

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC134967.10 + phase: 0 /pseudo
(624 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
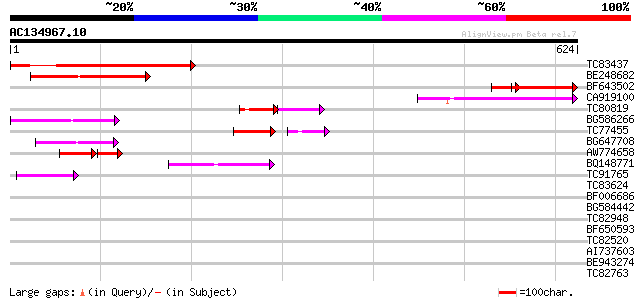
Score E
Sequences producing significant alignments: (bits) Value
TC83437 weakly similar to PIR|D86384|D86384 unknown protein [imp... 272 2e-73
BE248682 similar to GP|18568269|gb putative gag-pol polyprotein ... 125 5e-29
BF643502 120 1e-27
CA919100 homologue to PIR|G90291|G902 endoglucanase precursor [i... 100 2e-21
TC80819 weakly similar to GP|10140689|gb|AAG13524.1 putative non... 67 4e-21
BG586266 similar to GP|7267666|em RNA-directed DNA polymerase-li... 86 4e-17
TC77455 similar to GP|22335695|dbj|BAC10549. nine-cis-epoxycarot... 51 2e-11
BG647708 weakly similar to GP|13786450|gb| putative reverse tran... 65 8e-11
AW774658 similar to GP|2808681|emb| Hcr9-4B {Lycopersicon hirsut... 45 2e-09
BQ148771 55 1e-07
TC91765 weakly similar to GP|19881779|gb|AAM01180.1 Putative ret... 45 8e-05
TC83624 homologue to PIR|G84581|G84581 copia-like retroelement p... 41 0.002
BF006686 41 0.002
BG584442 40 0.002
TC82948 40 0.003
BF650593 39 0.008
TC82520 37 0.017
AI737603 37 0.017
BE943274 33 0.43
TC82763 33 0.43
>TC83437 weakly similar to PIR|D86384|D86384 unknown protein [imported] -
Arabidopsis thaliana, partial (6%)
Length = 951
Score = 272 bits (696), Expect = 2e-73
Identities = 145/204 (71%), Positives = 157/204 (76%)
Frame = +2
Query: 1 WRKWIKECVSTATASVLVNGSPTDEFPLKRGLRQGDPLSPFLFLLAAEGLNIVMKSLVEA 60
WRKWIKECVSTAT SVLVNGSPT N++MKSLV+
Sbjct: 398 WRKWIKECVSTATTSVLVNGSPT---------------------------NVLMKSLVQT 496
Query: 61 QLFSGYSIGAANPIVVSHLQFADDTLLLGSKSWANVRALRAALVLFEAMSGLKVNFHKSM 120
QLF+ YS G NP+VVSHLQFA+DTLLL +K+WAN+RALRAALV+F AMSGLKVNFHKS
Sbjct: 497 QLFTRYSFGVVNPVVVSHLQFANDTLLLETKNWANIRALRAALVIF*AMSGLKVNFHKSG 676
Query: 121 LVGVNIGSSWLYEAASVLSCKVGIVPFLYLDMPIGGNPRRLCFWEPIVNRIKSRLSGWNS 180
LV VNI SWL EAASVLS KVG VPFLYL MPI GN RRL FWEPIVNRIK+RL+GWNS
Sbjct: 677 LVCVNIAPSWLSEAASVLSWKVGKVPFLYLGMPIEGNSRRLSFWEPIVNRIKARLTGWNS 856
Query: 181 RFLSFGGRLVLLKSVLTSLPVYAL 204
RFLSFGGRLVLLKSVLTSL VYAL
Sbjct: 857 RFLSFGGRLVLLKSVLTSLSVYAL 928
>BE248682 similar to GP|18568269|gb putative gag-pol polyprotein {Zea mays},
partial (1%)
Length = 441
Score = 125 bits (314), Expect = 5e-29
Identities = 63/132 (47%), Positives = 89/132 (66%)
Frame = +3
Query: 24 DEFPLKRGLRQGDPLSPFLFLLAAEGLNIVMKSLVEAQLFSGYSIGAANPIVVSHLQFAD 83
+E ++RGL+QGDPL+PFLFLL AEG++ +MK+ V LF G+ + V SHLQ+AD
