

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC133779.5 + phase: 1 /pseudo
(547 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
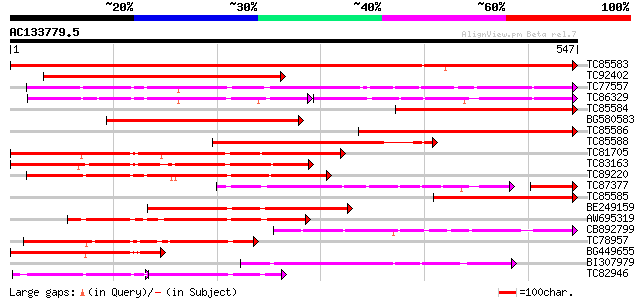
Score E
Sequences producing significant alignments: (bits) Value
TC85583 similar to PIR|T51335|T51335 subtilisin-like proteinase ... 825 0.0
TC92402 similar to GP|20198252|gb|AAM15483.1 subtilisin-like ser... 447 e-126
TC77557 similar to PIR|JC7519|JC7519 subtilisin-like serine prot... 392 e-109
TC86329 similar to PIR|T07171|T07171 subtilisin-like proteinase ... 206 3e-95
TC85584 weakly similar to PIR|A84473|A84473 probable serine prot... 342 2e-94
BG580583 weakly similar to GP|9757901|dbj serine protease-like p... 337 5e-93
TC85586 similar to GP|20521377|dbj|BAB91889. putative subtilisin... 329 2e-90
TC85588 similar to GP|20198169|gb|AAM15440.1 subtilisin-like ser... 280 1e-75
TC81705 weakly similar to GP|6721520|dbj|BAA89562.1 EST AU029428... 262 3e-70
TC83163 weakly similar to GP|6721520|dbj|BAA89562.1 EST AU029428... 252 3e-67
TC89220 similar to PIR|T05768|T05768 subtilisin-like proteinase ... 248 3e-66
TC87377 weakly similar to PIR|T07171|T07171 subtilisin-like prot... 191 1e-60
TC85585 similar to GP|22773236|gb|AAN06842.1 Putatvie subtilisin... 214 7e-56
BE249159 weakly similar to GP|20198252|gb subtilisin-like serine... 214 7e-56
AW695319 similar to GP|6708179|gb| subtilisin-type serine endope... 187 9e-48
CB892799 weakly similar to PIR|T05768|T05 subtilisin-like protei... 185 4e-47
TC78957 similar to GP|14091078|gb|AAK53589.1 subtilisin-like pro... 181 5e-46
BG449655 similar to GP|20198252|gb subtilisin-like serine protea... 178 4e-45
BI307979 weakly similar to GP|20521377|db putative subtilisin-li... 175 4e-44
TC82946 weakly similar to GP|20160863|dbj|BAB89802. putative sub... 96 9e-41
>TC85583 similar to PIR|T51335|T51335 subtilisin-like proteinase AIR3
auxin-induced [imported] - Arabidopsis thaliana
(fragment), partial (40%)
Length = 2691
Score = 825 bits (2130), Expect = 0.0
Identities = 415/563 (73%), Positives = 462/563 (81%), Gaps = 16/563 (2%)
Frame = +2
Query: 1 DLLGSILGSKENAKEAIIYSYNKQINGFAAMLEEEEAAQIAKNPKVVSVFLSKEHKLHTT 60
DLL S LGS+E AKEAIIYSYNK INGFAA+LE+EEAA IAK VVSVFLSK HKLHTT
Sbjct: 203 DLLSSTLGSREKAKEAIIYSYNKHINGFAALLEDEEAADIAKKRNVVSVFLSKPHKLHTT 382
Query: 61 RSWEFLGLRGNDINSAWQKGRFGENTIIGNIDTGVWPESKSFSDRGIGPIPAKWRGGNIC 120
RSWEFLGLR N N+AWQKG+FGENTII NIDTGVWPESKSF+D+G GP+P+KWRGG C
Sbjct: 383 RSWEFLGLRRNAKNTAWQKGKFGENTIIANIDTGVWPESKSFNDKGYGPVPSKWRGGKAC 562
Query: 121 QLDKLNTSKKVPCNRKLIGARFFNKAYQKRNGKLPRSQQTARDFVGHGTHTLSTAGGNFV 180
++ K + KK PCNRKLIGARFF+ AY+ N KLP Q+TARDF+GHGTHTLSTAGGNFV
Sbjct: 563 EISKFSKYKKNPCNRKLIGARFFSNAYEAYNDKLPSWQRTARDFLGHGTHTLSTAGGNFV 742
Query: 181 PGASIFNIGNGTIKGGSPRARVATYKVCWSLTDATSCFGADVLSAIDQAIDDGVDIISVS 240
P AS+F IGNGT+KGGSPRARVATYKVCWSL D CFGADVL+AIDQAI DGVDIIS+S
Sbjct: 743 PDASVFAIGNGTVKGGSPRARVATYKVCWSLLDLEDCFGADVLAAIDQAISDGVDIISLS 922
Query: 241 AGGPSSTNSEEIFTDEISIGAFHALARNILLVASAGNEGPTPGSVVNVAPWVFTVAASTL 300
G S E+IFTDE+SIGAFHAL+RNILLVASAGNEGPT GSVVNVAPWVFT+AASTL
Sbjct: 923 LAGHSLVYPEDIFTDEVSIGAFHALSRNILLVASAGNEGPTGGSVVNVAPWVFTIAASTL 1102
Query: 301 DRDFSSVMTIGNKTLTGASLFVNLPPNQDFTIVTSTDAKLANATNRDARFCRPRTLDPSK 360
