

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
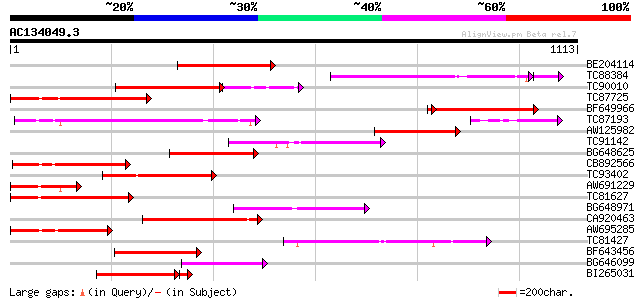
Query= AC134049.3 - phase: 0
(1113 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
BE204114 weakly similar to GP|15553678|gb I2C-5 {Lycopersicon pi... 300 3e-81
TC88384 homologue to GP|21626661|gb|AAF47055.2 CG13560-PA {Droso... 200 2e-62
TC90010 similar to GP|5817349|gb|AAD52718.1| putative NBS-LRR ty... 194 9e-61
TC87725 similar to GP|5817341|gb|AAD52714.1| putative NBS-LRR ty... 226 5e-59
BF649966 220 4e-58
TC87193 similar to GP|21616922|gb|AAM66423.1 NBS-LRR protein {Or... 223 4e-58
AW125982 201 1e-51
TC91142 weakly similar to PIR|T02213|T02213 NBS-LRR type resista... 198 8e-51
BG648625 weakly similar to GP|20385440|gb| resistance gene analo... 194 2e-49
CB892566 weakly similar to GP|15289776|db Lycopersicon esculentu... 184 1e-46
TC93402 weakly similar to GP|16588610|gb|AAL26857.1 NBS-type put... 178 9e-45
AW691229 175 8e-44
TC81627 weakly similar to GP|18652501|gb|AAL77135.1 Putative NBS... 173 4e-43
BG648971 similar to GP|20514808|gb| Putative NBS-LRR type resist... 173 4e-43
CA920463 weakly similar to GP|2852684|gb|A resistance protein ca... 173 4e-43
AW695285 similar to GP|15487944|gb| NBS/LRR resistance protein-l... 161 1e-39
TC81427 weakly similar to GP|11994216|dbj|BAB01338. disease resi... 152 7e-37
BF643456 weakly similar to GP|14348622|gb NBS-LRR resistance-lik... 149 6e-36
BG646099 weakly similar to GP|13957628|gb putative resistance pr... 139 5e-33
BI265031 weakly similar to GP|13872974|db putative NBS-LRR type ... 124 1e-31
>BE204114 weakly similar to GP|15553678|gb I2C-5 {Lycopersicon
pimpinellifolium}, partial (3%)
Length = 583
Score = 300 bits (767), Expect = 3e-81
Identities = 142/193 (73%), Positives = 167/193 (85%)
Frame = +3
Query: 329 KQRAFGPNEVQQKELVIVGKEIIKKCGGFPLAAIALGSLLRFKREEKEWLYVKESKLWNL 388
+QRAFGPNE + ++LV++GKEI+KKC PLAAIALGSLLRFKREEKEWLYVKESKLW+L
Sbjct: 3 RQRAFGPNEAEDEKLVVIGKEILKKCV*VPLAAIALGSLLRFKREEKEWLYVKESKLWSL 182
Query: 389 QGEAYVMPALRLSYLHLPVKLRQCFSFCALFPKDEIISKQLLIDLWTANGFISSNQMLEA 448
+GE YV AL+LSYL+LPVKLRQCFSFCALFPKDEI+SK +I+LW ANGFISSNQML+A
Sbjct: 183 EGEDYVKSALKLSYLNLPVKLRQCFSFCALFPKDEIMSKHFMIELWIANGFISSNQMLDA 362
Query: 449 DDIGNEVWNELYWRSFFENTENVGFGQITIFKMHDLVHDLAGSVTQDVCCITDDNSMRTM 508
+ +GNEVWNELYWRSFF++TE FGQIT FKMHDLVH+LA SVT++V CIT +N + T+
Sbjct: 363 EGVGNEVWNELYWRSFFQDTETDEFGQITSFKMHDLVHELAESVTREVWCITYNNDLPTV 542
Query: 509 SEETRHLLIYNRN 521
SE RHL IY N
Sbjct: 543 SESIRHLFIYKEN 581
>TC88384 homologue to GP|21626661|gb|AAF47055.2 CG13560-PA {Drosophila
melanogaster}, partial (3%)
Length = 1570
Score = 200 bits (509), Expect(2) = 2e-62
Identities = 153/413 (37%), Positives = 224/413 (54%), Gaps = 15/413 (3%)
Frame = -3
Query: 631 LQNLSLRDCDSLTSLPRQIGKLTSLNTLSKYIVGEERGFLLEELGQLNLKGQLHIKNLER 690
L++L ++DC+SL S+P +I KLTSL TLS +IV + GF L EL L L G+LHIK LE
Sbjct: 1490 LRHLVIKDCNSLYSMPSKISKLTSLKTLSIFIVVLKEGFGLAELNDLQLGGRLHIKGLEN 1311
Query: 691 LKSVTDAKKANM-SRKKLNQLWLSWERNEVSQ-LQENVEQILEALQPYAQKLYSFGVGGY 748
+ S DAK+AN+ +K+LN+L+LSW + SQ + +VEQ+LEAL+P+ L FG+ GY
Sbjct: 1310 VSSEWDAKEANLIGKKELNRLYLSWGSHANSQGIDTDVEQVLEALEPHT-GLKGFGIEGY 1134
Query: 749 TGAYFPQWISIPS-LNDLKSLELVDCKSCLNLPELWKLPSLKYLKLSNMIHVIYL---FH 804
G +FP W+ S L L + +C +C LP L KLP L L + M + Y+ +
Sbjct: 1133 VGIHFPHWMRNASILEGLVDITFYNCNNCQRLPPLGKLPCLTTLYVFGMRDLKYIDDDIY 954
Query: 805 ESYDGEGLMALKTLFLEKLPNLIGLSREERV-MFPRLKALEITECPNLLGLPCLPSLSDL 863
ES ++LK L L LPNL + + E V M P+L I+ P L LP LPS+ L
Sbjct: 953 ESTSKRAFISLKNLTLLGLPNLERMLKAEGVEMLPQLSYFNISNVPK-LALPSLPSIELL 777
