

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC135162.11 + phase: 0 /pseudo
(385 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
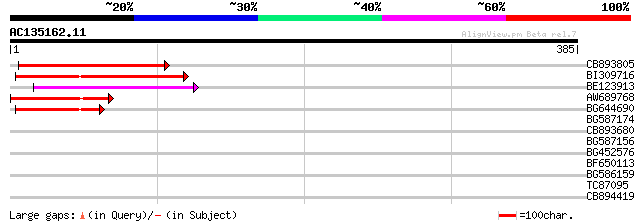
Score E
Sequences producing significant alignments: (bits) Value
CB893805 similar to GP|10177935|d copia-type polyprotein {Arabid... 96 2e-20
BI309716 weakly similar to PIR|G96722|G9 hypothetical protein F2... 87 8e-18
BE123913 weakly similar to GP|22093573|d polyprotein {Oryza sati... 72 3e-13
AW689768 weakly similar to GP|10177485|d polyprotein {Arabidopsi... 52 3e-07
BG644690 weakly similar to GP|18542179|gb putative pol protein {... 49 4e-06
BG587174 similar to PIR|A47759|A4775 retrovirus-related reverse ... 37 0.013
CB893680 weakly similar to GP|1167523|db ORF(AA 1-1338) {Nicotia... 37 0.013
BG587156 similar to PIR|G85055|G8 probable polyprotein [imported... 36 0.029
BG452576 weakly similar to GP|12005223|gb|A reverse transcriptas... 35 0.049
BF650113 weakly similar to GP|4753889|emb| Tpv2-1c {Phaseolus vu... 34 0.11
BG586159 weakly similar to PIR|T47841|T4 hypothetical protein T2... 29 3.5
TC87095 homologue to GP|4165183|emb|CAA10643.1 SAP1 protein {Ant... 28 6.0
CB894419 weakly similar to GP|10177935|db copia-type polyprotein... 28 7.8
>CB893805 similar to GP|10177935|d copia-type polyprotein {Arabidopsis
thaliana}, partial (14%)
Length = 778
Score = 95.9 bits (237), Expect = 2e-20
Identities = 47/102 (46%), Positives = 68/102 (66%)
Frame = +3
Query: 7 RLHKALYGLKQSLRAWNKRIVNFLVQQEFVKCKAEYNVYVKNSKDSNLVHICLYVDDLIV 66
R+ +ALYGLKQ+ RAW RI + ++ F KC E+ ++VK S+ ++ I LYVDDLI
Sbjct: 462 RVKRALYGLKQAPRAWYSRIEAYFTKEGFEKCPYEHTLFVKLSEGGKILIISLYVDDLIF 641
Query: 67 TDSNLAEIEAFKGHMMKKFNMTDLGKLTYFLGMEFTKTSEGL 108
++ E FK M K+FNM+DLGK+ YFLG+E T+ +G+
Sbjct: 642 IGNDENMFEEFKKSMKKEFNMSDLGKMHYFLGVEVTQNEKGI 767
>BI309716 weakly similar to PIR|G96722|G9 hypothetical protein F20P5.25
[imported] - Arabidopsis thaliana, partial (10%)
Length = 744
Score = 87.4 bits (215), Expect = 8e-18
Identities = 36/117 (30%), Positives = 78/117 (65%)
Frame = +2
Query: 5 IFRLHKALYGLKQSLRAWNKRIVNFLVQQEFVKCKAEYNVYVKNSKDSNLVHICLYVDDL 64
+ L K++YGLKQ+ R W ++ L+ +++ ++++++ K KDS+ + +YVDD+
Sbjct: 20 VCELQKSIYGLKQASRQWYSKLSESLISFGYLQSSSDFSLFTK-FKDSSFTTLLVYVDDI 196
Query: 65 IVTDSNLAEIEAFKGHMMKKFNMTDLGKLTYFLGMEFTKTSEGLVKHQKKYASDILK 121
++ ++++EI+ K ++ +F + DLG L YFLG+E ++ +G++ +Q+KY ++L+
Sbjct: 197 VLAGNDISEIQHVKCFLIDRFKIKDLGSLRYFLGLEVARSKQGILLNQRKYTLELLE 367
>BE123913 weakly similar to GP|22093573|d polyprotein {Oryza sativa (japonica
cultivar-group)}, partial (8%)
Length = 503
Score = 72.4 bits (176), Expect = 3e-13
Identities = 35/112 (31%), Positives = 62/112 (55%)
Frame = +1
Query: 17 QSLRAWNKRIVNFLVQQEFVKCKAEYNVYVKNSKDSNLVHICLYVDDLIVTDSNLAEIEA 76
