGTHATGHTMSRKLLRAGYYWMAMEHDCYQHARKCHKCQIYADKIHVPPH 1880
+L+ + HD T H G + +++ ++W + R C C IH+
Sbjct: 176 KLVQESHDSTAAGHP-GRNGTLEIVSRKFFWPGQSQTVRRFVRNCDVC----GGIHIWRQ 340
Query: 1881 ALNVMSSPWPF-----SMWGIDMIGRIEPKASNGHRFILVAIDYFTKWVEAASYTNVTKQ 1935
A P P S +D I + P G +++ V +D +K V + +
Sbjct: 341 AKRGFLKPLPVPNRLHSDLSMDFITSLPPTRGRGSQYLWVIVDRLSKSVTLEEMDTMEAE 520
Query: 1936 VVAKFIKNNIICRYGVPSKIITDNGTNLNNNVVQALCEEFKIEHHNSSPYRPQMNGAVEA 1995
A+ + +G+P I++D G+N + C + S+ Y PQ +G E
Sbjct: 521 ACAQRFLSCHYRFHGMPQSIVSDRGSNWVGRFWREFCRLTGVTQLLSTSYHPQTDGGTER 700
Query: 1996 ANKNIKRIVQKMVTTYKD-WHEMLPYALHGYRTTVRSSTGATPFSLVYGMEAVLPLEVEI 2054
N+ I+ +++ V +D W ++LP R SS GATPF + +G V P+
Sbjct: 701 WNQEIQAVLRAYVCWSQDNWGDLLPTVQLALRNRHNSSIGATPFFVEHGYH-VDPIPTVE 877
Query: 2055 PSLRVIMEAKLSEAEWCQSRYDQLNLIEEKRMDAMARGQSYQARMKTAFDKKVHPREFKV 2114
+ V+ E + + + D I+ + + A R ++ + + D+ ++V
Sbjct: 878 DTGGVVSEGEAAAQLLVKRMKDVTGFIQAEIVAAQQRSEASANKRRCPADR------YQV 1039
Query: 2115 GELV---LKRRISQQPDPRGKW------TPNYEGPYVVKKAFSGGALILTHMD 2158
G+ V + S +P + W + P+VV+ G H+D
Sbjct: 1040 GDKVWLNVSNYKSPRPSKKLDWLHHKYEVTRFVTPHVVELNVPGTVYPKFHVD 1198
>BE942785
Length = 460
Score = 71.6 bits (174), Expect = 3e-12
Identities = 45/77 (58%), Positives = 54/77 (69%)
Frame = -3
Query: 779 LQSMVKKHT*SANCLLFLVLKQGLLKGLLSKV*PLKVQSLRKLGLLWLH*RMLREPSKKV 838
L M +KHT* A+ L L KQGL +G L K * KV+S R + L WL *RM REPSK+V
Sbjct: 365 LPFMERKHT*LASYHLSLA*KQGLQRGPLFKD*LSKVRSPRGMELQWLL*RMHREPSKRV 186
Query: 839 RLPAGAR*YSSVRTSVK 855
+LPAGA *YS VRTS++
Sbjct: 185 KLPAGAG*YSFVRTSIR 135
>BG587127 weakly similar to PIR|H84506|H84 probable retroelement pol
polyprotein [imported] - Arabidopsis thaliana, partial
(13%)
Length = 415
Score = 65.5 bits (158), Expect = 2e-10
Identities = 37/109 (33%), Positives = 55/109 (49%)
Frame = +3
Query: 1610 GAVIVSPQGHHIPFTTRILFECTNNMAEYEACIFGIEEAIDMRIKHLDIYGDSALIINQI 1669
G + SP + + R+ F +NN YEA I G+ A ++I+++ Y DS L+ +Q
Sbjct: 3 GIRLTSPTNEILKQSFRLEFHASNNETNYEALIAGVRLAHGLKIRNIHAYCDSQLVASQF 182
Query: 1670 KGEWETHHANLIPYRDYARRLLTYFTKVELHHIPRDENQMADALATLSS 1718
GE+E + Y ++L L IPR EN ADALA L+S
Sbjct: 183 SGEYEARDELMDTYLKLVQKLAQKLDYFALTRIPRSENVQADALAALAS 329
>BF647164 weakly similar to GP|6691191|gb F7F22.15 {Arabidopsis thaliana},
partial (6%)
Length = 469
Score = 49.7 bits (117), Expect = 1e-05
Identities = 35/101 (34%), Positives = 52/101 (50%), Gaps = 3/101 (2%)
Frame = +1
