

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
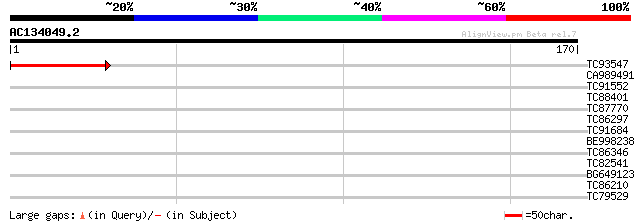
Query= AC134049.2 - phase: 0
(170 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC93547 weakly similar to GP|13605637|gb|AAK32812.1 At1g27150/T7... 51 2e-07
CA989491 similar to PIR|T10971|T109 Pv42 protein - kidney bean, ... 28 2.3
TC91552 28 2.3
TC88401 homologue to GP|20907370|gb|AAM32440.1 conserved protein... 28 2.3
TC87770 similar to GP|14970762|emb|CAC44460. protein tyrosine ph... 27 3.0
TC86297 similar to GP|15912283|gb|AAL08275.1 At1g30270/F12P21_6 ... 27 5.2
TC91684 weakly similar to PIR|A96574|A96574 protein F12M16.30 [i... 26 6.8
BE998238 similar to PIR|T39202|T39 heat shock protein 90 homolog... 26 6.8
TC86346 homologue to GP|22136752|gb|AAM91695.1 unknown protein {... 26 6.8
TC82541 similar to GP|13937149|gb|AAK50068.1 At1g79380/T8K14_20 ... 26 8.9
BG649123 homologue to GP|22597160|gb| hypothetical protein {Glyc... 26 8.9
TC86210 homologue to GP|13194621|gb|AAK15493.1 brassinosteroid b... 26 8.9
TC79529 similar to GP|20197375|gb|AAM15048.1 expressed protein {... 26 8.9
>TC93547 weakly similar to GP|13605637|gb|AAK32812.1 At1g27150/T7N9_21
{Arabidopsis thaliana}, partial (34%)
Length = 679
Score = 51.2 bits (121), Expect = 2e-07
Identities = 23/30 (76%), Positives = 26/30 (86%)
Frame = +1
Query: 1 MGINHHVLPQNEGENFIYGMLAFPLLELGQ 30
+ + + VLPQNEG NFIYGMLAFPLLELGQ
Sbjct: 532 LSLVNQVLPQNEGXNFIYGMLAFPLLELGQ 621
>CA989491 similar to PIR|T10971|T109 Pv42 protein - kidney bean, partial
(14%)
Length = 352
Score = 27.7 bits (60), Expect = 2.3
Identities = 9/29 (31%), Positives = 17/29 (58%)
Frame = +1
Query: 59 LQYECRFREAVEFMEECSPSWNSFLSFML 87
L Y+ + E V + ++C SW + S+M+
Sbjct: 184 LCYQMQMHECVRYKQQCVLSWRKYCSYMI 270
>TC91552
Length = 1001
Score = 27.7 bits (60), Expect = 2.3
Identities = 10/24 (41%), Positives = 17/24 (70%)
Frame = -2
Query: 29 GQMKEAEEAAKRGFEINNQDGWSQ 52
G + EE+ KRG + NN++GW++
Sbjct: 184 GGWRNGEESGKRGTKGNNRNGWNR 113
>TC88401 homologue to GP|20907370|gb|AAM32440.1 conserved protein
{Methanosarcina mazei Goe1}, partial (3%)
Length = 569
Score = 27.7 bits (60), Expect = 2.3
Identities = 10/24 (41%), Positives = 17/24 (70%)
Frame = +2
Query: 29 GQMKEAEEAAKRGFEINNQDGWSQ 52
G + EE+ KRG + NN++GW++
Sbjct: 380 GGWRNGEESGKRGTKGNNRNGWNR 451
>TC87770 similar to GP|14970762|emb|CAC44460. protein tyrosine phosphatase
{Lycopersicon esculentum}, partial (82%)
Length = 1714
Score = 27.3 bits (59), Expect = 3.0
Identities = 12/41 (29%), Positives = 16/41 (38%)
Frame = -3
Query: 51 SQHATCHVLQYECRFREAVEFMEECSPSWNSFLSFMLTHNW 91
S H CHV+ RF + C +W S + H W
Sbjct: 560 STHGLCHVITVYLRFHLLLNIRATCLCTWYRTRSSLK*HGW 438
>TC86297 similar to GP|15912283|gb|AAL08275.1 At1g30270/F12P21_6
{Arabidopsis thaliana}, partial (90%)
Length = 2043
Score = 26.6 bits (57), Expect = 5.2
Identities = 15/42 (35%), Positives = 23/42 (54%)
Frame = +1
Query: 67 EAVEFMEECSPSWNSFLSFMLTHNWWHVALCYLEGNAPMQRV 108
+++ F EC S +S + +HN W+ LC G+ P QRV
Sbjct: 862 QSIGFWIECIASTSSG-RWTTSHNMWYTKLCCSRGD-PKQRV 981
>TC91684 weakly similar to PIR|A96574|A96574 protein F12M16.30 [imported] -