Query: 864 YIQGKYNQQLPSSIHKLGSLESLHFSDNEELIYFPDGILRNLASPLKTLGFHRHSKLKML 923
+ + LP +K+ SL FP+ I+ ++ + LK L KLK+L
Sbjct: 776 DVGENKYRFLPQ--YKVVSL-------------FPERIVCSMHN-LKLLIIGNFHKLKVL 645
Query: 924 PTEMIHIHALQQLYINDCRNIEELPNEVMQRLHSLKELDIVGCDKL-KLSSDFQYLTCLE 982
P ++ + L++L+I+ C +E +Q + SL+ L I C KL LS L LE
Sbjct: 644 PDDLHCLSVLEELHISRCDELESFSMHALQGMISLRVLTIDLCGKLISLSEGMGDLASLE 465
Query: 983 TLAIGSCSEVEGFHEALQHMTTLKSLTLSD-------LPNLEYLPECIGNLTL 1028
L I CS++ + +T+L+ + +S L LE +P + NLTL
Sbjct: 464 RLVIHGCSQLV-LPSKMNKLTSLRQVNISHSGRNNRILQGLEVIPS-LQNLTL 312
Score = 58.5 bits (140), Expect(2) = 2e-62
Identities = 29/60 (48%), Positives = 33/60 (54%)
Frame = -2
Query: 1028 LLHEINIYSCPKLACLPTSIQQISGLEILSIHDCSKLEKRCQKEIGEDWPKIVHVQYIEI 1087
L E SC LP S Q + L L I CSKLEKRC+K GEDW KI HV +E+
Sbjct: 267 LFKE*KFISCTNAKSLPNSFQNLINLHTLLIVGCSKLEKRCKKGTGEDWQKIAHVPELEL 88
>TC90010 similar to GP|5817349|gb|AAD52718.1| putative NBS-LRR type disease
resistance protein {Pisum sativum}, partial (88%)
Length = 1107
Score = 194 bits (494), Expect(2) = 9e-61
Identities = 90/217 (41%), Positives = 139/217 (63%), Gaps = 3/217 (1%)
Frame = +3
Query: 208 KIVNHFELKIWVCVSEDFTLKRMTKAIIEGATKKSCEDLDLELLQRKLQDLLRRKRYLLV 267
++ HF+LK WVCVSEDF + R+TK+++E T + + +L++L+ +L+ + R KR+L V
Sbjct: 12 EVQQHFDLKAWVCVSEDFDIMRVTKSLLESVTSTTWDSKNLDVLRVELKKISREKRFLFV 191
Query: 268 LDDVWNDKQENWQRLKSVLACGGKGASILVTTRLPKVAKIMGTIPHHELSRLSDEDCWEL 327
DD+WND +W L S G G+ +++TTR KVA++ T P H+L LS+EDCW L
Sbjct: 192 FDDLWNDNYNDWNELASPFINGKPGSMVIITTRQQKVAEVAHTFPIHKLKLLSNEDCWSL 371
Query: 328 FKQRAFGPNEVQQKE---LVIVGKEIIKKCGGFPLAAIALGSLLRFKREEKEWLYVKESK 384
+ A G +E L G++I +KCGG P+AA +G LLR K + EW + S
Sbjct: 372 LSKHALGSDEFHHSSNTALEETGRKIARKCGGLPIAAKTIGGLLRSKVDITEWTSILNSN 551
Query: 385 LWNLQGEAYVMPALRLSYLHLPVKLRQCFSFCALFPK 421
+WNL+ + ++PAL LSY +LP L++CF++C++F K
Sbjct: 552 IWNLRND-NILPALHLSYQYLPSHLKRCFAYCSIFSK 659
Score = 58.9 bits (141), Expect(2) = 9e-61
Identities = 49/161 (30%), Positives = 79/161 (48%), Gaps = 2/161 (1%)
Frame = +1
Query: 419 FPKDEIISKQLLIDLWTANGFISSNQM-LEADDIGNEVWNELYWRSFFENTENVGFGQIT 477
FPKD + ++ L+ LW A GF+ +Q + +++G++ + EL RS + + G+
Sbjct: 652 FPKDYPLDRKELVLLWMAEGFLDCSQRGKKMEELGDDCFAELLSRSLIQQLSDDDRGE-- 825
Query: 478 IFKMHDLVHDLAGSVTQDVCCITDDNSMRTMSEETRHLLIYNRNSFAEANSIQLHHVKSL 537
F MHDLV+DLA V+ CC + + E RH N +LH+ K L
Sbjct: 826 KFVMHDLVNDLATFVSGKSCCRLECGD---IPENVRHFSYNQENYDIFMKFEKLHNFKCL 996
Query: 538 KTYMEFNFDVYEAGQLSPQVLNCYSLRVLLSH-RLNNLSSS 577
++++ + LS +V+N +L SH RL LS S
Sbjct: 997 RSFLFICLMTWRDNYLSFKVVN----DLLPSHKRLRVLSLS 1107
>TC87725 similar to GP|5817341|gb|AAD52714.1| putative NBS-LRR type disease
resistance protein {Pisum sativum}, partial (44%)
Length = 899
Score = 226 bits (575), Expect = 5e-59
Identities = 124/279 (44%), Positives = 176/279 (62%), Gaps = 2/279 (0%)
Frame = +3
Query: 1 MAEAVLEIVLGSLSELIRKEISLFLGFDQEFNRLASLLTTIKATLEDAEEKQFSDSEIGR 60
MA+A+L +V +L+ L + E S G + +L+ L IKA LEDAE+KQF + I
Sbjct: 69 MADALLGVVFENLTALHQNEFSTISGIKSKAQKLSDNLVHIKAVLEDAEKKQFKELSI-- 242
Query: 61 DVKDWLLKLKDAAYTLDDIMDECATEALEMEYKASKCGLSHKHIAFRYKLAKKMKRIGVW 120
K WL LKDA Y LDDI+DE + ++ ++ S L K+I FR ++ ++K I
Sbjct: 243 --KLWLQDLKDAVYVLDDILDEYSIKSGQLRGSTS---LKPKNIKFRNEIGNRLKEITRR 407
Query: 121 LDDIAAEKNKFHLTE--IVRERSGVVPDWRQTTSIVTQPLVYGRNEDKDKIVDFLVGDAS 178
LD+IA KNKF L +RE V + RQT SI+ +P V+GR DK+KIV+FL+ A
Sbjct: 408 LDNIAESKNKFSLQMGGTLREIPDQVAEGRQTGSIIAEPKVFGREVDKEKIVEFLLTQAK 587
Query: 179 EQEDLSVYPIVGLGGLGKTTLAQLVFNHDKIVNHFELKIWVCVSEDFTLKRMTKAIIEGA 238
+ + +SVYPI GLGG+GKTTL QL+FN ++ HF+ K+WVCVSE F++KR+ +I E