Query: 2077 QLNLIEEKRMDAMARGQSYQARMKTAFDKKVHPREFKVGELVL--KRRISQQPDPRGKWT 2134
+LN +EE R+DA + Y+ R K D+++ REF+ GELVL R+ P GK
Sbjct: 103 ELNELEELRLDAYENAKIYKERTKKWHDRRIIRREFREGELVLLFNSRLKLFP---GKLR 273
Query: 2135 PNYEGPYVVKKAFSGGAL-ILTHMDGIELPNPVNADIVKKY 2174
++ GP+ VK GA+ + + G P VN +K Y
Sbjct: 274 SHWSGPFQVKNVMPSGAVEVWSESTG---PFTVNGQRLKHY 387
>CA860311 weakly similar to GP|7289872|gb|A CG17427 gene product {Drosophila
melanogaster}, partial (20%)
Length = 192
Score = 48.9 bits (115), Expect = 2e-05
Identities = 24/56 (42%), Positives = 31/56 (54%)
Frame = +1
Query: 1440 DESMGCVLGQQDETGKKEHAIYYLSKKFTDCETRYTMLEKTCCALAWATKRLRHYL 1495
D +G L Q+DE EH I Y S+ T E YT++E+ C A WA + RHYL
Sbjct: 7 DWGIGAGLSQKDEENH-EHPIAYASRLLTAAERNYTVVERECLAAIWAIRNFRHYL 171
>BG644740 similar to PIR|A84460|A84 probable retroelement pol polyprotein
[imported] - Arabidopsis thaliana, partial (4%)
Length = 754
Score = 48.5 bits (114), Expect = 3e-05
Identities = 22/49 (44%), Positives = 30/49 (60%)
Frame = -1
Query: 1174 ANIVPVPKKDGKVRMCVDFRDLNKASPKDNFPLPHIDVLVDNTAQSKVF 1222
A ++ V KKDG RMC+D+R NK + K+ +PLP ID L D + F
Sbjct: 238 AALLFVRKKDGYFRMCIDYRQFNKVTTKNKYPLPRIDNLFDKIQEDCYF 92
>TC92306
Length = 521
Score = 48.1 bits (113), Expect = 4e-05
Identities = 24/73 (32%), Positives = 37/73 (49%)
Frame = +2
Query: 1642 IFGIEEAIDMRIKHLDIYGDSALIINQIKGEWETHHANLIPYRDYARRLLTYFTKVELHH 1701
I G+ EA + +H+ + GD + Q G W+ ++ NL + A L + F V + H
Sbjct: 2 ILGLNEARNQGYEHVHVRGDFQXVCKQFXGSWKVNNPNLRNLCNXAVELKSNFKSVSVEH 181
Query: 1702 IPRDENQMADALA 1714
+PR N ADA A
Sbjct: 182 VPRGXNXAADAQA 220
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.340 0.148 0.492
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 71,070,307
Number of Sequences: 36976
Number of extensions: 1089453
Number of successful extensions: 8683
Number of sequences better than 10.0: 56
Number of HSP's better than 10.0 without gapping: 4612
Number of HSP's successfully gapped in prelim test: 381
Number of HSP's that attempted gapping in prelim test: 3503
Number of HSP's gapped (non-prelim): 5748
length of query: 2175
length of database: 9,014,727
effective HSP length: 111
effective length of query: 2064
effective length of database: 4,910,391
effective search space: 10135047024
effective search space used: 10135047024
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.9 bits)
S2: 66 (30.0 bits)
Medicago: description of AC135162.7


