

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC134521.1 - phase: 0 /pseudo
(324 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
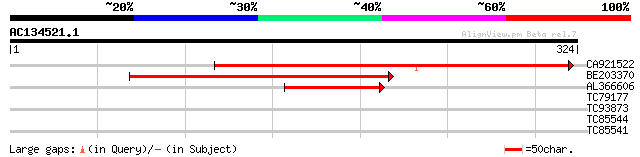
Score E
Sequences producing significant alignments: (bits) Value
CA921522 203 6e-53
BE203370 183 7e-47
AL366606 73 2e-13
TC79177 28 6.3
TC93873 similar to SP|P57703|METE_PSEAE 5-methyltetrahydropteroy... 28 6.3
TC85544 homologue to GP|6984142|gb|AAF34771.1| 40S ribosomal pro... 27 8.2
TC85541 homologue to GP|166867|gb|AAA32866.1|| ribosomal protein... 27 8.2
>CA921522
Length = 767
Score = 203 bits (517), Expect = 6e-53
Identities = 100/208 (48%), Positives = 137/208 (65%), Gaps = 3/208 (1%)
Frame = +3
Query: 118 EVADIKDKFTTKDGLPCLPYNFLHQKATTCFEKSKTDDFESILALLIYGIVLFPKVDKFV 177
E ++K T+ +FL+++ T + F SILALL+YG+VLFP +D FV
Sbjct: 3 ETDEVKTHLITRGKFLGFSTDFLYERTTFFDKMGVAYAFNSILALLVYGLVLFPSLDNFV 182
Query: 178 DMNAIRIFLTQNPVPVLLADTYVAIHDRTSHGKGTIVCCTPLLHRWITSHLPRP---SIR 234
D+ AI+IFL++NPVP LL DTY++IH RT G+GTI+CC LL+RWITSHLPR +
Sbjct: 183 DIKAIQIFLSRNPVPTLLGDTYLSIHRRTQAGRGTILCCAQLLYRWITSHLPRTPRFTTN 362
Query: 235 PEPIPWSQKLMTWTPKDIIWFNPSCDPEFIIDSCGDFNNVPLLGTQGGISYSPILARRQF 294
PE + WS++LM+ TP +++W++ D IIDSCG F NVPLLG +GGISY+P LAR QF
Sbjct: 363 PENLLWSKRLMSLTPAEVVWYDRVYDKGTIIDSCGKFANVPLLGMEGGISYNPTLARHQF 542
Query: 295 EYYTEMKPVYLILDRDFFLYKKDDANQR 322
Y E KP+ + L+ ++L D R
Sbjct: 543 GYPMERKPLSIYLENVYYLNADDSTGMR 626
>BE203370
Length = 454
Score = 183 bits (465), Expect = 7e-47
Identities = 93/151 (61%), Positives = 111/151 (72%)
Frame = +1
Query: 69 HGFTFPDFQLFPTLEEYSHWVGLPVLDEVPFHAFGPIPKIPTIAKALHLEVADIKDKFTT 128
H FTFPD+QL PTLEEYS+ VG PVLD+VPF F PIPK TIA ALHLE + IK K T
Sbjct: 1 HCFTFPDYQLAPTLEEYSYLVG*PVLDKVPFTGFEPIPKAATIADALHLETSLIKAKLTI 180
Query: 129 KDGLPCLPYNFLHQKATTCFEKSKTDDFESILALLIYGIVLFPKVDKFVDMNAIRIFLTQ 188
K LP LP FL+Q+A+ + D F SILALLIYG+VLFP VD +VD++AI++FLT+
Sbjct: 181 KGNLPSLPTKFLYQQASDFAKVDNVDAFYSILALLIYGLVLFPNVDNYVDIHAIQMFLTK 360
Query: 189 NPVPVLLADTYVAIHDRTSHGKGTIVCCTPL 219
NPVP LLAD Y +IHDR G+G I+ C PL
Sbjct: 361 NPVPTLLADMYHSIHDRNQVGRGAILGCAPL 453
>AL366606
Length = 393
Score = 72.8 bits (177), Expect = 2e-13
Identities = 34/57 (59%), Positives = 43/57 (74%)
Frame = -1
Query: 158 SILALLIYGIVLFPKVDKFVDMNAIRIFLTQNPVPVLLADTYVAIHDRTSHGKGTIV 214
+I L G++LFP +D FVDMNAI +F ++NPVP LLADTY AIHDRT G+G I+
Sbjct: 381 AIXLCLSTGLILFPNLDNFVDMNAIEVFHSKNPVPTLLADTYHAIHDRTLKGRGYIL 211
>TC79177
Length = 1704
Score = 27.7 bits (60), Expect = 6.3
Identities = 16/47 (34%), Positives = 26/47 (55%), Gaps = 1/47 (2%)
Frame = +3
Query: 179 MNAIRIFLTQNPVPVLLADTYVAI-HDRTSHGKGTIVCCTPLLHRWI 224
+N +R+ +T N +LL Y +I +RTS GT +C ++ WI
Sbjct: 819 VNHMRMLVTVNHKRMLLTVNY*SILTNRTSIAAGTCICPAMMIGTWI 959
>TC93873 similar to SP|P57703|METE_PSEAE
5-methyltetrahydropteroyltriglutamate--homocysteine
methyltransferase (EC 2.1.1.14), partial (14%)
Length = 517
Score = 27.7 bits (60), Expect = 6.3
Identities = 11/33 (33%), Positives = 18/33 (54%)
Frame = +2
Query: 227 HLPRPSIRPEPIPWSQKLMTWTPKDIIWFNPSC 259
H PR + E Q+++ + K++IW NP C
Sbjct: 206 HSPRVPSKEELKARLQEILKYISKNLIWVNPDC 304
>TC85544 homologue to GP|6984142|gb|AAF34771.1| 40S ribosomal protein S11
{Euphorbia esula}, partial (76%)
Length = 1335
Score = 27.3 bits (59), Expect = 8.2
Identities = 15/46 (32%), Positives = 24/46 (51%), Gaps = 2/46 (4%)
Frame = -2
Query: 108 IPTIAK--ALHLEVADIKDKFTTKDGLPCLPYNFLHQKATTCFEKS 151
+P++A L+ D+K++F GL LP F H+ CF K+
Sbjct: 173 VPSMASLGVLNPRPTDLKNRFPPFPGLFPLPLFFEHRNTLGCFRKA 36
>TC85541 homologue to GP|166867|gb|AAA32866.1|| ribosomal protein S11
(probable start codon at bp 67) {Arabidopsis thaliana},
partial (87%)
Length = 883
Score = 27.3 bits (59), Expect = 8.2
Identities = 15/46 (32%), Positives = 24/46 (51%), Gaps = 2/46 (4%)
Frame = -1
Query: 108 IPTIAK--ALHLEVADIKDKFTTKDGLPCLPYNFLHQKATTCFEKS 151
+P++A L+ D+K++F GL LP F H+ CF K+
Sbjct: 268 VPSMASLGVLNPRPTDLKNRFPPFPGLFPLPLFFEHRNTLGCFRKA 131
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.324 0.144 0.458
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 11,709,183
Number of Sequences: 36976
Number of extensions: 201338
Number of successful extensions: 929
Number of sequences better than 10.0: 14
Number of HSP's better than 10.0 without gapping: 921
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 926
length of query: 324
length of database: 9,014,727
effective HSP length: 96
effective length of query: 228
effective length of database: 5,465,031
effective search space: 1246027068
effective search space used: 1246027068
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (22.0 bits)
S2: 58 (26.9 bits)
Medicago: description of AC134521.1