Sbjct: 588 DSDFISVYPIFGLGGIGKTTLVQLIFNDVRVSGHFDKKVWVCVSETFSVKRILCSIFESI 767
Query: 239 TKKSCEDLDLELLQRKLQDLLRRKRYLLVLDDVWNDKQE 277
T C D + +++ K+Q L + KRYLL LDDVWN ++
Sbjct: 768 TLXKCPDFEYAVMEGKVQGLXQGKRYLLXLDDVWNQNEQ 884
>BF649966
Length = 657
Score = 220 bits (561), Expect(2) = 4e-58
Identities = 115/207 (55%), Positives = 149/207 (71%)
Frame = +3
Query: 831 REERVMFPRLKALEITECPNLLGLPCLPSLSDLYIQGKYNQQLPSSIHKLGSLESLHFSD 890
R E+ F L L+IT+CP L GLP LPSL+D+ ++ K NQ L SSIHK SLE++ F+
Sbjct: 39 RIEKTCFHXLSTLQITKCPKLSGLPYLPSLNDMRVREKCNQGLLSSIHKHQSLETIRFAH 218
Query: 891 NEELIYFPDGILRNLASPLKTLGFHRHSKLKMLPTEMIHIHALQQLYINDCRNIEELPNE 950
NEEL+YFPD +L+NL S LK L SKL+ LPTE + ++++Q++YI+ +++ LP+E
Sbjct: 219 NEELVYFPDRMLQNLTS-LKVLDIFELSKLEKLPTEFVSLNSIQEIYISGSNSLKSLPDE 395
Query: 951 VMQRLHSLKELDIVGCDKLKLSSDFQYLTCLETLAIGSCSEVEGFHEALQHMTTLKSLTL 1010
V+Q L+SLK LDIV C K LS+ FQYLTCLE L I S SE+E HEALQHMT+L+SL L
Sbjct: 396 VLQGLNSLKILDIVRCPKFNLSASFQYLTCLEKLMIESSSEIEXLHEALQHMTSLQSLIL 575
Query: 1011 SDLPNLEYLPECIGNLTLLHEINIYSC 1037
DLPNL LP+ GNL LLHE+ I C
Sbjct: 576 CDLPNLPSLPDWXGNLGLLHELIISKC 656
Score = 37.7 bits (86), Expect = 0.025
Identities = 29/112 (25%), Positives = 52/112 (45%), Gaps = 2/112 (1%)
Frame = +3
Query: 956 HSLKELDIVGCDKLKLSSDFQYLTCLETLAIGS-CSEVEGFHEALQHMTTLKSLTLSDLP 1014
H L L I C KL S YL L + + C++ G ++ +L+++ +
Sbjct: 60 HXLSTLQITKCPKL---SGLPYLPSLNDMRVREKCNQ--GLLSSIHKHQSLETIRFAHNE 224
Query: 1015 NLEYLPE-CIGNLTLLHEINIYSCPKLACLPTSIQQISGLEILSIHDCSKLE 1065
L Y P+ + NLT L ++I+ KL LPT ++ ++ + I + L+
Sbjct: 225 ELVYFPDRMLQNLTSLKVLDIFELSKLEKLPTEFVSLNSIQEIYISGSNSLK 380
Score = 30.8 bits (68), Expect = 3.0
Identities = 20/58 (34%), Positives = 31/58 (52%)
Frame = +3
Query: 601 LCKLCNLEVLKLDGCVSLQKLPGGLTRLKRLQNLSLRDCDSLTSLPRQIGKLTSLNTL 658
L L +L+VL + L+KLP L +Q + + +SL SLP ++ L LN+L
Sbjct: 252 LQNLTSLKVLDIFELSKLEKLPTEFVSLNSIQEIYISGSNSLKSLPDEV--LQGLNSL 419
Score = 24.3 bits (51), Expect(2) = 4e-58
Identities = 14/20 (70%), Positives = 15/20 (75%), Gaps = 1/20 (5%)
Frame = +2
Query: 820 LEKLPNLIGLSREERV-MFP 838
LEKLPNL LS E+R MFP
Sbjct: 2 LEKLPNLKRLSWEDRENMFP 61
>TC87193 similar to GP|21616922|gb|AAM66423.1 NBS-LRR protein {Oryza sativa
(japonica cultivar-group)}, partial (26%)
Length = 3119
Score = 223 bits (567), Expect = 4e-58
Identities = 169/508 (33%), Positives = 249/508 (48%), Gaps = 24/508 (4%)
Frame = +3
Query: 9 VLGSLSELIRKEISLFLGFDQEFNRLASLLTTIKATLEDAEEKQFSDSEIGRDVKDWLLK 68
++ L+ +EI G E RL + +IKA L DAE+KQ E V+ W+ +
Sbjct: 114 LVNRLASAAFREIGRIYGVMDELERLKKTVESIKAVLLDAEDKQ----EQSHAVQLWVRR 281
Query: 69 LKDAAYTLDDIMDECATEALEMEYKASKC----------GLSHKHIAFRYKLAKKMKRIG 118
L D DD++DE E +M +K K S AFR K+A ++++I
Sbjct: 282 LNDLLLPADDLIDEFFIE--DMIHKRDKAHKNKVKQVFHSFSPSRTAFRRKMAHEIEKIQ 455
Query: 119 VWLDDIAAEKNKFHLTEIVRERSGVVPDWRQTTSIVTQPLVYGRNEDKDKIVDFLVGDAS 178
D+ + + L V + R+T S V + + GR +DK+KI+ L+ +
Sbjct: 456 RSFKDVEKDMSYLKLNSNVVVVAKTNNVRRETCSYVLESEIIGREDDKNKIIS-LLRQSH 632
Query: 179 EQEDLSVYPIVGLGGLGKTTLAQLVFNHDKIVNHFELKIWVCVSEDFTLKRMTKAIIEGA 238
E +++S+ IVG+GGLGKT LAQLV+ ++ N FE +WVCVS++F K + K ++
Sbjct: 633 EHQNVSLVAIVGIGGLGKTALAQLVYKDGEVKNLFEKHMWVCVSDNFDFKTILKNMVASL 812
Query: 239 TKKSCEDLDLELLQRKLQDLLRRKRYLLVLDDVWNDKQENWQRLKSVLACGGKGASILVT 298
TK+ + L+ LQ LQ L KRYLLVLDDVWN+ E W +L+ L CG +G+ +++T
Sbjct: 813 TKEDVVNKTLQELQSMLQVNLTGKRYLLVLDDVWNESFEKWDQLRPYLMCGAQGSKVVMT 992
Query: 299 TRLPKVAKIMGTIPHHELSRLSDEDCWELFKQRAFGPNEV-QQKELVIVGKEIIKKCGGF 357
T VA MG H L L+ E W LFK FG V + L +GK+I +KC G
Sbjct: 993 TCSKIVADRMGVSDQHVLRGLTPEKSWVLFKNIVFGDVTVGVNQPLESIGKKIAEKCKGV 1172