Sbjct: 6 EEISVQRGLKQGDPLAPFLFLLVAEGISGLMKNAVNRNLFQGFDVKRGGTRV-SHLQYAD 182
Query: 84 DTLLLGSKSWANVRALRAALVLFEAMSGLKVNFHKSMLVGVNIGSSWLYEAASVLSCKVG 143
DTL +G + N+ L+A L FE SGLKVNFHKS L+G+N+ ++ A L+C+
Sbjct: 183 DTLCIGMPTVDNLWTLKALLQGFEMASGLKVNFHKSSLIGINVPRDFMEAACRFLNCREE 362
Query: 144 IVPFLYLDMPIG 155
+PF+YL +P G
Sbjct: 363 SIPFIYLGLPGG 398
>BF643502
Length = 631
Score = 120 bits (302), Expect = 1e-27
Identities = 59/72 (81%), Positives = 64/72 (87%)
Frame = -3
Query: 553 YATNLASLCLGCLE*T**PYV*SCCYPYPLFT**D*VVVSRLVES*KSYLCFWDSAVVVK 612
YAT +AS+CLGCLE*T** +V*SCCYP+PLFT D*VVVSRLVES*K Y+CFWDSA VK
Sbjct: 500 YATYMASMCLGCLE*T**RHV*SCCYPHPLFTRQD*VVVSRLVES*KCYICFWDSAGAVK 321
Query: 613 PSCLFGYWLRFF 624
PSCLFGYWL FF
Sbjct: 320 PSCLFGYWLMFF 285
Score = 42.4 bits (98), Expect = 5e-04
Identities = 17/32 (53%), Positives = 21/32 (65%)
Frame = -1
Query: 531 R*YFSSFFTVHSFDRWWCCTFFYATNLASLCL 562
+* F FTVHSFDRWWCC F+ + LC+
Sbjct: 568 Q*IFQIMFTVHSFDRWWCC*TFFMQLIWLLCV 473
>CA919100 homologue to PIR|G90291|G902 endoglucanase precursor [imported] -
Sulfolobus solfataricus, partial (3%)
Length = 789
Score = 100 bits (249), Expect = 2e-21
Identities = 72/179 (40%), Positives = 96/179 (53%), Gaps = 4/179 (2%)
Frame = -1
Query: 450 GFGVAQTGSLEGFRFCVEAPP*SFTNKDKFD---CSTGVVV*HVFVCCWLRSS*IGSTSI 506
G +AQTGS EG R C+ A *S + + D C++ FVC WLR IG+T +
Sbjct: 591 GDDLAQTGSFEGIRLCLAASS*SVAYQIESDL*RCNSNR---SWFVCFWLRCLGIGTTFV 421
Query: 507 FIV*YFWFSVAFGA*LDRLLWGGHR*YFSSFFTVHSFDRWW-CCTFFYATNLASLCLGCL 565
++ F F++ G+*LD W GH + S +V S +RW + F LASLCLG +
Sbjct: 420 SLLFVFCFTLVVGS*LDWFCWSGH*CFV*SLCSVCSLNRWQ*GFSVFSPAYLASLCLGFM 241
Query: 566 E*T**PYV*SCCYPYPLFT**D*VVVSRLVES*KSYLCFWDSAVVVKPSCLFGYWLRFF 624
+*T* Y + * *+ VSRLVES*K + W +VVKP +FGYWL F
Sbjct: 240 D*T*QYVFQ*FYYSFTQVA*QS*IFVSRLVES*KRFFFVWHF*LVVKPVTMFGYWLTCF 64
>TC80819 weakly similar to GP|10140689|gb|AAG13524.1 putative non-LTR
retroelement reverse transcriptase {Oryza sativa
(japonica cultivar-group)}, partial (2%)
Length = 1262
Score = 66.6 bits (161), Expect(2) = 4e-21
Identities = 31/43 (72%), Positives = 35/43 (81%)
Frame = +1
Query: 254 GLGVRKLKEFNLALLGKWCWRMLVDRGGLWYRVLVARYGELGG 296
GLGV FNL+LLGKWCWR+LVD+ GLW+RVL ARYGE GG
Sbjct: 31 GLGVGA---FNLSLLGKWCWRLLVDKEGLWHRVLKARYGEEGG 150
Score = 53.1 bits (126), Expect(2) = 4e-21
Identities = 23/52 (44%), Positives = 30/52 (57%)
Frame = +3
Query: 295 GGRLEVGGRSVSCWWSEVRRLRDGVGDGGSGWFGDQVLRRVGDGVVTSFWSD 346
GG L GGR S WW + ++R+GVG+G WF + + VGDG FW D
