

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC134322.16 - phase: 0
(690 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
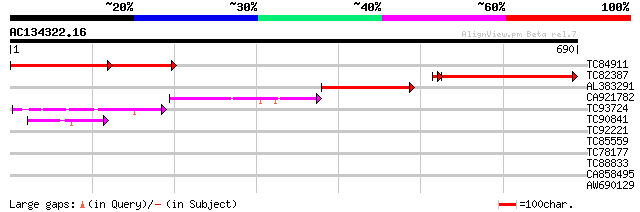
Sequences producing significant alignments: (bits) Value
TC84911 similar to PIR|G96635|G96635 hypothetical protein T7P1.1... 254 1e-95
TC82387 similar to PIR|G96635|G96635 hypothetical protein T7P1.1... 321 8e-90
AL383291 similar to PIR|G96635|G96 hypothetical protein T7P1.15 ... 214 1e-55
CA921782 similar to GP|9082326|gb| polyadenylation cleavage/spec... 98 1e-20
TC93724 similar to SP|Q9LKF9|CPSB_ARATH Cleavage and polyadenyla... 55 1e-07
TC90841 similar to GP|9758283|dbj|BAB08807.1 gene_id:MLE2.5~simi... 42 6e-04
TC92221 similar to GP|9758283|dbj|BAB08807.1 gene_id:MLE2.5~simi... 41 0.001
TC85559 similar to PIR|T07012|T07012 acetyl-CoA carboxylase (EC ... 32 0.62
TC78177 similar to PIR|T07145|T07145 epoxide hydrolase homolog -... 31 1.8
TC88833 similar to PIR|JQ1674|JQ1674 protein kinase TMK1 (EC 2.7... 30 3.1
CA858495 30 4.0
AW690129 similar to GP|21554386|gb| unknown {Arabidopsis thalian... 29 5.3
>TC84911 similar to PIR|G96635|G96635 hypothetical protein T7P1.15
[imported] - Arabidopsis thaliana, partial (24%)
Length = 617
Score = 254 bits (648), Expect(2) = 1e-95
Identities = 124/124 (100%), Positives = 124/124 (100%)
Frame = +3
Query: 2 SSVKKRESNGGTINRETEDQLIVTPLGAGNEVGRSCVYMTYKGKTVLFDCGIHPGYSGMA 61
SSVKKRESNGGTINRETEDQLIVTPLGAGNEVGRSCVYMTYKGKTVLFDCGIHPGYSGMA
Sbjct: 3 SSVKKRESNGGTINRETEDQLIVTPLGAGNEVGRSCVYMTYKGKTVLFDCGIHPGYSGMA 182
Query: 62 ALPYFDEIDPSTVDVLLITHFHLDHAASLPYFLEKTTFKGRVFMTYATKAIYKLLLSDYV 121
ALPYFDEIDPSTVDVLLITHFHLDHAASLPYFLEKTTFKGRVFMTYATKAIYKLLLSDYV
Sbjct: 183 ALPYFDEIDPSTVDVLLITHFHLDHAASLPYFLEKTTFKGRVFMTYATKAIYKLLLSDYV 362
Query: 122 KVSK 125
KVSK
Sbjct: 363 KVSK 374
Score = 115 bits (288), Expect(2) = 1e-95
Identities = 61/85 (71%), Positives = 66/85 (76%), Gaps = 2/85 (2%)
Frame = +1
Query: 121 VKVSKVSVDDMLYDEQDINRSMDKIEVIDFHQTVEVNGIRFWCYTAGHVLGAA-MFMVDI 179
+K +VSVDDMLYDEQDINRSMDKIEVIDFHQTVEVNGIRFWCYTAGHVL ++ +
Sbjct: 361 LKSVRVSVDDMLYDEQDINRSMDKIEVIDFHQTVEVNGIRFWCYTAGHVLWCCYVYGWTL 540
Query: 180 AGVRVLYTGDYSRE-EDRHLRAAET 203
RVLYTGDYSRE AAET
Sbjct: 541 LVFRVLYTGDYSREGRQTSFEAAET 615
>TC82387 similar to PIR|G96635|G96635 hypothetical protein T7P1.15
[imported] - Arabidopsis thaliana, partial (14%)
Length = 824
Score = 321 bits (823), Expect(2) = 8e-90
Identities = 165/165 (100%), Positives = 165/165 (100%)
Frame = +3
Query: 526 GAFCVIQSRLKQIYESVEPSVDEESGVPMLLVHDRVTVKHESEKHVSLHWASDPINDMVS 585
GAFCVIQSRLKQIYESVEPSVDEESGVPMLLVHDRVTVKHESEKHVSLHWASDPINDMVS