DRDFSS +TIGN+T+ GASLFVNLPPNQ F ++ STD KLANATN DA+FC+P TLDPSK
Sbjct: 1103DRDFSSTITIGNQTIRGASLFVNLPPNQAFPLIVSTDGKLANATNHDAQFCKPGTLDPSK 1282
Query: 361 VNGKIVACDREGKIKSVAEGQEALSAGAKGVILRNQPEINGKTLLSEPHVLSTISYPGN- 419
V GKIV C REG IKSVAEGQEALSAGAKG++L NQP+ GKT L+EPH LS + P +
Sbjct: 1283VKGKIVECIREGNIKSVAEGQEALSAGAKGMLLSNQPK-QGKTTLAEPHTLSCVEVPHHA 1459
Query: 420 ---------------HSRTTGRSLDIIPSDIKSGTKLRMSPAKTLNRRKPAPVMASYSSR 464
S + + S +K+GT ++ S AKTL RKPAPVMAS+SSR
Sbjct: 1460PKPPKPKKSAEQERAGSHAPAFDITSMDSKLKAGTTIKFSGAKTLYGRKPAPVMASFSSR 1639
Query: 465 GPNKVQPSILKPDVTAPGVNILAAYSLFASASNLITDTRRGFPFNVMQGTSMSCPHVAGT 524
GPNK+QPSILKPDVTAPGVNILAAYSL+ASASNL TD R FPFNV+QGTSMSCPHVAG
Sbjct: 1640GPNKIQPSILKPDVTAPGVNILAAYSLYASASNLKTDNRNNFPFNVLQGTSMSCPHVAGI 1819
Query: 525 AGLIKTLHPNWSPAAIKSAIRTT 547
AGLIKTLHPNWSPAAIKSAI TT
Sbjct: 1820AGLIKTLHPNWSPAAIKSAIMTT 1888
>TC92402 similar to GP|20198252|gb|AAM15483.1 subtilisin-like serine
protease AIR3 {Arabidopsis thaliana}, partial (16%)
Length = 704
Score = 447 bits (1150), Expect = e-126
Identities = 215/234 (91%), Positives = 225/234 (95%)
Frame = +1
Query: 33 EEEEAAQIAKNPKVVSVFLSKEHKLHTTRSWEFLGLRGNDINSAWQKGRFGENTIIGNID 92
EEEEAAQ+AKNPKVVSVFLSKEHKLHTTRSWEFLGL GNDINSAWQKGRFGENTII NID
Sbjct: 1 EEEEAAQLAKNPKVVSVFLSKEHKLHTTRSWEFLGLHGNDINSAWQKGRFGENTIIANID 180
Query: 93 TGVWPESKSFSDRGIGPIPAKWRGGNICQLDKLNTSKKVPCNRKLIGARFFNKAYQKRNG 152
TGVWPES+SFSDRGIGPIPAKWRGGN+CQ++KL SKKVPCNRKLIGARFF+ AY++ NG
Sbjct: 181 TGVWPESRSFSDRGIGPIPAKWRGGNVCQINKLRGSKKVPCNRKLIGARFFSDAYERYNG 360
Query: 153 KLPRSQQTARDFVGHGTHTLSTAGGNFVPGASIFNIGNGTIKGGSPRARVATYKVCWSLT 212
KLP SQ+TARDFVGHGTHTLSTAGGNFVPGASIFNIGNGTIKGGSPRARVATYKVCWSLT
Sbjct: 361 KLPTSQRTARDFVGHGTHTLSTAGGNFVPGASIFNIGNGTIKGGSPRARVATYKVCWSLT 540
Query: 213 DATSCFGADVLSAIDQAIDDGVDIISVSAGGPSSTNSEEIFTDEISIGAFHALA 266
DA SCFGADVLSAIDQAIDDGVDII VSAGGPSSTNSEEIFTDE+SIGAFHALA
Sbjct: 541 DAASCFGADVLSAIDQAIDDGVDIIFVSAGGPSSTNSEEIFTDEVSIGAFHALA 702
>TC77557 similar to PIR|JC7519|JC7519 subtilisin-like serine proteinase (EC
3.4.21.-) - Arabidopsis thaliana, partial (88%)
Length = 2682
Score = 392 bits (1006), Expect = e-109
Identities = 228/535 (42%), Positives = 321/535 (59%), Gaps = 4/535 (0%)
Frame = +3
Query: 17 IIYSYNKQINGFAAMLEEEEAAQIAKNPKVVSVFLSKEHKLHTTRSWEFLGLRGNDINSA 76
++Y+Y I+GF+ L EEA + +++V +++LHTTR+ +FLGL +
Sbjct: 429 MMYTYENAIHGFSTRLTPEEARLLESQTGILAVLPEVKYELHTTRTPQFLGL--DKSADM 602
Query: 77 WQKGRFGENTIIGNIDTGVWPESKSFSDRGIGPIPAKWRGGNICQLDKLNTSKKVPCNRK 136
+ + G ++G +DTGVWPESKSF+D G GPIP W+G C+ T+ CN+K
Sbjct: 603 FPESSSGNEVVVGVLDTGVWPESKSFNDAGFGPIPTTWKGA--CESGTNFTAAN--CNKK 770
Query: 137 LIGARFFNKAYQKRNGKLPRSQQTA--RDFVGHGTHTLSTAGGNFVPGASIFNIGNGTIK 194
LIGARFF+K + G + + ++ RD GHGTHT STA G+ VP AS+F +GT +
Sbjct: 771 LIGARFFSKGVEAMLGPIDETTESKSPRDDDGHGTHTSSTAAGSVVPDASLFGYASGTAR 950
Query: 195 GGSPRARVATYKVCWSLTDATSCFGADVLSAIDQAIDDGVDIISVSAGGPSSTNSEEIFT 254
G + RARVA YKVCW CF +D+L+AID+AI D V+++S+S GG S + F
Sbjct: 951 GMATRARVAVYKVCWK----GGCFSSDILAAIDKAISDNVNVLSLSLGGGMS----DYFR 1106
Query: 255 DEISIGAFHALARNILLVASAGNEGPTPGSVVNVAPWVFTVAASTLDRDFSSVMTIGN-K 313
D ++IGAF A+ + IL+ SAGN GP+ S+ NVAPW+ TV A TLDRDF + +++GN
Sbjct: 1107DSVAIGAFSAMEKGILVSCSAGNAGPSAYSLSNVAPWITTVGAGTLDRDFPASVSLGNGL 1286