Query: 358 PLAAIALGSLLRFKREEKEWLYVKESKLWNLQGEAYVMPALRLSYLHLPVKL---RQCFS 414
PLA +LG +LR + +E EW+ V LQGE +R + + K+ S
Sbjct: 1173PLAIRSLGGILRSESKESEWINV-------LQGECLENCVIRENSIIAGPKIELPELITS 1331
Query: 415 FCALFPKDEIISKQLLIDLWTANGFISSNQMLEADDIGNEVWNELYWRSFFENTENV--- 471
A+F IS L I N + ++ + +E W S +N
Sbjct: 1332TKAMFCLLLFISSGLGI*EG*VNSNVDGTRLSWL--FS*KAMHERCW*SICKNFLEEFIL 1505
Query: 472 -------GFGQITIFKMHDLVHDLAGSV 492
+ +T FKMHDL+HDLA V
Sbjct: 1506PRCKFE*SWVMVTGFKMHDLMHDLATQV 1589
Score = 43.9 bits (102), Expect = 3e-04
Identities = 46/184 (25%), Positives = 77/184 (41%), Gaps = 2/184 (1%)
Frame = +1
Query: 904 NLASPLKTLGFHRHSKLKMLPTEMIHIHALQQLYINDCRNIEELPNEVMQRLHSLKELDI 963
N+A ++GF S LK HI I +E +P + ++ L SL+ LD+
Sbjct: 2506 NIAESQYSIGFPPLSILKSF-----HI-------IETIMGMENVPKDWLKNLTSLENLDL 2649
Query: 964 VGCDKLKLSSDFQYLTCLETLAIGSCSEVEGFHEALQHMTTLKSLTLSDLPNLEYLPECI 1023
D L S F+ + F + L + +L+ + + L+ LP+ I
Sbjct: 2650 ---DSLS-SQQFEVIEMW-------------FKDDLICLPSLQKIQIRTC-RLKALPDWI 2775
Query: 1024 GNLTLLHEINIYSCPKLACLPTSIQQISGLEILSIHDC--SKLEKRCQKEIGEDWPKIVH 1081
N++ L + +Y C L LP + +++ L L I C S ++ G W KI H
Sbjct: 2776 CNISSLQHLAVYGCESLVDLPEGMPRLTNLHTLEIIGCPISYYYDEFLRKTGATWSKIAH 2955
Query: 1082 VQYI 1085
+ I
Sbjct: 2956 IPKI 2967
>AW125982
Length = 527
Score = 201 bits (512), Expect = 1e-51
Identities = 111/173 (64%), Positives = 128/173 (73%), Gaps = 4/173 (2%)
Frame = +3
Query: 716 RNEVSQLQENVEQILEALQPYAQKLYSFGVGGYTGAYFPQWISIPSLNDLKSLELVDCKS 775
RNE SQLQENVE+ILE LQP++Q+L S GVGGYTGAYFPQW+S PSL L LELVDC +
Sbjct: 9 RNEDSQLQENVEKILEELQPHSQQLQSLGVGGYTGAYFPQWMSSPSLKYLTQLELVDCNN 188
Query: 776 CLNLPELWKLPSLKYLKLSNMIHVIYLFHESYDG---EGLMALKTLFLEKLPNLIGLSRE 832
CL+LP L KL SL L + NM H+ YL+ ESY G G +K L LEKLP+L+ LSRE
Sbjct: 189 CLHLPLLGKLSSLNSLTVCNMSHLKYLYEESYIGGVAGGYTTVKILILEKLPDLVRLSRE 368
Query: 833 ER-VMFPRLKALEITECPNLLGLPCLPSLSDLYIQGKYNQQLPSSIHKLGSLE 884
+R +FP L L+ITECP LLGLP LPSLSDL + GK NQ L SSIHK SLE
Sbjct: 369 DRDNIFPCLSTLQITECPILLGLPSLPSLSDLRVIGKCNQHLLSSIHKQHSLE 527
>TC91142 weakly similar to PIR|T02213|T02213 NBS-LRR type resistance protein
- rice (fragment), partial (6%)
Length = 944
Score = 198 bits (504), Expect = 8e-51
Identities = 122/323 (37%), Positives = 194/323 (59%), Gaps = 15/323 (4%)
Frame = +1
Query: 430 LIDLWTANGFISSNQMLEADDIGNEVWNELYWRSFFENTENVGFGQITIFKMHDLVHDLA 489
LI LW ANG ++S L+ + +G++VWN+L+ RSFF+ ++ G IT F+MHD +HDLA
Sbjct: 7 LIHLWMANGRVTSTGNLQMEHVGDDVWNQLWQRSFFQEVKSDLAGNIT-FRMHDFIHDLA 183
Query: 490 GSVTQDVCCITDDNSMRTMSEETRHLLIYNRNS-----FAEA---NSIQLHHVKSLKTYM 541
S+ + C D + +S HL I+++ F ++ + I V L+T++
Sbjct: 184 QSIMEKECISYDVSHSTNVSIGVHHLSIFDKKPNICFFFLKSKYDHIIPFQKVDFLRTFL 363
Query: 542 EF-----NFDVYEAGQLSPQVLNCYSLRVLLSHRLNNLSSSIGRLKYLRYLDISEGRFKN 596
E+ N DV+ L+ SLRVLL+ +N S + + +LRYL+I +
Sbjct: 364 EYKPPSKNLDVF---------LSSTSLRVLLTR--SNELSLLKSIVHLRYLEIYDSNITT 510
Query: 597 LPNSLCKLCNLEVLKLDGCVSLQKLPGGLTRLKRLQNLSLRDCDSLTSLPRQIGKLTSLN 656
LP S+C+L L+ LKL+ C L P T+LK L++L +++C SL S P +IG+LTSL
Sbjct: 511 LPGSVCRLQKLQTLKLERCHLLSSFPKQFTKLKDLRHLMIKNCHSLISAPFRIGQLTSLK 690
Query: 657 TLSKYIVGEERGFLLEELGQLNLKGQLHIKNLERLKSVTDAKKANM-SRKKLNQLWLSWE 715
TL+ +IVG + G+ L +L L L G+LHIK LE + + DA++ N+ S+K L++L+LSW
Sbjct: 691 TLTIFIVGSKTGYGLAQLHNLQLGGKLHIKCLENVSNEEDARETNLISKKDLDRLYLSWG 870
Query: 716 RNEVSQL-QENVEQILEALQPYA 737
+ SQ+ + E++LEAL+P++
Sbjct: 871 NDTNSQVGSVDAERVLEALEPHS 939
>BG648625 weakly similar to GP|20385440|gb| resistance gene analog {Vitis
vinifera}, partial (21%)
Length = 521
Score = 194 bits (493), Expect = 2e-49