Sbjct: 144 GGWLCEGGRQSSMWWKTICKVREGVGEGVGNWFEENIRMVVGDGRDAFFWYD 299
>BG586266 similar to GP|7267666|em RNA-directed DNA polymerase-like protein
{Arabidopsis thaliana}, partial (18%)
Length = 789
Score = 85.9 bits (211), Expect = 4e-17
Identities = 47/121 (38%), Positives = 67/121 (54%)
Frame = -3
Query: 1 WRKWIKECVSTATASVLVNGSPTDEFPLKRGLRQGDPLSPFLFLLAAEGLNIVMKSLVEA 60
W WI ECVST + S L+NG P RGLRQGDPLSP+LF+L E L+ + + +
Sbjct: 475 WISWIMECVSTVSYSFLINGGPQGRVLPSRGLRQGDPLSPYLFILCTEVLSGLCQQALRK 296
Query: 61 QLFSGYSIGAANPIVVSHLQFADDTLLLGSKSWANVRALRAALVLFEAMSGLKVNFHKSM 120
G + A N ++HL FADDT+ G + ++ L + + + A SG +N KS
Sbjct: 295 GTLPGVKV-ARNCPPINHLLFADDTMFFGKSNASSCAILLSIMDKYRAASGRCIN*TKSA 119
Query: 121 L 121
+
Sbjct: 118 I 116
>TC77455 similar to GP|22335695|dbj|BAC10549. nine-cis-epoxycarotenoid
dioxygenase1 {Pisum sativum}, partial (43%)
Length = 1865
Score = 51.2 bits (121), Expect(2) = 2e-11
Identities = 23/46 (50%), Positives = 30/46 (65%)
Frame = -3
Query: 247 CLKKECGGLGVRKLKEFNLALLGKWCWRMLVDRGGLWYRVLVARYG 292
CL + GGLGVR ++ N++LL KW WR+L D+ LW VL YG
Sbjct: 969 CLPRCKGGLGVRDIRLVNVSLLAKWWWRLLQDQSSLWKEVLEDIYG 832
Score = 35.4 bits (80), Expect(2) = 2e-11
Identities = 17/47 (36%), Positives = 26/47 (55%)
Frame = -2
Query: 306 SCWWSEVRRLRDGVGDGGSGWFGDQVLRRVGDGVVTSFWSDR*GSSL 352
S WW ++ L + G WF +++R+VGDG + FW D SS+
Sbjct: 778 SRWWKDLMSLEEV---GRVRWFPRELIRKVGDGRSSFFWKDAWDSSV 647
>BG647708 weakly similar to GP|13786450|gb| putative reverse transcriptase
{Oryza sativa}, partial (9%)
Length = 708
Score = 65.1 bits (157), Expect = 8e-11
Identities = 35/91 (38%), Positives = 53/91 (57%)
Frame = +1
Query: 29 KRGLRQGDPLSPFLFLLAAEGLNIVMKSLVEAQLFSGYSIGAANPIVVSHLQFADDTLLL 88
++GLRQGDPLSP+LF+L A L+ ++K Q G + ++P ++HL FADD+LL
Sbjct: 7 EKGLRQGDPLSPYLFILCANVLSGLLKREGNKQNLHGIQVARSDP-KITHLLFADDSLLF 183
Query: 89 GSKSWANVRALRAALVLFEAMSGLKVNFHKS 119
+ + L +++ SG VNF KS
Sbjct: 184 ARANLTEAATIMQVLHSYQSASGQLVNFEKS 276
>AW774658 similar to GP|2808681|emb| Hcr9-4B {Lycopersicon hirsutum}, partial
(4%)
Length = 665
Score = 45.4 bits (106), Expect(2) = 2e-09
Identities = 23/40 (57%), Positives = 27/40 (67%)
Frame = -1
Query: 56 SLVEAQLFSGYSIGAANPIVVSHLQFADDTLLLGSKSWAN 95
+L+ + SIG + V SHLQFADDTLLLG KSWAN
Sbjct: 446 NLLSHSICLNLSIGMHSLTVFSHLQFADDTLLLGVKSWAN 327
Score = 35.0 bits (79), Expect(2) = 2e-09
Identities = 16/28 (57%), Positives = 22/28 (78%)
Frame = -3
Query: 97 RALRAALVLFEAMSGLKVNFHKSMLVGV 124
RALR+ LV+FE MSGLKVN + +++ V