Query: 314 TLTGASLF-VNLPPNQDFTIVTSTDAKLANATNRDARFCRPRTLDPSKVNGKIVACDREG 372
+G SL+ N P ++ + +A NATN C TL P V GKIV CDR G
Sbjct: 1287NYSGVSLYRGNALPESPLPLIYAGNA--TNATN--GNLCMTGTLSPELVAGKIVLCDR-G 1451
Query: 373 KIKSVAEGQEALSAGAKGVILRNQPEINGKTLLSEPHVLSTISYPGNHSRTTGRSLDIIP 432
V +G +AG G++L N NG+ L+++ H+L + + L
Sbjct: 1452MNARVQKGAVVKAAGGLGMVLSN-TAANGEELVADTHLLPATAVGEREGNAIKKYLF--- 1619
Query: 433 SDIKSGTKLRMSPAKTLNRRKPAPVMASYSSRGPNKVQPSILKPDVTAPGVNILAAYSLF 492
S+ K K+ K +P+PV+A++SSRGPN + P ILKPD+ APGVNILA +S
Sbjct: 1620SEAKPTVKIVFQGTKV--GVEPSPVVAAFSSRGPNSITPQILKPDLIAPGVNILAGWSKA 1793
Query: 493 ASASNLITDTRRGFPFNVMQGTSMSCPHVAGTAGLIKTLHPNWSPAAIKSAIRTT 547
+ L D RR FN++ GTSMSCPHV+G A LIK+ HP+WSPAA++SA+ TT
Sbjct: 1794VGPTGLAVDERR-VDFNIISGTSMSCPHVSGLAALIKSAHPDWSPAAVRSALMTT 1955
>TC86329 similar to PIR|T07171|T07171 subtilisin-like proteinase (EC
3.4.21.-) 1 - tomato, partial (91%)
Length = 2671
Score = 206 bits (524), Expect(2) = 3e-95
Identities = 118/279 (42%), Positives = 165/279 (58%), Gaps = 4/279 (1%)
Frame = +2
Query: 18 IYSYNKQINGFAAMLEEEEAAQIAKNPKVVSVFLSKEHKLHTTRSWEFLGLRGNDINSAW 77
+Y+Y +GF+ L +EA + K P ++SV ++LHTTR+ EFLGL I
Sbjct: 284 MYTYKHVAHGFSTRLTTQEADLLTKQPGILSVIPDVRYELHTTRTPEFLGLE-KTITLLP 460
Query: 78 QKGRFGENTIIGNIDTGVWPESKSFSDRGIGPIPAKWRGGNICQLDKLNTSKKVPCNRKL 137
G+ E I+G IDTGVWPE KSF D G+GP+P W+G C+ K S CN+KL
Sbjct: 461 SSGKQSE-VIVGVIDTGVWPELKSFDDTGLGPVPKSWKGE--CETGKTFNSSN--CNKKL 625
Query: 138 IGARFFNKAYQKRNGKLPRSQQTA--RDFVGHGTHTLSTAGGNFVPGASIFNIGNGTIKG 195
+GARFF K Y+ G + + ++ RD GHG+HT +TA G+ V GAS+F +GT KG
Sbjct: 626 VGARFFAKGYEAAFGPIDENTESKSPRDDDGHGSHTSTTAAGSAVAGASLFGFASGTAKG 805
Query: 196 GSPRARVATYKVCWSLTDATSCFGADVLSAIDQAIDDGVDIIS--VSAGGPSSTNSEEIF 253
+ +ARVA YKVCW CF +D+ +AID+AID + VS G T + +
Sbjct: 806 MATQARVAAYKVCW----LGGCFTSDIAAAIDKAIDTQARTAAYKVSRLGGGLT---DYY 964
Query: 254 TDEISIGAFHALARNILLVASAGNEGPTPGSVVNVAPWV 292
D +++G F A+ IL+ +SAGN GP+ S+ NVAPW+
Sbjct: 965 KDTVAMGTFAAIEHGILVSSSAGNGGPSKASLANVAPWI 1081
Score = 161 bits (407), Expect(2) = 3e-95
Identities = 103/260 (39%), Positives = 145/260 (55%), Gaps = 6/260 (2%)
Frame = +3
Query: 294 TVAASTLDRDFSSVMTIGN-KTLTGASLFVN-LPPNQDFTIVTSTDAKLANATNRDARFC 351
TV A T+DRDF + +T+GN G SL+ LPPN +V + + ++ N C
Sbjct: 1086 TVGARTIDRDFPAYITLGNGNRYNGVSLYNGKLPPNSPLPLVYAANVSQDSSDN----LC 1253
Query: 352 RPRTLDPSKVNGKIVACDREGKIKSVAEGQEALSAGAKGVILRNQPEINGKTLLSEPHVL 411
+L PSKV+GKIV CDR G ++ + AG G+IL N + G+ L+++ +L
Sbjct: 1254 STDSLIPSKVSGKIVICDRGGNPRA-EKSLVVKRAGGIGMILANNQDY-GEELVADSFLL 1427
Query: 412 STISYPGNHSRTTGRSLDIIPSDIKS----GTKLRMSPAKTLNRRKPAPVMASYSSRGPN 467
+ S + P+ GT+ + +P+PV+A++SSRGPN
Sbjct: 1428 PAAALGEKASNEIKKYASSAPNPTAKIAFGGTRFGV---------QPSPVVAAFSSRGPN 1580
Query: 468 KVQPSILKPDVTAPGVNILAAYSLFASASNLITDTRRGFPFNVMQGTSMSCPHVAGTAGL 527
+ P ILKPD+ APGVNILA +S + L DTR FN++ GTSMSCPHV+G A L
Sbjct: 1581 ILTPKILKPDLIAPGVNILAGWSGKVGPTGLSVDTRH-VSFNIISGTSMSCPHVSGLAAL 1757
Query: 528 IKTLHPNWSPAAIKSAIRTT 547
+K HP WSPAAI+SA+ TT
Sbjct: 1758 LKGAHPEWSPAAIRSALMTT 1817
>TC85584 weakly similar to PIR|A84473|A84473 probable serine proteinase
[imported] - Arabidopsis thaliana, partial (19%)
Length = 1241
Score = 342 bits (878), Expect = 2e-94