Identities = 95/174 (54%), Positives = 124/174 (70%)
Frame = +3
Query: 314 HELSRLSDEDCWELFKQRAFGPNEVQQKELVIVGKEIIKKCGGFPLAAIALGSLLRFKRE 373
H L++LSD+D LFKQ AFG N + ELV +G+++++KC G PLAA LGSLLRFK +
Sbjct: 3 HPLAQLSDDDI*SLFKQHAFGANREGRAELVEIGQKLVRKCVGSPLAAKVLGSLLRFKSD 182
Query: 374 EKEWLYVKESKLWNLQGEAYVMPALRLSYLHLPVKLRQCFSFCALFPKDEIISKQLLIDL 433
E +W V ES+ WNL + +VM ALRLSY +L + LR CF+FCA+FPKD + K+ I L
Sbjct: 183 EHQWTSVVESEFWNLADDNHVMSALRLSYFNLKLSLRPCFTFCAVFPKDFEMEKEFFIQL 362
Query: 434 WTANGFISSNQMLEADDIGNEVWNELYWRSFFENTENVGFGQITIFKMHDLVHD 487
W ANG ++S L+ + +GNEVWNELY RSFF+ ++ G IT FKMHDLVHD
Sbjct: 363 WMANGLVTSRGNLQMEHVGNEVWNELYQRSFFQEIKSDLVGNIT-FKMHDLVHD 521
>CB892566 weakly similar to GP|15289776|db Lycopersicon esculentum resistance
complex protein I2C-2 like protein, partial (1%)
Length = 688
Score = 184 bits (468), Expect = 1e-46
Identities = 102/235 (43%), Positives = 152/235 (64%), Gaps = 3/235 (1%)
Frame = +2
Query: 5 VLEIVLGSLSELIRKEISLFLGFDQEFNRLASLLTTIKATLEDAEEKQFSDSEIGRDVKD 64
+L I++ +L +++E++ +LG + L+ LT I+A L+DAE+KQ ++ VK+
Sbjct: 2 LLGILIQNLGSFVQEELATYLGVGELTQSLSRKLTLIRAVLKDAEKKQITNDA----VKE 169
Query: 65 WLLKLKDAAYTLDDIMDECATEALEMEYKASKCGLSHKH---IAFRYKLAKKMKRIGVWL 121
WL +L+DAAY LDDI+DEC+ + ++ + ++ H I R + K+MK I +
Sbjct: 170 WLQQLRDAAYVLDDILDECS---ITLKAHGNNKRITRFHPMKILVRRNIGKRMKEIAKEI 340
Query: 122 DDIAAEKNKFHLTEIVRERSGVVPDWRQTTSIVTQPLVYGRNEDKDKIVDFLVGDASEQE 181
DDIA E+ KF L V ER RQTTS++T+ VYGR++DK+ IV+FL+ A + E
Sbjct: 341 DDIAEERMKFGLHVGVIERQPEDEGRRQTTSVITESKVYGRDKDKEHIVEFLLRHAGDSE 520
Query: 182 DLSVYPIVGLGGLGKTTLAQLVFNHDKIVNHFELKIWVCVSEDFTLKRMTKAIIE 236
+LSVY IVG GG GKTTLAQ VFN +++ HF+LKIWVCVS D ++ ++IIE
Sbjct: 521 ELSVYSIVGHGGYGKTTLAQTVFNDERVKTHFDLKIWVCVSGDINAMKVLESIIE 685
>TC93402 weakly similar to GP|16588610|gb|AAL26857.1 NBS-type putative
resistance protein {Glycine max}, partial (61%)
Length = 676
Score = 178 bits (452), Expect = 9e-45
Identities = 103/226 (45%), Positives = 142/226 (62%), Gaps = 3/226 (1%)
Frame = +2
Query: 183 LSVYPIVGLGGLGKTTLAQLVFNHDKIVNHFELKIWVCVSEDFTLKRMTKAIIEG-ATKK 241
+S IVGLGG+GKTTLAQLV+N +I FELK WV VSE F + +T AI+ T +
Sbjct: 5 VSTISIVGLGGMGKTTLAQLVYNDRRIQETFELKAWVYVSEYFDVIGLTNAILRKFGTAE 184
Query: 242 SCEDLDLELLQRKLQDLLRRKRYLLVLDDVWNDKQENWQRLKSVLACGGKGASILVTTRL 301
+ EDLDL LQ +LQ L K YLLV+DDVW +E+W++L G G+ I++TTR
Sbjct: 185 NSEDLDL--LQWQLQKKLTGKNYLLVMDDVWKLNEESWEKLLLPFNYGSSGSKIILTTRD 358
Query: 302 PKVAKIMGTIPHHELSRLSDEDCWELFKQRAF-GPNEVQQKELVIVGKEIIKKCGGFPLA 360
KVA I+ + +L +L ++DCW LFK+ AF G N + +L +GK I+ KCGG PL
Sbjct: 359 KKVALIVKSTELVDLEQLKNKDCWSLFKRLAFHGSNVSEYPKLESIGKNIVDKCGGLPLT 538
Query: 361 AIALGSLLRFKREEKEWLYVKESKLWNL-QGEAYVMPALRLSYLHL 405
+G+LLR K + EW + E+ +W L ++ + ALRLSY +L
Sbjct: 539 GKTMGNLLRKKFTQSEWEKILEADMWRLTDDDSNINSALRLSYHNL 676
>AW691229
Length = 487
Score = 175 bits (444), Expect = 8e-44
Identities = 97/152 (63%), Positives = 116/152 (75%), Gaps = 12/152 (7%)
Frame = +1
Query: 1 MAEAVLEIVLGSLSELIRKEISLFLGFDQEFNRLASLLTTIKATLEDAEEKQFSDSEIGR 60
MAEAV+E+VL +LS LI+KE+ LFLG D+E L+SLLTTIKATLEDAEEKQFS+ R
Sbjct: 43 MAEAVIEVVLDNLSTLIQKELGLFLGVDRELKSLSSLLTTIKATLEDAEEKQFSN----R 210
Query: 61 DVKDWLLKLKDAAYTLDDIMDECATEALEMEYKASKCG------------LSHKHIAFRY 108
+KDWLLKLKDAA+ LDDI+DECAT+ALE EYK K G L+ K++AF Y
Sbjct: 211 AIKDWLLKLKDAAHVLDDILDECATKALEPEYKGFKYGPSQKVQSSCLSSLNPKNVAFLY 390
Query: 109 KLAKKMKRIGVWLDDIAAEKNKFHLTEIVRER 140
K+AKK+KRI LD IA E++KFHLTEIVRER
Sbjct: 391 KIAKKIKRIRERLDGIAEERSKFHLTEIVRER 486
>TC81627 weakly similar to GP|18652501|gb|AAL77135.1 Putative NBS-LRR type
resistance protein {Oryza sativa}, partial (6%)