Sbjct: 54 GAFCVIQSRLKQIYESVEPSVDEESGVPMLLVHDRVTVKHESEKHVSLHWASDPINDMVS 233
Query: 586 DSVVALVLNINRDLPKIVAESDATKIEEENEKKTEKVMQALLNSLFGNVKVGENGKLIIN 645
DSVVALVLNINRDLPKIVAESDATKIEEENEKKTEKVMQALLNSLFGNVKVGENGKLIIN
Sbjct: 234 DSVVALVLNINRDLPKIVAESDATKIEEENEKKTEKVMQALLNSLFGNVKVGENGKLIIN 413
Query: 646 IDGNVAELNKESGEVESENEGLKERVRTAFRRIQSSVKPIPLSAP 690
IDGNVAELNKESGEVESENEGLKERVRTAFRRIQSSVKPIPLSAP
Sbjct: 414 IDGNVAELNKESGEVESENEGLKERVRTAFRRIQSSVKPIPLSAP 548
Score = 28.5 bits (62), Expect(2) = 8e-90
Identities = 12/12 (100%), Positives = 12/12 (100%)
Frame = +2
Query: 515 NVTQRITIPYSG 526
NVTQRITIPYSG
Sbjct: 20 NVTQRITIPYSG 55
>AL383291 similar to PIR|G96635|G96 hypothetical protein T7P1.15 [imported] -
Arabidopsis thaliana, partial (16%)
Length = 341
Score = 214 bits (544), Expect = 1e-55
Identities = 108/113 (95%), Positives = 108/113 (95%)
Frame = +1
Query: 380 KEVTLMNGLSAPLHMQVHYISFSAHADSAQTSAFLEELNPPNIILVHGAANEMGRLKQKL 439
KEVTLMNGLSAPLHMQVHYISFSAHADSAQTSAFLEELNPPNIILVHGA NEMGRLKQ L
Sbjct: 1 KEVTLMNGLSAPLHMQVHYISFSAHADSAQTSAFLEELNPPNIILVHGAXNEMGRLKQXL 180
Query: 440 MTQFADRNTKILTPKNCQSVEMYFNSQKMAKTIGKLAEKTPEVGETVSGLLVK 492
MT FADRNTKILTPKNCQSVEMYFNSQ MA TIGKLAEKTPEVGETVSGLLVK
Sbjct: 181 MTXFADRNTKILTPKNCQSVEMYFNSQXMAXTIGKLAEKTPEVGETVSGLLVK 339
>CA921782 similar to GP|9082326|gb| polyadenylation cleavage/specificity
factor 100 kDa subunit {Arabidopsis thaliana}, partial
(31%)
Length = 792
Score = 97.8 bits (242), Expect = 1e-20
Identities = 63/193 (32%), Positives = 99/193 (50%), Gaps = 8/193 (4%)
Frame = +1
Query: 195 DRHLRAAETPQFSPDVCIIESTYGVQHHQP-RHTREKRFTDVIHSTISQGGRVLIPAYAL 253
+RHL F +I Y ++QP R ++K F D++ T+ GG VL+P
Sbjct: 1 ERHLNGTVLGSFVRPAVLITDAYNALNNQPYRRQKDKEFGDILKKTLRAGGNVLLPVDTA 180
Query: 254 GRAQELLLILDEYWANHPELQNIPIYYASPLAKKCLTVYETYTLSMNDRI----QNAKSN 309
GR EL+L+L+ YWA+ E N PIY+ + +A + +++ M+D I + + N
Sbjct: 181 GRILELILMLESYWAD--ENLNYPIYFLTYVASSTIDYVKSFLEWMSDSIAKSFEQTREN 354
Query: 310 PFAFKHISALSS---IDIFKDVGPSVVMASPGGLQSGLSRQLFDMWCSDKKNSCVIPGYV 366
F K+I+ L S +D D GP VV+AS L++G S +F W +D KN +
Sbjct: 355 IFLLKNITLLVSKADLDNAPD-GPKVVLASMASLEAGFSHDIFVEWGNDVKNLVLFTERG 531
Query: 367 VEGTLAKTILNEP 379
GTLA+ + +P
Sbjct: 532 QFGTLARMLQADP 570
>TC93724 similar to SP|Q9LKF9|CPSB_ARATH Cleavage and polyadenylation
specificity factor 100 kDa subunit (CPSF 100 kDa
subunit)., partial (23%)
Length = 664
Score = 54.7 bits (130), Expect = 1e-07
Identities = 53/196 (27%), Positives = 94/196 (47%), Gaps = 8/196 (4%)
Frame = +1
Query: 4 VKKRESNGGTINRETEDQLIVTPL-GAGNEVGRSCVYMTYKGKTVLFDCGIHPGYSGMAA 62
+K+R+ G ++ VTPL G NE S + ++ +L DCG + +
Sbjct: 115 IKRRQIMGTSVQ--------VTPLCGVYNENPLSYL-VSIDSFNILIDCGWNDHFDPSLL 267