Sbjct: 327 RALRSILVIFENMSGLKVNLREEVIIYV 244
>BQ148771
Length = 680
Score = 54.7 bits (130), Expect = 1e-07
Identities = 34/117 (29%), Positives = 54/117 (46%)
Frame = -3
Query: 175 LSGWNSRFLSFGGRLVLLKSVLTSLPVYALSFFKAPTGIISSIDSLFINFFLGGGGGSED 234
L+ W + LS R+ L KSV+ ++P+Y + P I I L F G +E
Sbjct: 573 LANWKANHLSLARRVTLAKSVIEAVPLYPMMTTIIPKACIEEIQKLQRKFV---WGDTEV 403
Query: 235 NRKISWLSWQTVCLKKECGGLGVRKLKEFNLALLGKWCWRMLVDRGGLWYRVLVARY 291
+R+ + W+T+ K GLG+R+L N A + K W + L V+ +Y
Sbjct: 402 SRRYHAVGWETMSKPKTIYGLGLRRLDVMNKACIMKLGWSIYSGSNSLCTEVMRGKY 232
>TC91765 weakly similar to GP|19881779|gb|AAM01180.1 Putative retroelement
{Oryza sativa (japonica cultivar-group)}, partial (1%)
Length = 625
Score = 45.1 bits (105), Expect = 8e-05
Identities = 26/68 (38%), Positives = 35/68 (51%)
Frame = +2
Query: 8 CVSTATASVLVNGSPTDEFPLKRGLRQGDPLSPFLFLLAAEGLNIVMKSLVEAQLFSGYS 67
CV + VLVN D RGL+QGD LSP++F++ EGL+ ++ E G S
Sbjct: 104 CVESNDYYVLVNNDAVDPIIPSRGLQQGDHLSPYIFIICVEGLSFLIPHAKERGDTHGTS 283
Query: 68 IGAANPIV 75
I P V
Sbjct: 284 I*RGAPPV 307
>TC83624 homologue to PIR|G84581|G84581 copia-like retroelement pol
polyprotein [imported] - Arabidopsis thaliana, partial
(1%)
Length = 831
Score = 40.8 bits (94), Expect = 0.002
Identities = 17/57 (29%), Positives = 34/57 (58%)
Frame = +1
Query: 116 FHKSMLVGVNIGSSWLYEAASVLSCKVGIVPFLYLDMPIGGNPRRLCFWEPIVNRIK 172
F + + +N+ S++ + + L C V VPF +L +PIG NP+R +P+++ ++
Sbjct: 256 FSRVNFMALNLEESFVEASPNFLLCNVNEVPFCFLGLPIGANPKRSSTRKPVLDSLQ 426
>BF006686
Length = 325
Score = 40.8 bits (94), Expect = 0.002
Identities = 16/39 (41%), Positives = 23/39 (58%)
Frame = +3
Query: 154 IGGNPRRLCFWEPIVNRIKSRLSGWNSRFLSFGGRLVLL 192
+ G L WEP++ + L W ++ LSFGGR+VLL
Sbjct: 207 VEGRGEILPMWEPLLEHVNKMLKSWGNKLLSFGGRIVLL 323
>BG584442
Length = 775
Score = 40.4 bits (93), Expect = 0.002
Identities = 25/94 (26%), Positives = 45/94 (47%)
Frame = +1
Query: 190 VLLKSVLTSLPVYALSFFKAPTGIISSIDSLFINFFLGGGGGSEDNRKISWLSWQTVCLK 249
V++K L S+ Y +S F + I+ + F G E+ + + W+S + + +
Sbjct: 430 VMIKYALQSISSYVMSIFLLLNSQVDEIEKIMNTFSWVHVG--ENRKGMHWMS*EKLFVH 603
Query: 250 KECGGLGVRKLKEFNLALLGKWCWRMLVDRGGLW 283
K GG+G FN+ +LGK L++R L+
Sbjct: 604 KNYGGMGFTDFTTFNIPMLGKQV*SFLLNRTTLF 705
>TC82948
Length = 705
Score = 40.0 bits (92), Expect = 0.003
Identities = 19/48 (39%), Positives = 28/48 (57%)
Frame = +3
Query: 237 KISWLSWQTVCLKKECGGLGVRKLKEFNLALLGKWCWRMLVDRGGLWY 284
K+ +SW+ VC + G LG+R L + N AL K CW M++ + WY
Sbjct: 255 KVVKVSWEKVCRPIKEGSLGIRSLSKLNEALNLKLCWDMMISK-EQWY 395