Length = 890
Score = 173 bits (438), Expect = 4e-43
Identities = 97/243 (39%), Positives = 149/243 (60%), Gaps = 1/243 (0%)
Frame = +2
Query: 1 MAEAVLEIVLGSLSELIRKEISLFLGFDQEFNRLASLLTTIKATLEDAEEKQFSDSEIGR 60
M + +L IV+ +L +R+E+S FLG + +L LT I+A L+DAEEKQ + R
Sbjct: 146 MTDVLLGIVIENLISFVREELSTFLGVGELTQKLCGNLTAIRAVLQDAEEKQITS----R 313
Query: 61 DVKDWLLKLKDAAYTLDDIMDECATEALEMEYKASKCGLSHKHIAFRYKLAKKMKRIGVW 120
VKDWL KL DAA+ LDDI+D+C+T + K I + + K+MK +
Sbjct: 314 AVKDWLQKLADAAHVLDDILDDCSTTSKAHGDNKWIARFHPKKILAQRAIGKRMKEVAKK 493
Query: 121 LDDIAAEKNKFHL-TEIVRERSGVVPDWRQTTSIVTQPLVYGRNEDKDKIVDFLVGDASE 179
+D+IA + K+ L ++ E +WRQTTS++T+P VYGR+ D +KIV+ LV A +
Sbjct: 494 IDEIAEGRIKYGLQVKVTEEHQRGDDEWRQTTSVITEPKVYGRDHDIEKIVEVLVSHAID 673
Query: 180 QEDLSVYPIVGLGGLGKTTLAQLVFNHDKIVNHFELKIWVCVSEDFTLKRMTKAIIEGAT 239
E+LSVY IVG+GGLGKTTLAQ+VFN +++ +E + +S ++ ++ ++II+
Sbjct: 674 SEELSVYSIVGVGGLGKTTLAQVVFNDERLDTLYERLTNMKLSSSSSMMKILESIIKSTV 853
Query: 240 KKS 242
K+
Sbjct: 854 GKN 862
>BG648971 similar to GP|20514808|gb| Putative NBS-LRR type resistance protein
{Oryza sativa (japonica cultivar-group)}, partial (2%)
Length = 830
Score = 173 bits (438), Expect = 4e-43
Identities = 99/269 (36%), Positives = 154/269 (56%), Gaps = 1/269 (0%)
Frame = +1
Query: 439 FISSNQMLEADDIGNEVWNELYWRSFFENTENVGFGQITIFKMHDLVHDLAGSVTQDVCC 498
F+ L+ + +GNEVWNELY RSFF+ ++ G IT FKMHDLVHDLA S+ + C
Sbjct: 4 FLHLEGNLQMELLGNEVWNELYQRSFFQEVKSDIVGNIT-FKMHDLVHDLAQSIMGEECV 180
Query: 499 ITDDNSMRTMSEETRHLLIYNRNSFAEANSIQLHHVKSLKTYMEFNFDVYEAGQLSPQVL 558
++ +S+ +S H+ + + I + ++SL+T++EF + L P
Sbjct: 181 ASEVSSLADLSIRVHHISFIDSKEKLDYKMIPFNKIESLRTFLEFRPSTKKLDVLPP--- 351
Query: 559 NCYSLRVLLSHRLNNLS-SSIGRLKYLRYLDISEGRFKNLPNSLCKLCNLEVLKLDGCVS 617
+ +L + R ++ S++ L +LRYL++ R LP S+C+L L+ LKL C
Sbjct: 352 ----INLLRALRTSSFGLSALRNLMHLRYLELCHSRITTLPGSVCRLQKLQTLKLKDCPY 519
Query: 618 LQKLPGGLTRLKRLQNLSLRDCDSLTSLPRQIGKLTSLNTLSKYIVGEERGFLLEELGQL 677
P LT+L+ L+++ + +C SL S P +IG+LT L TL+ +IVG + GF L EL L
Sbjct: 520 FSHFPKQLTQLQELRHIVIENCFSLVSTPFRIGELTCLKTLTVFIVGSKTGFGLAELHNL 699
Query: 678 NLKGQLHIKNLERLKSVTDAKKANMSRKK 706
L G LHI+ LE + + DA++AN+ K
Sbjct: 700 QLGGMLHIRGLENVSNDGDAREANLIGNK 786
Score = 41.6 bits (96), Expect = 0.002
Identities = 28/87 (32%), Positives = 42/87 (48%), Gaps = 2/87 (2%)
Frame = +1
Query: 917 HSKLKMLPTEMIHIHALQQLYINDCRNIEELPNEVMQRLHSLKELDIVGCDKLKLSSDFQ 976
HS++ LP + + LQ L + DC P ++ Q L L+ + I C L +S+ F+
Sbjct: 439 HSRITTLPGSVCRLQKLQTLKLKDCPYFSHFPKQLTQ-LQELRHIVIENCFSL-VSTPFR 612
Query: 977 Y--LTCLETLAIGSCSEVEGFHEALQH 1001
LTCL+TL + GF A H
Sbjct: 613 IGELTCLKTLTVFIVGSKTGFGLAELH 693
>CA920463 weakly similar to GP|2852684|gb|A resistance protein candidate
{Lactuca sativa}, partial (6%)
Length = 810
Score = 173 bits (438), Expect = 4e-43
Identities = 98/237 (41%), Positives = 150/237 (62%), Gaps = 3/237 (1%)
Frame = -1
Query: 262 KRYLLVLDDVWNDKQENWQRLKSVLACGGKGASILVTTRLPKVAKIMGTIPHHELSRLSD 321
K++LL LDDVW++ + W +K++L G +G+ +LVTTR +AK+M T + L LS
Sbjct: 756 KKFLLRLDDVWSEDRVKWIEVKNLLQVGDEGSKVLVTTRSHSIAKMMCTNTSYTLQGLSR 577
Query: 322 EDCWELFKQRAFGPNEVQQ-KELVIVGKEIIKKCGGFPLAAIALGSLLRFKREEKEWLYV 380
ED +F + AF E ++ +L+ +GKEI++KCGG PLA LGS L K + +EW +V
Sbjct: 576 EDSLSVFVKWAFKEGEEKKYPKLIEIGKEIVQKCGGLPLALRTLGSSLFLKDDIEEWKFV 397
Query: 381 KESKLWNL-QGEAYVMPALRLSYLHLPVKLRQCFSFCALFPKDEIISKQLLIDLWTANGF 439
+++++WNL Q E ++PAL+LS+ LP L++CF+ +LF KD S + LW A F
Sbjct: 396 RDNEIWNLPQKEDDILPALKLSFDQLPSYLKRCFACFSLFVKDFHFSNYSVTVLWEALDF 217
Query: 440 I-SSNQMLEADDIGNEVWNELYWRSFFENTENVGFGQITIFKMHDLVHDLAGSVTQD 495
+ S N+ +D+GN+ +EL RSF + + G + +FK+HDLVHDLA V +D