Identities = 173/175 (98%), Positives = 174/175 (98%)
Frame = +1
Query: 373 KIKSVAEGQEALSAGAKGVILRNQPEINGKTLLSEPHVLSTISYPGNHSRTTGRSLDIIP 432
+IKSVAEGQEALSAGAKGVILRNQPEINGKTLLSEPHVLSTISYPGNHSRTTGRSLDIIP
Sbjct: 1 EIKSVAEGQEALSAGAKGVILRNQPEINGKTLLSEPHVLSTISYPGNHSRTTGRSLDIIP 180
Query: 433 SDIKSGTKLRMSPAKTLNRRKPAPVMASYSSRGPNKVQPSILKPDVTAPGVNILAAYSLF 492
SDIKSGTKLRMSPAKTLNRRKPAPVMASYSSRGPNKVQPSILKPDVTAPGVNILAAYSLF
Sbjct: 181 SDIKSGTKLRMSPAKTLNRRKPAPVMASYSSRGPNKVQPSILKPDVTAPGVNILAAYSLF 360
Query: 493 ASASNLITDTRRGFPFNVMQGTSMSCPHVAGTAGLIKTLHPNWSPAAIKSAIRTT 547
ASASNLITDTRRGFPFNVMQGTSMSCPHVAGTAGLIKTLHPNWSPAAIKSAI TT
Sbjct: 361 ASASNLITDTRRGFPFNVMQGTSMSCPHVAGTAGLIKTLHPNWSPAAIKSAIMTT 525
>BG580583 weakly similar to GP|9757901|dbj serine protease-like protein
{Arabidopsis thaliana}, partial (13%)
Length = 592
Score = 337 bits (865), Expect = 5e-93
Identities = 162/190 (85%), Positives = 173/190 (90%)
Frame = +3
Query: 94 GVWPESKSFSDRGIGPIPAKWRGGNICQLDKLNTSKKVPCNRKLIGARFFNKAYQKRNGK 153
GVWPES+SF+DRGIGPIP +WRGGNICQLDKLNTSKKVPCNRKLIGARFFNKAY+ +GK
Sbjct: 21 GVWPESESFNDRGIGPIPLRWRGGNICQLDKLNTSKKVPCNRKLIGARFFNKAYEAFHGK 200
Query: 154 LPRSQQTARDFVGHGTHTLSTAGGNFVPGASIFNIGNGTIKGGSPRARVATYKVCWSLTD 213
LP SQQTARDFVGHGTHTLSTAGGNFV A+IF IGNGTIKGGSPR+RVATYK CWSLTD
Sbjct: 201 LPSSQQTARDFVGHGTHTLSTAGGNFVQNATIFGIGNGTIKGGSPRSRVATYKACWSLTD 380
Query: 214 ATSCFGADVLSAIDQAIDDGVDIISVSAGGPSSTNSEEIFTDEISIGAFHALARNILLVA 273
CFGADVL+AIDQ+I DG D+ISVSAGG +TN E IFTDEISIGAFHALARNILLVA
Sbjct: 381 VVDCFGADVLAAIDQSIYDGADLISVSAGGKPNTNPEVIFTDEISIGAFHALARNILLVA 560
Query: 274 SAGNEGPTPG 283
SAGNEGPTPG
Sbjct: 561 SAGNEGPTPG 590
>TC85586 similar to GP|20521377|dbj|BAB91889. putative subtilisin-like
protease {Oryza sativa (japonica cultivar-group)},
partial (17%)
Length = 1460
Score = 329 bits (843), Expect = 2e-90
Identities = 169/212 (79%), Positives = 183/212 (85%), Gaps = 1/212 (0%)
Frame = +1
Query: 337 DAKLANATNRDARFCRPRTLDPSKVNGKIVACDREGKIKSVAEGQEALSAGAKGVILRNQ 396
DAK +NAT RDARFCRPRTLDPSKV GKIVAC REGKIKSVAEGQEALSAGAKG+ L NQ
Sbjct: 4 DAKFSNATTRDARFCRPRTLDPSKVKGKIVACAREGKIKSVAEGQEALSAGAKGMFLENQ 183
Query: 397 PEINGKTLLSEPHVLSTISYPGNHSRTTGRSLDIIPSD-IKSGTKLRMSPAKTLNRRKPA 455
P+++G TLLSEPHVLST+ G + T L + +D I+SGTK+R S A TL RKPA
Sbjct: 184 PKVSGNTLLSEPHVLSTVGGNGQAAITAPPRLGVTATDTIESGTKIRFSQAITLIGRKPA 363
Query: 456 PVMASYSSRGPNKVQPSILKPDVTAPGVNILAAYSLFASASNLITDTRRGFPFNVMQGTS 515
PVMAS+SSRGPN+VQP ILKPDVTAPGVNILAAYSLFASASNL+TD RRGFPFNVMQGTS
Sbjct: 364 PVMASFSSRGPNQVQPYILKPDVTAPGVNILAAYSLFASASNLLTDNRRGFPFNVMQGTS 543
Query: 516 MSCPHVAGTAGLIKTLHPNWSPAAIKSAIRTT 547
MSCPHVAGTAGLIKTLHPNWSPAAIKSAI TT
Sbjct: 544 MSCPHVAGTAGLIKTLHPNWSPAAIKSAIMTT 639
>TC85588 similar to GP|20198169|gb|AAM15440.1 subtilisin-like serine
protease AIR3 {Arabidopsis thaliana}, partial (7%)
Length = 572
Score = 280 bits (715), Expect = 1e-75
Identities = 146/217 (67%), Positives = 163/217 (74%)
Frame = +1
Query: 196 GSPRARVATYKVCWSLTDATSCFGADVLSAIDQAIDDGVDIISVSAGGPSSTNSEEIFTD 255
GSPRARVATYKVCWSL D CFGADVL+AIDQAI DGVDIIS+S G S E+IFTD
Sbjct: 1 GSPRARVATYKVCWSLLDLEDCFGADVLAAIDQAISDGVDIISLSLAGHSLVYPEDIFTD 180
Query: 256 EISIGAFHALARNILLVASAGNEGPTPGSVVNVAPWVFTVAASTLDRDFSSVMTIGNKTL 315
E+SIGAFHAL+RNILLVASAGNEGPT GSVVNV PWVFT+AASTLDRDFSS +TIGN+T+
