

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC134322.2 + phase: 0 /pseudo
(840 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
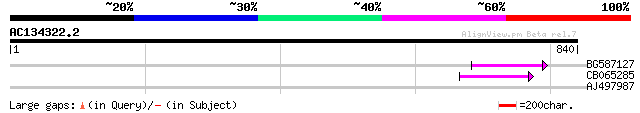
BG587127 weakly similar to PIR|H84506|H84 probable retroelement ... 72 1e-12
CB065285 weakly similar to PIR|A84500|A84 probable retroelement ... 67 3e-11
AJ497987 weakly similar to GP|9927273|dbj Similar to Arabidopsis... 42 0.001
>BG587127 weakly similar to PIR|H84506|H84 probable retroelement pol
polyprotein [imported] - Arabidopsis thaliana, partial
(13%)
Length = 415
Score = 71.6 bits (174), Expect = 1e-12
Identities = 39/112 (34%), Positives = 60/112 (52%)
Frame = +3
Query: 685 GVTLEGPYGVLIEQSLRFDFKSSNNQAEYEALFAGMRLALKVVVKELEAKSDSQLVFRQV 744
G+ L P +++QS R +F +SNN+ YEAL AG+RLA + ++ + A DSQLV Q
Sbjct: 3 GIRLTSPTNEILKQSFRLEFHASNNETNYEALIAGVRLAHGLKIRNIHAYCDSQLVASQF 182
Query: 745 TGNFQTKDPQLLMYLESVMYLTQKFVSFKYYMCLESRTVEQTCWLSWLAPSD 796
+G ++ +D + YL+ V L QK F S V+ + + SD
Sbjct: 183 SGEYEARDELMDTYLKLVQKLAQKLDYFALTRIPRSENVQADALAALASSSD 338
>CB065285 weakly similar to PIR|A84500|A84 probable retroelement gag/pol
polyprotein [imported] - Arabidopsis thaliana, partial
(2%)
Length = 592
Score = 67.0 bits (162), Expect = 3e-11
Identities = 36/109 (33%), Positives = 54/109 (49%)
Frame = -2
Query: 667 AKWVLSVDGSSNLKGSGAGVTLEGPYGVLIEQSLRFDFKSSNNQAEYEALFAGMRLALKV 726
+KW L DG+ N G G G + P G I + R F+ +NN AEYEA G+ A+ +
Sbjct: 540 SKWGLVFDGAVNAYGKGIGAVIVSPQGHYIPFTARILFECTNNMAEYEACIFGIEEAIDM 361
Query: 727 VVKELEAKSDSQLVFRQVTGNFQTKDPQLLMYLESVMYLTQKFVSFKYY 775
+K L+ DS LV Q+ G ++T L+ Y + L F + +
Sbjct: 360 RIKHLDIYGDSALVINQIKGEWETHHANLIPYRDYARRLLTYFTKVELH 214
>AJ497987 weakly similar to GP|9927273|dbj Similar to Arabidopsis thaliana
chromosome II BAC F26H6; putative retroelement pol
polyprotein, partial (1%)
Length = 636
Score = 42.0 bits (97), Expect = 0.001
Identities = 22/52 (42%), Positives = 28/52 (53%)
Frame = -2
Query: 669 WVLSVDGSSNLKGSGAGVTLEGPYGVLIEQSLRFDFKSSNNQAEYEALFAGM 720
W L DG+ N+ G+G G L P G I + R F +NN AEYEA G+
Sbjct: 290 WGLIFDGAVNVYGNGIGAVLLTPKGAHIPFTARLRFDCTNNIAEYEACIMGI 135
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.358 0.156 0.572
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 26,587,149
Number of Sequences: 36976
Number of extensions: 368965
Number of successful extensions: 3485
Number of sequences better than 10.0: 6
Number of HSP's better than 10.0 without gapping: 1012
Number of HSP's successfully gapped in prelim test: 109
Number of HSP's that attempted gapping in prelim test: 2452
Number of HSP's gapped (non-prelim): 1178
length of query: 840
length of database: 9,014,727
effective HSP length: 104
effective length of query: 736
effective length of database: 5,169,223
effective search space: 3804548128
effective search space used: 3804548128
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 14 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 37 (21.8 bits)
S2: 63 (28.9 bits)
Medicago: description of AC134322.2