Sbjct: 216 LPSPNKGKTLEDVGNQFLHELQSRSFLQ--DFYVSGNVCVFKLHDLVHDLALYVARD 52
>AW695285 similar to GP|15487944|gb| NBS/LRR resistance protein-like protein
{Theobroma cacao}, partial (9%)
Length = 618
Score = 161 bits (408), Expect = 1e-39
Identities = 93/205 (45%), Positives = 126/205 (61%), Gaps = 3/205 (1%)
Frame = +2
Query: 1 MAEAVLEIVLGSLSELIRKEISLFLGFDQEFNRLASLLTTIKATLEDAEEKQFSDSEIGR 60
MAEA+L ++ L++ E S G + L++ L I+A L DAE++Q DS I
Sbjct: 23 MAEALLRAAFEKVNSLLQSEFSTISGIKSKAKNLSTSLNHIEAVLVDAEKRQVKDSYI-- 196
Query: 61 DVKDWLLKLKDAAYTLDDIMDECATEALEMEYKASKCGLSHKHIAFRYKLAKKMKRIGVW 120
K WL +LKDA Y LDDI+DEC+ E+ + + K+I FR ++ ++K I
Sbjct: 197 --KVWLQQLKDAVYVLDDILDECSIESARL---GGSFSFNPKNIVFRRQIGNRLKEITRR 361
Query: 121 LDDIAAEKNKFHL---TEIVRERSGVVPDWRQTTSIVTQPLVYGRNEDKDKIVDFLVGDA 177
LDDIA KNKF L T VRE S V +WRQ SI+ +P V+GR +DK+KI +FL+ A
Sbjct: 362 LDDIADIKNKFLLRDGTVYVRESSDEVDEWRQINSIIAKPEVFGRKDDKEKIFEFLLTHA 541
Query: 178 SEQEDLSVYPIVGLGGLGKTTLAQL 202
+ + LSVYPIVGLGG+GKTTL QL
Sbjct: 542 RDSDFLSVYPIVGLGGIGKTTLVQL 616
>TC81427 weakly similar to GP|11994216|dbj|BAB01338. disease resistance
comples protein {Arabidopsis thaliana}, partial (6%)
Length = 1253
Score = 152 bits (384), Expect = 7e-37
Identities = 129/426 (30%), Positives = 209/426 (48%), Gaps = 18/426 (4%)
Frame = +3
Query: 538 KTYMEFNFD--VYEAGQLSPQVLNCYS----LRV--LLSHRLNNLSSSIGRLKYLRYLDI 589
+T+M F F V G +S V LRV L + + L SSIG L +LRYLD+
Sbjct: 3 RTFMPFKFYPVVPSLGGISASVSTLLKKPKPLRVFSLSEYPITLLPSSIGHLLHLRYLDL 182
Query: 590 SEGRFKNLPNSLCKLCNLEVLKLDGCVSLQKLPGGLTRLKRLQNLSLRDCDSLTSLPRQI 649
S +LP+S+C L NL L L GC L LP ++L L+ L + + +P +
Sbjct: 183 SRTPITSLPDSICNLYNLXALLLVGCADLTLLPTKTSKLINLRQLDISG-SGIKKMPTNL 359
Query: 650 GKLTSLNTLSKYIVGEERGFLLEELGQ-LNLKGQLHIKNLERLKSVTDAKKANMSRKK-L 707
GKL SL +L +++V + G + ELG+ L L+G L I NLE + A A + RKK L
Sbjct: 360 GKLKSLQSLPRFVVSNDGGSNVGELGEMLELRGSLSIVNLENVLLKEXASNAGLKRKKYL 539
Query: 708 NQLWLSWERNEVSQLQENVEQILEALQPYAQKLYSFGVGGYTGAYFPQWISIPSLNDLKS 767
+++ W SQ EN+ I + L P+ + L + + G FP W+ S + + S
Sbjct: 540 HEVEFKWTTPTHSQESENI--IFDMLXPH-RNLKRLKINNFGGEKFPNWLGSNSGSTMMS 710
Query: 768 LELVDCKSCLNLPELWKLPSLKYLKLSNMIHVIYLFHESYDGEGLMALKTLFLEKLPNLI 827
L L +C +CL+LP L +L +L+ + ++++ + + E Y G G A +L + K +++
Sbjct: 711 LYLDECGNCLSLPSLGQLSNLREIYITSVTRLQKVGPEFY-GNGFEAFSSLRIIKFKDML 887
Query: 828 GL------SREERVMFPRLKALEITECPNLLG-LP-CLPSLSDLYIQGKYNQQLPSSIHK 879
++ F L+ L I CP L+G LP LPSL L I Q L ++
Sbjct: 888 NWEEWSVNNQSGSEGFTLLQELYIENCPKLIGKLPGNLPSLDKLVITS--CQTLSDTMPC 1061
Query: 880 LGSLESLHFSDNEELIYFPDGILRNLASPLKTLGFHRHSKLKMLPTEMIHIHALQQLYIN 939
+ L L S E + + +++ L+T+ L +P + + L+ L ++
Sbjct: 1062VPRLRELKISGCEAFVSLSEQMMK-CNDCLQTMAISNCPSLVSIPMDCVS-RTLKSLKVS 1235
Query: 940 DCRNIE 945
DC+ ++
Sbjct: 1236DCQKLQ 1253
Score = 31.2 bits (69), Expect = 2.3
Identities = 20/61 (32%), Positives = 30/61 (48%)
Frame = +3
Query: 998 ALQHMTTLKSLTLSDLPNLEYLPECIGNLTLLHEINIYSCPKLACLPTSIQQISGLEILS 1057
++ H+ L+ L LS P + LP+ I NL L + + C L LPT ++ L L
Sbjct: 144 SIGHLLHLRYLDLSRTP-ITSLPDSICNLYNLXALLLVGCADLTLLPTKTSKLINLRQLD 320
Query: 1058 I 1058
I
Sbjct: 321 I 323
Score = 30.8 bits (68), Expect = 3.0
Identities = 33/138 (23%), Positives = 59/138 (41%), Gaps = 8/138 (5%)
Frame = +3
Query: 933 LQQLYINDCRNIEELPNEVMQRLHSLKELDIVGCDKL-KLSSDFQYLTCLETLAIGSCSE 991
+ LY+++C N LP+ + +L +L+E+ I +L K+ +F
Sbjct: 702 MMSLYLDECGNCLSLPS--LGQLSNLREIYITSVTRLQKVGPEF---------------- 827
Query: 992 VEGFHEALQHMTTLKSLTLSDLPNLEYLPECIGN------LTLLHEINIYSCPKL-ACLP 1044
+ + ++L+ + D+ N E + N TLL E+ I +CPKL LP