QS R W R + + +++C+ ++ +++K+S + +YVDD+ +T + I+
Sbjct: 1 QSPRDWFDRFT*VVKKFGYIQCQTDHAMFIKHSSTVKKAILIVYVDDIFLTGDHGK*IKR 180
Query: 77 FKGHMMKKFNMTDLGKLTYFLGMEFTKTSEGLVKHQKKYASDILKRFNMLNC 128
K + ++F + DLG L YFLGME + +G Q+KY D+LK M+ C
Sbjct: 181 LKNLLAEEFEIKDLGNLKYFLGMEVARWKKGSSISQRKYVLDLLKETRMIGC 336
>AW689768 weakly similar to GP|10177485|d polyprotein {Arabidopsis thaliana},
partial (9%)
Length = 675
Score = 52.4 bits (124), Expect = 3e-07
Identities = 22/70 (31%), Positives = 48/70 (68%)
Frame = +1
Query: 1 NEDMIFRLHKALYGLKQSLRAWNKRIVNFLVQQEFVKCKAEYNVYVKNSKDSNLVHICLY 60
N+ ++ +L+K+LYGLKQ+ RAW + + + +Q F K + + ++ + N ++ +++ +Y
Sbjct: 463 NKSLVCKLNKSLYGLKQAPRAWYEXLTSAQIQFGFTKSRCDPSLLIYN-QNGACIYLXIY 639
Query: 61 VDDLIVTDSN 70
VDD+++T S+
Sbjct: 640 VDDILITGSS 669
>BG644690 weakly similar to GP|18542179|gb putative pol protein {Zea mays},
partial (22%)
Length = 629
Score = 48.5 bits (114), Expect = 4e-06
Identities = 24/60 (40%), Positives = 41/60 (68%)
Frame = -2
Query: 5 IFRLHKALYGLKQSLRAWNKRIVNFLVQQEFVKCKAEYNVYVKNSKDSNLVHICLYVDDL 64
+FRL+K LYGLKQ+ RAW +R+ FL++ F + K + +++ ++ L+ I +YVDD+
Sbjct: 178 VFRLNKTLYGLKQAPRAWYERLSKFLLKNGFKRGKIDNTLFLL-KRE*ELLIIQVYVDDI 2
>BG587174 similar to PIR|A47759|A4775 retrovirus-related reverse
transcriptase homolog - rape retrotransposon copia-like
(fragment), partial (84%)
Length = 249
Score = 37.0 bits (84), Expect = 0.013
Identities = 21/64 (32%), Positives = 32/64 (49%)
Frame = -1
Query: 2 EDMIFRLHKALYGLKQSLRAWNKRIVNFLVQQEFVKCKAEYNVYVKNSKDSNLVHICLYV 61
ED + +L K+LYGLKQS R W KR ++ + S +++ LYV
Sbjct: 192 EDHVCKLRKSLYGLKQSPRQWYKRFDSYRSSWATTGVLMTVVST*TR*RMSRYIYLVLYV 13
Query: 62 DDLI 65
DD++
Sbjct: 12 DDML 1
>CB893680 weakly similar to GP|1167523|db ORF(AA 1-1338) {Nicotiana tabacum},
partial (7%)
Length = 780
Score = 37.0 bits (84), Expect = 0.013
Identities = 25/72 (34%), Positives = 36/72 (49%), Gaps = 2/72 (2%)
Frame = -3
Query: 64 LIVTDSNLAEIEAFKGHMMKKFNMTDLGKLTYFLGME--FTKTSEGLVKHQKKYASDILK 121
L+V SN+ EI+ K K+ +M DLG +GM+ K L Q +Y + +L+
Sbjct: 271 LLVVGSNIDEIKNLKTRFSKEIDMKDLGPAKKIIGMQIMIDKQKGVL*LSQVEYITRVLQ 92
Query: 122 RFNMLNCNPAST 133
FNM N ST
Sbjct: 91 IFNMGNAILVST 56
>BG587156 similar to PIR|G85055|G8 probable polyprotein [imported] -
Arabidopsis thaliana, partial (17%)
Length = 618
Score = 35.8 bits (81), Expect = 0.029
Identities = 16/41 (39%), Positives = 26/41 (63%)
Frame = -1
Query: 5 IFRLHKALYGLKQSLRAWNKRIVNFLVQQEFVKCKAEYNVY 45
+ RL KA+YGLKQS RAW ++ L + F K + ++ ++
Sbjct: 141 VLRLKKAIYGLKQSPRAWYNKLSTTLNGRGFRKSELDHTLF 19
>BG452576 weakly similar to GP|12005223|gb|A reverse transcriptase-like
protein {Spiranthes hongkongensis}, partial (25%)
Length = 676
Score = 35.0 bits (79), Expect = 0.049
Identities = 17/68 (25%), Positives = 37/68 (54%)
Frame = +2