Sbjct: 181 EVSIGAFHALSRNILLVASAGNEGPTGGSVVNVTPWVFTIAASTLDRDFSSTITIGNQTI 360
Query: 316 TGASLFVNLPPNQDFTIVTSTDAKLANATNRDARFCRPRTLDPSKVNGKIVACDREGKIK 375
GASLFVNLPPNQ F ++ STD KLANATN DA+FC+P TLDPSKV
Sbjct: 361 RGASLFVNLPPNQAFPLIVSTDGKLANATNHDAQFCKPGTLDPSKV-------------- 498
Query: 376 SVAEGQEALSAGAKGVILRNQPEINGKTLLSEPHVLS 412
KG++L NQP+ GKT L+EPH LS
Sbjct: 499 -------------KGMLLSNQPK-QGKTTLAEPHTLS 567
>TC81705 weakly similar to GP|6721520|dbj|BAA89562.1 EST AU029428(E30359)
corresponds to a region of the predicted gene.~Similar
to antifreeze-like, partial (33%)
Length = 1443
Score = 262 bits (669), Expect = 3e-70
Identities = 150/330 (45%), Positives = 202/330 (60%), Gaps = 6/330 (1%)
Frame = +1
Query: 1 DLLGSILGSKENAKEAIIYSYNKQINGFAAMLEEEEAAQIAKNPKVVSVFLSKEHKLHTT 60
+LL SI+ S+E+ + +I+ YN +GF+AML + EA+ ++ + VVSVF +LHTT
Sbjct: 151 ELLSSIIPSEESERIKLIHHYNHAFSGFSAMLTQSEASALSGHDGVVSVFPDPILELHTT 330
Query: 61 RSWEFLG--LRGNDINSAWQKGRFGENTIIGNIDTGVWPESKSFSDRGIGPIPAKWRGGN 118
RSW+FL L + + II IDTG+WPES SF+D GIG IP+ W+G
Sbjct: 331 RSWDFLDSDLGMKPSTNVLTHQHSSNDIIIALIDTGIWPESPSFTDEGIGKIPSVWKG-- 504
Query: 119 ICQLDKLNTSKKVPCNRKLIGARFFNK--AYQKRNGKLPRSQQTARDFVGHGTHTLSTAG 176
IC + + KK CNRKLIGAR++N + + ++ + RD VGHGTHT STA
Sbjct: 505 ICM--EGHDFKKSNCNRKLIGARYYNTQDTFGSNKTHIGGAKGSPRDTVGHGTHTASTAA 678
Query: 177 GNFVPGASIFNIGNGTIKGGSPRARVATYKVCWSLTDATSCFGADVLSAIDQAIDDGVDI 236
G V A+ + + GT +GGSP R+A YK C C G+ +L A+D AI DGVDI
Sbjct: 679 GVNVNNANYYGLAKGTARGGSPSTRIAAYKTC----SEEGCSGSTILKAMDDAIKDGVDI 846
Query: 237 ISVSAGGPSSTNSEEIFTDEISIGAFHALARNILLVASAGNEGPTPGSVVNVAPWVFTVA 296
IS+S G SS + D I+IGAFHA R + V SAGN+GP P +VVN APW+FTVA
Sbjct: 847 ISISIG-LSSLMQSDYLNDPIAIGAFHAEQRGVTAVCSAGNDGPDPNTVVNTAPWIFTVA 1023
Query: 297 ASTLDRDFSSVMTIGN-KTLTGASL-FVNL 324
AS +DR+F S + +GN K+ GA + F NL
Sbjct: 1024ASNIDRNFQSTIVLGNGKSFQGAGINFSNL 1113
>TC83163 weakly similar to GP|6721520|dbj|BAA89562.1 EST AU029428(E30359)
corresponds to a region of the predicted gene.~Similar
to antifreeze-like, partial (29%)
Length = 990
Score = 252 bits (643), Expect = 3e-67
Identities = 145/297 (48%), Positives = 187/297 (62%), Gaps = 4/297 (1%)
Frame = +2
Query: 1 DLLGSILGSKENAKEAIIYSYNKQINGFAAMLEEEEAAQIAKNPKVVSVFLSKEHKLHTT 60
DLL SI+ S+E+ + A+I+ Y+ NGF+AML + EA+ +A N VVSVF +LHTT
Sbjct: 161 DLLSSIIPSEESERIALIHHYSHAFNGFSAMLTQSEASALAGNDGVVSVFEDPFLELHTT 340
Query: 61 RSWEF----LGLRGNDINSAWQKGRFGENTIIGNIDTGVWPESKSFSDRGIGPIPAKWRG 116
RSW+F LG+R + I + + IIG IDTG+WPES SF D GIG IP++W+G
Sbjct: 341 RSWDFLESDLGMRPHGI---LKHQHSSNDIIIGVIDTGIWPESPSFKDEGIGKIPSRWKG 511
Query: 117 GNICQLDKLNTSKKVPCNRKLIGARFFNKAYQKRNGKLPRSQQTARDFVGHGTHTLSTAG 176
+C + + KK CNRKLIGAR++NK K P+ + RDF GHGTHT STA
Sbjct: 512 --VCM--EAHDFKKSNCNRKLIGARYYNK-------KDPKG--SPRDFNGHGTHTASTAA 652
Query: 177 GNFVPGASIFNIGNGTIKGGSPRARVATYKVCWSLTDATSCFGADVLSAIDQAIDDGVDI 236
G V AS + + GT +GGSP AR+A YK C C G +L AID AI DGVDI
Sbjct: 653 GVIVNNASYYGLAKGTARGGSPSARIAAYKAC----SGEGCSGGTLLKAIDDAIKDGVDI 820
Query: 237 ISVSAGGPSSTNSEEIFTDEISIGAFHALARNILLVASAGNEGPTPGSVVNVAPWVF 293
IS+S G S SE + +D I+IGAFHA R +++V SAGNEGP +VVN PW+F
Sbjct: 821 ISISIGFSSEFLSEYL-SDPIAIGAFHAEQRGVMVVCSAGNEGPDHYTVVNTTPWIF 988
>TC89220 similar to PIR|T05768|T05768 subtilisin-like proteinase (EC