Sbjct: 828 ---YGNGFEAFSSLRIIKFKDMLNWEEWS--VNNQSGSEGFTLLQELYIENCPKLIGKLP 992
Query: 1045 TSIQQISGLEILSIHDCS 1062
++ + L I S S
Sbjct: 993 GNLPSLDKLVITSCQTLS 1046
>BF643456 weakly similar to GP|14348622|gb NBS-LRR resistance-like protein
J78 {Phaseolus vulgaris}, partial (6%)
Length = 521
Score = 149 bits (376), Expect = 6e-36
Identities = 75/173 (43%), Positives = 113/173 (64%), Gaps = 2/173 (1%)
Frame = +2
Query: 206 HDKIVNHFELKIWVCVSEDFTLKRMTKAIIEGATKKSCEDLDLELLQRKLQDLLRRKRYL 265
+D I +F+++ W CVS+ F ++TKAI+E T+ +C ++ELL L++ L K++L
Sbjct: 2 NDNIKQNFDVQAWACVSDHFDEFKVTKAIMEAVTRSACNINNIELLHLDLKEKLAGKKFL 181
Query: 266 LVLDDVWNDKQENWQRLKSVLACGGKGASILVTTRLPKVAKIMGTIPHHELSRLSDEDCW 325
+VLDDVW + + W+ L L KG+ ILVTTR+ KVA ++ T + L +LSDEDCW
Sbjct: 182 IVLDDVWTEDYDAWKSLLRPLQYSDKGSKILVTTRIKKVASMLQTFQGYSLEQLSDEDCW 361
Query: 326 ELFKQRA--FGPNEVQQKELVIVGKEIIKKCGGFPLAAIALGSLLRFKREEKE 376
+F+ A + +++ EL +GKEII+KC G PLAA +LG LLR KR+ K+
Sbjct: 362 SVFENHACLSLKHSIEKMELQKIGKEIIRKCQGLPLAAQSLGGLLRSKRDIKD 520
>BG646099 weakly similar to GP|13957628|gb putative resistance protein {Oryza
sativa}, partial (4%)
Length = 681
Score = 139 bits (351), Expect = 5e-33
Identities = 74/171 (43%), Positives = 101/171 (58%), Gaps = 2/171 (1%)
Frame = -3
Query: 338 VQQKELVIVGKEIIKKCGGFPLAAIALGSLLRFKREEKEWLYVKESKLWNL-QGEAYVMP 396
V + L +GK+I +KC G PLA LG LL+ K EE EW+ V + W L + E +MP
Sbjct: 673 VLNQNLEAIGKKIAEKCSGVPLAIRTLGGLLQGKSEETEWVGVLQDDFWKLCEDEETIMP 494
Query: 397 ALRLSYLHLPVKLRQCFSFCALFPKDEIISKQLLIDLWTANGFIS-SNQMLEADDIGNEV 455
L+LSY +L +LRQCFS+C+L+PKD I K LI LW A G++ S Q +DIGN+
Sbjct: 493 VLKLSYQNLSPQLRQCFSYCSLYPKDWEIKKDELIQLWMAQGYLECSTQKQGPEDIGNQF 314
Query: 456 WNELYWRSFFENTENVGFGQITIFKMHDLVHDLAGSVTQDVCCITDDNSMR 506
+SFF++ + G I FKMHDL+HDLA + + CC + R
Sbjct: 313 VKIFLMKSFFQDAKIDDCGDICSFKMHDLMHDLAMQIAGNDCCYIGSETKR 161
>BI265031 weakly similar to GP|13872974|db putative NBS-LRR type resistance
protein {Oryza sativa (japonica cultivar-group)},
partial (3%)
Length = 662
Score = 124 bits (311), Expect(2) = 1e-31
Identities = 66/163 (40%), Positives = 99/163 (60%)
Frame = +1
Query: 171 DFLVGDASEQEDLSVYPIVGLGGLGKTTLAQLVFNHDKIVNHFELKIWVCVSEDFTLKRM 230
+FL+ D+ + + + IVG+ G+GKTT+A+LV+N KI FELK WV VSE F L +
Sbjct: 91 EFLLSDSYSETFVPIISIVGVIGMGKTTIARLVYNDHKIHEQFELKAWVYVSESFDLVHL 270
Query: 231 TKAIIEGATKKSCEDLDLELLQRKLQDLLRRKRYLLVLDDVWNDKQENWQRLKSVLACGG 290
T+AI+ D+E+LQR+LQ L K+YLLVLD++WN+ E ++L + G
Sbjct: 271 TQAILREFHSSETYSEDMEILQRQLQQRLAGKKYLLVLDNIWNENVECRKKLLLPFSNGS 450
Query: 291 KGASILVTTRLPKVAKIMGTIPHHELSRLSDEDCWELFKQRAF 333
G+ ++V T +VA IM + L++L++ D W LF AF
Sbjct: 451 SGSKLIVRTPHNEVASIMASTRLLRLNQLNESDSWSLFVHHAF 579
Score = 32.0 bits (71), Expect(2) = 1e-31
Identities = 13/27 (48%), Positives = 17/27 (62%)
Frame = +3
Query: 333 FGPNEVQQKELVIVGKEIIKKCGGFPL 359
FG N + L +GK+I+ KCGG PL
Sbjct: 579 FGQNIFEYPNLESIGKKIVXKCGGLPL 659
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.320 0.138 0.409
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 36,826,226
Number of Sequences: 36976
Number of extensions: 551778
Number of successful extensions: 3665
Number of sequences better than 10.0: 323
Number of HSP's better than 10.0 without gapping: 3171
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 3465
length of query: 1113
length of database: 9,014,727
effective HSP length: 106
effective length of query: 1007
effective length of database: 5,095,271
effective search space: 5130937897
effective search space used: 5130937897
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 64 (29.3 bits)
Medicago: description of AC134049.3