Query: 63 LPYFDEIDPSTVDVLLITHFHLDHAASLPYFLEKTTFKGRVFMTYATKAIYKL-LLSDYV 121
P ST+D +L++H H A+LPY ++ V Y+T+ +Y+L LL+ Y
Sbjct: 268 QPLSRVA--STIDAVLLSHPDTLHLAALPYAIKHLGLSAPV---YSTEPVYRLGLLTMYD 432
Query: 122 K-VSKVSVDDM-LYDEQDINRSMDKIEVIDF----HQTVEVNGIRFWCYTAGHVLGAAMF 175
+S+ V D L+ DI+ + + + + H + + GI +TAGH+LG ++
Sbjct: 433 HFLSRKQVSDFDLFTLDDIDSAFQTVTRLTYXQNHHLSXKGEGIVIAPHTAGHLLGGTIW 612
Query: 176 MVDIAGVRVLYTGDYS 191
+ G V+Y D++
Sbjct: 613 KITKDGEDVIYAXDFN 660
>TC90841 similar to GP|9758283|dbj|BAB08807.1 gene_id:MLE2.5~similar to
unknown protein~sp|P54122 {Arabidopsis thaliana},
partial (27%)
Length = 658
Score = 42.4 bits (98), Expect = 6e-04
Identities = 29/109 (26%), Positives = 48/109 (43%), Gaps = 10/109 (9%)
Frame = +1
Query: 22 LIVTPLGAGNEVGRSCVYMTYKGKTVLFDCGI-HPGYSGMAALPYFDEIDPST------- 73
L V P+G E+G +C+ + + +L D GI P Y + +I P T
Sbjct: 325 LRVLPIGGLGEIGMNCMLVGNHDRYILIDAGIMFPDYDDLGV----QKIIPDTTFIRKWS 492
Query: 74 --VDVLLITHFHLDHAASLPYFLEKTTFKGRVFMTYATKAIYKLLLSDY 120
++ L+ITH H DH +LP+ + +F + T + K L +
Sbjct: 493 HKIEALVITHGHEDHIGALPWVIPALDSNTPIFASSFTMELIKKRLKXH 639
>TC92221 similar to GP|9758283|dbj|BAB08807.1 gene_id:MLE2.5~similar to
unknown protein~sp|P54122 {Arabidopsis thaliana},
partial (37%)
Length = 956
Score = 41.2 bits (95), Expect = 0.001
Identities = 28/109 (25%), Positives = 50/109 (45%), Gaps = 10/109 (9%)
Frame = +3
Query: 22 LIVTPLGAGNEVGRSCVYMTYKGKTVLFDCGI-HPGYSGMAALPYFDEIDPST------- 73
L V P+G E+G +C+ + + +L D G+ PG + +I P T
Sbjct: 387 LRVLPIGGLGEIGMNCMLVGNHDRYILVDAGVMFPGDDELGV----QKIIPDTTFIKKWS 554
Query: 74 --VDVLLITHFHLDHAASLPYFLEKTTFKGRVFMTYATKAIYKLLLSDY 120
++ ++ITH H DH +LP+ + + VF + T + + L D+
Sbjct: 555 HKIEAVVITHGHEDHIGALPWVIPMLDSQTPVFASSFTMELIRKRLKDH 701
>TC85559 similar to PIR|T07012|T07012 acetyl-CoA carboxylase (EC 6.4.1.2) -
potato chloroplast, partial (66%)
Length = 5011
Score = 32.3 bits (72), Expect = 0.62
Identities = 13/52 (25%), Positives = 31/52 (59%)
Frame = +1
Query: 552 VPMLLVHDRVTVKHESEKHVSLHWASDPINDMVSDSVVALVLNINRDLPKIV 603
VP+ ++ R+T++ + + + + P+ D+++D + L+L++ DL IV
Sbjct: 2557 VPLYIISLRITMRIIRRRRIKMKNYTSPVIDLITDLITDLMLDLKSDLKLIV 2712
>TC78177 similar to PIR|T07145|T07145 epoxide hydrolase homolog - soybean,
partial (89%)
Length = 1037
Score = 30.8 bits (68), Expect = 1.8
Identities = 14/27 (51%), Positives = 22/27 (80%)
Frame = +2
Query: 134 DEQDINRSMDKIEVIDFHQTVEVNGIR 160
++++INR M++IE H+TVEVNGI+
Sbjct: 5 NKKEINREMERIE----HRTVEVNGIK 73
>TC88833 similar to PIR|JQ1674|JQ1674 protein kinase TMK1 (EC 2.7.1.-)
receptor type precursor - Arabidopsis thaliana, partial
(36%)
Length = 1382
Score = 30.0 bits (66), Expect = 3.1