3.4.21.-) - Arabidopsis thaliana, partial (39%)
Length = 1149
Score = 248 bits (634), Expect = 3e-66
Identities = 126/302 (41%), Positives = 195/302 (63%), Gaps = 8/302 (2%)
Frame = +3
Query: 17 IIYSYNKQINGFAAMLEEEEAAQIAKNPKVVSVFLSKEHKLHTTRSWEFLGLRGNDINSA 76
I+++Y+ +GF+A+L ++ A I+ +P +++VF + +LHTTRS +FLGLR
Sbjct: 228 ILHTYDTAFHGFSAVLTRQQVASISNHPSILAVFEDRRRQLHTTRSPQFLGLRNQ--RGL 401
Query: 77 WQKGRFGENTIIGNIDTGVWPESKSFSDRGIGPIPAKWRGGNICQLDKLNTSKKVPCNRK 136
W + +G + I+G DTG+WPE +SFSD +GPIP +W+G +C+ + + + CNRK
Sbjct: 402 WSESDYGSDVIVGVFDTGIWPERRSFSDMNLGPIPRRWKG--VCESGEKFSPRN--CNRK 569
Query: 137 LIGARFFNKAYQKRNGKL----PRSQ----QTARDFVGHGTHTLSTAGGNFVPGASIFNI 188
LIGAR+F+K ++ G P ++ ++ RD GHGTHT STA G + A++
Sbjct: 570 LIGARYFSKGHEVGAGSAGPLNPINETVEFRSPRDADGHGTHTASTAAGRYAFQANMSGY 749
Query: 189 GNGTIKGGSPRARVATYKVCWSLTDATSCFGADVLSAIDQAIDDGVDIISVSAGGPSSTN 248
+G KG +P+AR+A YKVCW + CF +D+L+A D A++DGVD+IS+S GG
Sbjct: 750 ASGIAKGVAPKARLAVYKVCWK---NSGCFDSDILAAFDAAVNDGVDVISISIGGGDGIA 920
Query: 249 SEEIFTDEISIGAFHALARNILLVASAGNEGPTPGSVVNVAPWVFTVAASTLDRDFSSVM 308
S + D I+IG++ A++R + + +SAGN+GP+ SV N+APW+ TV A T+DRDF S +
Sbjct: 921 SP-YYLDPIAIGSYGAVSRGVFVSSSAGNDGPSGMSVTNLAPWLTTVGAGTIDRDFPSQI 1097
Query: 309 TI 310
I
Sbjct: 1098II 1103
>TC87377 weakly similar to PIR|T07171|T07171 subtilisin-like proteinase (EC
3.4.21.-) 1 - tomato, partial (54%)
Length = 2071
Score = 191 bits (484), Expect(2) = 1e-60
Identities = 122/294 (41%), Positives = 177/294 (59%), Gaps = 6/294 (2%)
Frame = +2
Query: 200 ARVATYKVCWSLTDATSCFGADVLSAIDQAIDDGVDIISVSAGGPSSTNSEEIFTDEISI 259
ARVA YKVCW + CF +D+ + +D+AI+DGV+I+S+S GG + + + D I+I
Sbjct: 11 ARVAAYKVCW----LSGCFTSDIAAGMDKAIEDGVNILSMSIGG----SIMDYYRDIIAI 166
Query: 260 GAFHALARNILLVASAGNEGPTPGSVVNVAPWVFTVAASTLDRDFSSVMTIGN-KTLTGA 318
GAF A++ IL+ +SAGN GP+ S+ NVAPW+ TV A T+DRDF S +T+GN KT TGA
Sbjct: 167 GAFTAMSHGILVSSSAGNGGPSAESLSNVAPWITTVGAGTIDRDFPSYITLGNGKTYTGA 346
Query: 319 SLFVNLPPNQDFTIVTSTDAKLANATNRDARFCRPRTLDPSKVNGKIVACDREGKIKSVA 378
SL+ N P+ D + ++ ++ C P +L SKV GKIV C+R G + V
Sbjct: 347 SLY-NGKPSSDSLLPVVYAGNVSESS--VGYLCIPDSLTSSKVLGKIVICERGGNSR-VE 514
Query: 379 EGQEALSAGAKGVILRNQPEINGKTLLSEPHVLSTISYPGNHSRTTGRSLDIIPSD---- 434
+G +AG G+IL N E G+ L+++ H+L + G S T + +
Sbjct: 515 KGLVVKNAGGVGMILVNN-EAYGEELIADSHLLPAAAL-GQKSSTVLKDYVFTTKNPRAK 688
Query: 435 -IKSGTKLRMSPAKTLNRRKPAPVMASYSSRGPNKVQPSILKPDVTAPGVNILA 487
+ GT L++ +P+PV+A++SSRGPN + P ILKPD+ APGVNILA
Sbjct: 689 LVFGGTHLQV---------QPSPVVAAFSSRGPNSLTPKILKPDLIAPGVNILA 823
Score = 61.2 bits (147), Expect(2) = 1e-60
Identities = 25/45 (55%), Positives = 34/45 (75%)
Frame = +1
Query: 503 RRGFPFNVMQGTSMSCPHVAGTAGLIKTLHPNWSPAAIKSAIRTT 547
+R FN++ GTSMSCPH +G A ++K +P WSPAAI+SA+ TT
Sbjct: 868 KRHVNFNIISGTSMSCPHASGLAAIVKGAYPEWSPAAIRSALMTT 1002
>TC85585 similar to GP|22773236|gb|AAN06842.1 Putatvie subtilisin-like
serine protease {Oryza sativa (japonica
cultivar-group)}, partial (21%)
Length = 1353
Score = 214 bits (545), Expect = 7e-56
Identities = 106/138 (76%), Positives = 115/138 (82%)
Frame = +2
Query: 410 VLSTISYPGNHSRTTGRSLDIIPSDIKSGTKLRMSPAKTLNRRKPAPVMASYSSRGPNKV 469
V+STI+Y S TT + +I P DIK+ +RMSPA LN RKPAPVMAS+SSRGPNKV
Sbjct: 5 VVSTINYHDPRSITTPKGSEITPEDIKTNATIRMSPANALNGRKPAPVMASFSSRGPNKV 184