>BF650593
Length = 486
Score = 38.5 bits (88), Expect = 0.008
Identities = 14/32 (43%), Positives = 20/32 (61%)
Frame = +1
Query: 281 GLWYRVLVARYGELGGRLEVGGRSVSCWWSEV 312
GLW+R LV +YG G + + R VS WW ++
Sbjct: 85 GLWFRALVNKYGLNRGSITIENRGVSLWWKDI 180
>TC82520
Length = 833
Score = 37.4 bits (85), Expect = 0.017
Identities = 24/68 (35%), Positives = 36/68 (52%), Gaps = 3/68 (4%)
Frame = +1
Query: 502 GSTSIFIV*YFWFSVAFGA*LDRLLWGG---HR*YFSSFFTVHSFDRWWCCTFFYATNLA 558
G+TS+F +*YFW S+ L G H+ F S + F + + +FF+A ++
Sbjct: 334 GNTSVFTL*YFWLSLVSCFALVTSFVGFTC*HKTIFHSIYFYGGFAKIY--SFFFANHVV 507
Query: 559 SLCLGCLE 566
LCLG LE
Sbjct: 508 RLCLGALE 531
>AI737603
Length = 531
Score = 37.4 bits (85), Expect = 0.017
Identities = 17/40 (42%), Positives = 21/40 (52%)
Frame = -2
Query: 302 GRSVSCWWSEVRRLRDGVGDGGSGWFGDQVLRRVGDGVVT 341
GR S W + +R GVG G GWF + + R VG G T
Sbjct: 125 GRRDSSVWRMLSAIRSGVGSGEGGWFEENIRRMVGGGSST 6
>BE943274
Length = 597
Score = 32.7 bits (73), Expect = 0.43
Identities = 15/31 (48%), Positives = 19/31 (60%)
Frame = -2
Query: 594 LVES*KSYLCFWDSAVVVKPSCLFGYWLRFF 624
+ ES* Y CFW +VV+P +FGY L F
Sbjct: 323 VAES*ACYFCFWYPNMVVEPLNVFGY*LALF 231
>TC82763
Length = 1376
Score = 32.7 bits (73), Expect = 0.43
Identities = 22/78 (28%), Positives = 36/78 (45%)
Frame = +1
Query: 243 WQTVCLKKECGGLGVRKLKEFNLALLGKWCWRMLVDRGGLWYRVLVARYGELGGRLEVGG 302
W +C + GG+ +R L+ FN+A + W + + + VL +YG LE
Sbjct: 244 WNWIC-RPIRGGIRIRNLRFFNIA-VR**SWTLKTETNSFHFMVLFHKYGSW---LEGVN 408
Query: 303 RSVSCWWSEVRRLRDGVG 320
S S WW + + G+G
Sbjct: 409 CSSSLWWRYLSSIDTGMG 462
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.348 0.155 0.570
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 23,842,144
Number of Sequences: 36976
Number of extensions: 397457
Number of successful extensions: 4276
Number of sequences better than 10.0: 50
Number of HSP's better than 10.0 without gapping: 2420
Number of HSP's successfully gapped in prelim test: 162
Number of HSP's that attempted gapping in prelim test: 1748
Number of HSP's gapped (non-prelim): 2743
length of query: 624
length of database: 9,014,727
effective HSP length: 102
effective length of query: 522
effective length of database: 5,243,175
effective search space: 2736937350
effective search space used: 2736937350
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 14 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 38 (21.7 bits)
S2: 61 (28.1 bits)
Medicago: description of AC134967.10