Identities = 31/137 (22%), Positives = 55/137 (39%), Gaps = 1/137 (0%)
Frame = +3
Query: 352 WCSDKKNSCVIPGYVVEGTLAKTILNEPKEVTLMNGLSAPLHMQVHYISFSAHADSAQTS 411
+C D ++ Y+ +GTL++ I N P+E G + L + + + S
Sbjct: 216 YCLDGNEKLLVYEYMPQGTLSRYIFNWPEEGLEPLGWNKRLVIALDVARGVEYLHSLAHQ 395
Query: 412 AFL-EELNPPNIILVHGAANEMGRLKQKLMTQFADRNTKILTPKNCQSVEMYFNSQKMAK 470
+F+ +L P NI+ L + + AD L P+ S+E ++A
Sbjct: 396 SFIHRDLKPSNIL-----------LGDDMRAKVADFGLVRLAPEGKASIE-----TRIAG 527
Query: 471 TIGKLAEKTPEVGETVS 487
T G LA + G +
Sbjct: 528 TFGYLAPEYAVTGRVTT 578
>CA858495
Length = 763
Score = 29.6 bits (65), Expect = 4.0
Identities = 51/232 (21%), Positives = 101/232 (42%), Gaps = 12/232 (5%)
Frame = +3
Query: 464 NSQKMAKTIGKLAEKTPEVGETVSGLLVK--------KGFTYQI-MAPDDLHVFSQLSTA 514
N+ + K +G LAEKT + +GL+ K +G ++ + +L FS+
Sbjct: 27 NNSLLQKEVG-LAEKTNLLLNEKAGLVEKVNMLLNEKEGLEQKVNILESNLSSFSEKEAG 203
Query: 515 NV--TQRITIPYSGAFCVIQSRLKQIYESVEPSVDEESGVPMLLVHDRVTVKHESEKHVS 572
+V T + G ++ +L + ++ ++E+G+ M + + +K
Sbjct: 204 SVDTTNLLLKEKEG----LEQKLNILESNLSSFSEKETGLEMRIAQLQSETNSLLQKETG 371
Query: 573 LHWASDPINDMVSDSVVALVLNINRDLPKIV-AESDATKIEEENEKKTEKVMQALLNSLF 631
+ N ++++ + L L ++ KI+ AESD + E+ E TE+V+ L
Sbjct: 372 F---VEKTNQLLNEKNI-LSLKVDSLERKIIHAESDLSSFVEK-ENSTEEVISNL----- 521
Query: 632 GNVKVGENGKLIINIDGNVAELNKESGEVESENEGLKERVRTAFRRIQSSVK 683
NG + + + G VAEL + + EN+ L+E ++ +S K
Sbjct: 522 -------NGSISM-LQGQVAELEESKNNLLLENQQLRENSSSSSASQDASAK 653
>AW690129 similar to GP|21554386|gb| unknown {Arabidopsis thaliana}, partial
(7%)
Length = 464
Score = 29.3 bits (64), Expect = 5.3
Identities = 26/88 (29%), Positives = 41/88 (46%), Gaps = 4/88 (4%)
Frame = +2
Query: 565 HESEKHVSLHWASDPINDMVSDSVVALVLNINRDLPKIVAESDATKIEEENEKKTEKV-M 623
H+S L SDP+N S S V + N N+ +P ++ ++ EE+ K E+V +
Sbjct: 98 HQSRSLDILSSESDPLNGTSSSSSVVVETNQNK-VPLLLRSTNNVVEEEKKSKNEEQVGL 274
Query: 624 QALLNSLFGNVKVGE---NGKLIINIDG 648
L + +GN V + K I DG
Sbjct: 275 APLWDDGYGNRTVEDYFAASKEICKFDG 358
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.317 0.134 0.383
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 18,494,352
Number of Sequences: 36976
Number of extensions: 239476
Number of successful extensions: 1222
Number of sequences better than 10.0: 24
Number of HSP's better than 10.0 without gapping: 1213
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1218
length of query: 690
length of database: 9,014,727
effective HSP length: 103
effective length of query: 587
effective length of database: 5,206,199
effective search space: 3056038813
effective search space used: 3056038813
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 62 (28.5 bits)
Medicago: description of AC134322.16