Query: 470 QPSILKPDVTAPGVNILAAYSLFASASNLITDTRRGFPFNVMQGTSMSCPHVAGTAGLIK 529
QP ILKPDVTAPGVNILAAYSL AS SNL+TD RRGFPFN+ QGTSMSCPHV GTAGLIK
Sbjct: 185 QPYILKPDVTAPGVNILAAYSLLASVSNLVTDNRRGFPFNIQQGTSMSCPHVVGTAGLIK 364
Query: 530 TLHPNWSPAAIKSAIRTT 547
TLHPNWSPAAIKSAI TT
Sbjct: 365 TLHPNWSPAAIKSAIMTT 418
>BE249159 weakly similar to GP|20198252|gb subtilisin-like serine protease
AIR3 {Arabidopsis thaliana}, partial (24%)
Length = 661
Score = 214 bits (545), Expect = 7e-56
Identities = 114/198 (57%), Positives = 137/198 (68%), Gaps = 1/198 (0%)
Frame = +1
Query: 134 NRKLIGARFFNKAYQKRNGKLPRSQQTARDFVGHGTHTLSTAGGNFVPGASIFNIGNGTI 193
+RKLIGAR F K Y+ GKL S TARD +GHG+HTLSTAGGNFV G S++ GNGT
Sbjct: 82 SRKLIGARAFYKGYEAYVGKLDASFYTARDTIGHGSHTLSTAGGNFVQGVSVYGNGNGTA 261
Query: 194 KGGSPRARVATYKVCWSLTDATSCFGADVLSAIDQAIDDGVDIISVSAGGPSSTNSEEIF 253
KGGSP+A VA YKVCW C ADVL+ + AI DGVD++SVS G + +F
Sbjct: 262 KGGSPKAHVAAYKVCWK----GGCSDADVLAGFEAAISDGVDVLSVSLG----MKTHNLF 417
Query: 254 TDEISIGAFHALARNILLVASAGNEGPTPGSVVNVAPWVFTVAASTLDRDFSSVMTIG-N 312
TD ISIG+FHA+A I++VASAGN GP G+V NVAPW+FTVAAST+DRDF+S +T+G N
Sbjct: 418 TDSISIGSFHAVANGIVVVASAGNSGPYFGTVSNVAPWLFTVAASTIDRDFASYVTLGDN 597
Query: 313 KTLTGASLFVNLPPNQDF 330
K G SL P F
Sbjct: 598 KHFKGTSLSSKDLPTHKF 651
>AW695319 similar to GP|6708179|gb| subtilisin-type serine endopeptidase XSP1
{Arabidopsis thaliana}, partial (27%)
Length = 648
Score = 187 bits (475), Expect = 9e-48
Identities = 111/237 (46%), Positives = 145/237 (60%), Gaps = 2/237 (0%)
Frame = +2
Query: 56 KLHTTRSWEFLGLRGNDINSAWQKGRFGENTIIGNIDTGVWPESKSFSDRGIGPIPAKWR 115
KLHTTRSW+F+GL +A +K + +TI+ +DTG+ PE +SF D G GP PAKW+
Sbjct: 5 KLHTTRSWDFIGLP----LTAKRKLKSEGDTIVALLDTGITPEFQSFKDDGFGPPPAKWK 172
Query: 116 GGNICQLDK-LNTSKKVPCNRKLIGARFFNKAYQKRNGKL-PRSQQTARDFVGHGTHTLS 173
G DK +N S CN K+IGA++F K +G+ P + D GHGTHT S
Sbjct: 173 G----TCDKYVNFSG---CNNKIIGAKYF-----KLDGRSNPSDILSPIDVEGHGTHTAS 316
Query: 174 TAGGNFVPGASIFNIGNGTIKGGSPRARVATYKVCWSLTDATSCFGADVLSAIDQAIDDG 233
TA GN VP AS+F + G +G AR+A YK+CW+ C D+L+A + AI DG
Sbjct: 317 TAAGNIVPNASLFGLAKGMARGAVHSARLAIYKICWT---EDGCADMDILAAFEAAIHDG 487
Query: 234 VDIISVSAGGPSSTNSEEIFTDEISIGAFHALARNILLVASAGNEGPTPGSVVNVAP 290
VD+ISVS GG +E D I+IGAFHA+ + I+ VASAGN GPT +VVN AP
Sbjct: 488 VDVISVSLGG----GNENYAQDSIAIGAFHAMRKGIITVASAGNGGPTMATVVNNAP 646
>CB892799 weakly similar to PIR|T05768|T05 subtilisin-like proteinase (EC
3.4.21.-) - Arabidopsis thaliana, partial (20%)
Length = 875
Score = 185 bits (469), Expect = 4e-47
Identities = 118/297 (39%), Positives = 177/297 (58%), Gaps = 4/297 (1%)
Frame = +2
Query: 255 DEISIGAFHALARNILLVASAGNEGPTPGSVVNVAPWVFTVAASTLDRDFSSVMTIGNKT 314
D ISIG+FHA R + +V+SAGNEG GS N+APW+ TVAA + DRDF+S + +GN
Sbjct: 2 DAISIGSFHAANRGLFVVSSAGNEGNL-GSATNLAPWMLTVAAGSTDRDFTSDIILGNGA 178
Query: 315 -LTGASLFVNLPPNQDFTIVTSTDAKLANATNRDARFCRPRTLDPSKVNGKIVAC---DR 370
+TG SL + N I+++++A T + +C +L+ +K GK++ C +R
Sbjct: 179 KITGESLSL-FEMNASTRIISASEAFAGYFTPYQSSYCLESSLNKTKTKGKVLVCRHVER 355
Query: 371 EGKIKSVAEGQEALSAGAKGVILRNQPEINGKTLLSEPHVLSTISYPGNHSRTTGRSLDI 430
+ K VA+ + AG G+IL ++ + + ++ P V+ + + + L
Sbjct: 356 STESK-VAKSKIVKEAGGVGMILIDETDQD----VAIPFVIPSAIVGKKKGQ---KILSY 511