Arabidopsis thaliana, partial (20%)
Length = 1085
Score = 26.2 bits (56), Expect = 6.8
Identities = 15/48 (31%), Positives = 25/48 (51%)
Frame = -1
Query: 94 VALCYLEGNAPMQRVLEVYDNYIWKELDKTDATVPEVYLNAVALLLRL 141
V+L + GN P++ + E IW+ L + P+ LN+ LLR+
Sbjct: 323 VSLANVAGNNPVKLLEERSSLSIWERLPSSGGRFPDN*LNSNTKLLRV 180
>BE998238 similar to PIR|T39202|T39 heat shock protein 90 homolog - fission
yeast (Schizosaccharomyces pombe), partial (30%)
Length = 654
Score = 26.2 bits (56), Expect = 6.8
Identities = 14/32 (43%), Positives = 18/32 (55%)
Frame = -2
Query: 132 LNAVALLLRLCVRDELEFFGDRLKMLADRLAD 163
L + LLL L DEL+ GD L +L D + D
Sbjct: 134 LIVIDLLLNLLGHDELDGVGDELGVLLDDILD 39
>TC86346 homologue to GP|22136752|gb|AAM91695.1 unknown protein {Arabidopsis
thaliana}, partial (89%)
Length = 2436
Score = 26.2 bits (56), Expect = 6.8
Identities = 13/31 (41%), Positives = 21/31 (66%)
Frame = +1
Query: 31 MKEAEEAAKRGFEINNQDGWSQHATCHVLQY 61
+++AEEAAKR + Q G S + TC V+++
Sbjct: 1069 IEDAEEAAKRLMKEAYQRGSSDNITCVVVRF 1161
>TC82541 similar to GP|13937149|gb|AAK50068.1 At1g79380/T8K14_20
{Arabidopsis thaliana}, partial (32%)
Length = 820
Score = 25.8 bits (55), Expect = 8.9
Identities = 11/32 (34%), Positives = 15/32 (46%)
Frame = -1
Query: 86 MLTHNWWHVALCYLEGNAPMQRVLEVYDNYIW 117
+L H WWH +E + QR L+ IW
Sbjct: 679 LLDHVWWHHQSQSMESSCLHQRELKSCQQLIW 584
>BG649123 homologue to GP|22597160|gb| hypothetical protein {Glycine max},
partial (18%)
Length = 536
Score = 25.8 bits (55), Expect = 8.9
Identities = 12/33 (36%), Positives = 15/33 (45%)
Frame = +1
Query: 54 ATCHVLQYECRFREAVEFMEECSPSWNSFLSFM 86
A CH Y C F + ++ C S LSFM
Sbjct: 40 ANCHFFSYSCAFAY*IFYLIPCL*SLYYILSFM 138
>TC86210 homologue to GP|13194621|gb|AAK15493.1 brassinosteroid biosynthetic
protein LKB {Pisum sativum}, partial (67%)
Length = 1377
Score = 25.8 bits (55), Expect = 8.9
Identities = 13/34 (38%), Positives = 18/34 (52%)
Frame = +2
Query: 17 IYGMLAFPLLELGQMKEAEEAAKRGFEINNQDGW 50
I+G L L G+ EEA K+G +IN+ W
Sbjct: 287 IWGRLPGSLCMTGRYASKEEAKKKGNKINSVGWW 388
>TC79529 similar to GP|20197375|gb|AAM15048.1 expressed protein {Arabidopsis
thaliana}, partial (74%)
Length = 1378
Score = 25.8 bits (55), Expect = 8.9
Identities = 13/31 (41%), Positives = 16/31 (50%)
Frame = +3
Query: 120 LDKTDATVPEVYLNAVALLLRLCVRDELEFF 150
LDK +P YL+A L LC+ EL F
Sbjct: 1101 LDKKIGVIPLFYLDA*FFFLELCLHSELISF 1193
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.323 0.137 0.436
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 5,951,741
Number of Sequences: 36976
Number of extensions: 86482
Number of successful extensions: 447
Number of sequences better than 10.0: 26
Number of HSP's better than 10.0 without gapping: 447
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 447
length of query: 170
length of database: 9,014,727
effective HSP length: 89
effective length of query: 81
effective length of database: 5,723,863
effective search space: 463632903
effective search space used: 463632903
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 55 (25.8 bits)
Medicago: description of AC134049.2