Query: 59 LYVDDLIVTDSNLAEIEAFKGHMMKKFNMTDLGKLTYFLGMEFTKTSEGLVKHQKKYASD 118
+Y+DD++ + +E + K H+ K F + LG L +++G++ + ++ Q KY +
Sbjct: 344 VYLDDIVFARDDHSETQLVKSHLDKNFIIIGLGTL-HYVGVKIA*SES*IIDDQCKYTIE 520
Query: 119 ILKRFNML 126
+L+ L
Sbjct: 521 LLEESGQL 544
>BF650113 weakly similar to GP|4753889|emb| Tpv2-1c {Phaseolus vulgaris},
partial (13%)
Length = 494
Score = 33.9 bits (76), Expect = 0.11
Identities = 29/74 (39%), Positives = 35/74 (47%)
Frame = +2
Query: 207 DTLREP*TMV*FFLNVSAKLTWS*LPTQMQIGVEICKIGKALHATCSNF*MLLYHGVQKS 266
D L E MV +FL S Q+ IGVEI G+AL CSN ML G +S
Sbjct: 212 DMLEELWNMVCYFLMELRVKCMSSYAIQIVIGVEI---GEALQVMCSNSMMLQSPGAPRS 382
Query: 267 NLY*YFQLVNLSTL 280
N + + LSTL
Sbjct: 383 NQSPHSHPMKLSTL 424
>BG586159 weakly similar to PIR|T47841|T4 hypothetical protein T2O9.150 -
Arabidopsis thaliana, partial (11%)
Length = 732
Score = 28.9 bits (63), Expect = 3.5
Identities = 20/62 (32%), Positives = 30/62 (48%), Gaps = 20/62 (32%)
Frame = +1
Query: 99 MEFTKTSEGLVKHQKKYASDILKRFNM----LNCNPA----------------STLFKQI 138
+E + EG+ Q+KY +D+L+RF M L+ NP +T +KQI
Sbjct: 1 VEVIQNEEGIYICQRKYVTDLLERFGMEKSNLSRNPIAPRCKLIKDENGVKVDATKYKQI 180
Query: 139 VG 140
VG
Sbjct: 181 VG 186
>TC87095 homologue to GP|4165183|emb|CAA10643.1 SAP1 protein {Antirrhinum
majus}, partial (4%)
Length = 987
Score = 28.1 bits (61), Expect = 6.0
Identities = 14/47 (29%), Positives = 23/47 (48%)
Frame = +3
Query: 15 LKQSLRAWNKRIVNFLVQQEFVKCKAEYNVYVKNSKDSNLVHICLYV 61
LK + WN +I QE++ C Y ++ + +HIC+YV
Sbjct: 378 LKL*FQVWNNQI------QEYIHCNKGRQRY*EDYNNWGCIHICVYV 500
>CB894419 weakly similar to GP|10177935|db copia-type polyprotein
{Arabidopsis thaliana}, partial (2%)
Length = 170
Score = 27.7 bits (60), Expect = 7.8
Identities = 13/21 (61%), Positives = 15/21 (70%)
Frame = +3
Query: 113 KKYASDILKRFNMLNCNPAST 133
KK ASDILK+F M + P ST
Sbjct: 3 KKCASDILKKFKMEHSKPIST 65
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.366 0.163 0.562
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 10,786,433
Number of Sequences: 36976
Number of extensions: 136490
Number of successful extensions: 1598
Number of sequences better than 10.0: 26
Number of HSP's better than 10.0 without gapping: 852
Number of HSP's successfully gapped in prelim test: 62
Number of HSP's that attempted gapping in prelim test: 737
Number of HSP's gapped (non-prelim): 927
length of query: 385
length of database: 9,014,727
effective HSP length: 98
effective length of query: 287
effective length of database: 5,391,079
effective search space: 1547239673
effective search space used: 1547239673
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 14 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 36 (21.6 bits)
S2: 59 (27.3 bits)
Medicago: description of AC135162.11