Query: 431 IPSDIKSGTKLRMSPAKTLNRRKPAPVMASYSSRGPNKVQPSILKPDVTAPGVNILAAYS 490
+ + K +K+ AKT+ + AP +A++SSRGPN + P ILKPD+TAPG+NILAA+S
Sbjct: 512 LKTTRKPMSKILR--AKTVIGAQSAPRVAAFSSRGPNALNPEILKPDITAPGLNILAAWS 685
Query: 491 LFASASNLITDTRRGFPFNVMQGTSMSCPHVAGTAGLIKTLHPNWSPAAIKSAIRTT 547
A G FN++ GTSM+CPHV G A L+K +HP+WSP+AIKSAI TT
Sbjct: 686 PVA-----------GNMFNILSGTSMACPHVTGIATLVKAVHPSWSPSAIKSAIMTT 823
>TC78957 similar to GP|14091078|gb|AAK53589.1 subtilisin-like protein
{Glycine max}, partial (28%)
Length = 838
Score = 181 bits (460), Expect = 5e-46
Identities = 105/233 (45%), Positives = 141/233 (60%), Gaps = 6/233 (2%)
Frame = +1
Query: 14 KEAIIYSYNKQINGFAAMLEEEEAAQIAKNPKVVSVFLSKEHKLHTTRSWEFLGLRGND- 72
++A++++Y +GFAA L + EAA IA+ P VVSVF KLHTT SW+FL L+ +
Sbjct: 181 EKALVHNYKHGFSGFAARLSKNEAASIAQQPGVVSVFPDPILKLHTTHSWDFLKLQTHVK 360
Query: 73 -----INSAWQKGRFGENTIIGNIDTGVWPESKSFSDRGIGPIPAKWRGGNICQLDKLNT 127
NS+ Q + +IG +D+G+WPE+ SFSD G+ PIP+ W+G IC N
Sbjct: 361 IDSTLSNSSSQSS--SSDIVIGMLDSGIWPEATSFSDNGMDPIPSGWKG--ICMTS--ND 522
Query: 128 SKKVPCNRKLIGARFFNKAYQKRNGKLPRSQQTARDFVGHGTHTLSTAGGNFVPGASIFN 187
CNRK+IGAR+ Y G R T RD VGHGTHT STA GN V GAS +
Sbjct: 523 FNSSNCNRKIIGARY----YPNLEGD-DRVAATTRDTVGHGTHTASTAAGNAVSGASYYG 687
Query: 188 IGNGTIKGGSPRARVATYKVCWSLTDATSCFGADVLSAIDQAIDDGVDIISVS 240
+ G KGGSP +R+A KVC ++ C G+ +L+A D AI DGVD++S+S
Sbjct: 688 LAEGIAKGGSPESRLAI*KVCSNI----GCSGSAILAAFDDAISDGVDVLSLS 834
>BG449655 similar to GP|20198252|gb subtilisin-like serine protease AIR3
{Arabidopsis thaliana}, partial (21%)
Length = 664
Score = 178 bits (452), Expect = 4e-45
Identities = 93/154 (60%), Positives = 108/154 (69%), Gaps = 4/154 (2%)
Frame = +1
Query: 1 DLLGSILGSKENAKEAIIYSYNKQINGFAAMLEEEEAAQIAKNPKVVSVFLSKEHKLHTT 60
+ LGS LGS + AKE+I YSY + INGFAA LEEE AA+IAK+PKV+SVF + KLHTT
Sbjct: 193 EFLGSFLGSSKTAKESIFYSYTRHINGFAATLEEEVAAEIAKHPKVLSVFENNGRKLHTT 372
Query: 61 RSWEFLGLRGN----DINSAWQKGRFGENTIIGNIDTGVWPESKSFSDRGIGPIPAKWRG 116
SW F+GL + +S W K RFG+ II N+DTGVWPESKSFSD G GPIP+KWRG
Sbjct: 373 HSWGFMGLEDSYGVIPSSSIWNKARFGDGIIIANLDTGVWPESKSFSDEGFGPIPSKWRG 552
Query: 117 GNICQLDKLNTSKKVPCNRKLIGARFFNKAYQKR 150
IC DK CNRKLIGAR+FNK Y R
Sbjct: 553 --IC--DK-GRDPSFHCNRKLIGARYFNKGYASR 639
>BI307979 weakly similar to GP|20521377|db putative subtilisin-like protease
{Oryza sativa (japonica cultivar-group)}, partial (18%)
Length = 777
Score = 175 bits (444), Expect = 4e-44
Identities = 105/268 (39%), Positives = 159/268 (59%), Gaps = 1/268 (0%)
Frame = +3
Query: 223 LSAIDQAIDDGVDIISVSAGGPSSTNSEEIFTDEISIGAFHALARNILLVASAGNEGPTP 282
L AID AI+DGVD+I++S G P+ E+ D I+ GA A+ +NI++V SAGN GP+P
Sbjct: 3 LKAIDDAIEDGVDVINLSIGFPAPLKYED---DVIAKGALQAVRKNIVVVCSAGNAGPSP 173
Query: 283 GSVVNVAPWVFTVAASTLDRDFSSVMTIGN-KTLTGASLFVNLPPNQDFTIVTSTDAKLA 341
S+ N +PW+ TV AST+DR F + + + N T+ G S+ N +V ++D + A
Sbjct: 174 HSLSNPSPWIITVGASTVDRTFLAPIKLSNGTTIEGRSITPLRMGNSFCPLVLASDVEYA 353
Query: 342 NATNRDARFCRPRTLDPSKVNGKIVACDREGKIKSVAEGQEALSAGAKGVILRNQPEING 401
+ ++ +C TLDPSKV GKIV C R G+ V + E AG G+IL N ++
Sbjct: 354 GILSANSSYCLDNTLDPSKVKGKIVLCMR-GQGGRVKKSLEVQRAGGVGLILGNN-KVYA 527
Query: 402 KTLLSEPHVLSTISYPGNHSRTTGRSLDIIPSDIKSGTKLRMSPAKTLNRRKPAPVMASY 461
+ S+P+ + T ++ + + P+ + ++ P +T+ KPAP MA +
Sbjct: 528 NDVPSDPYFIPTTGVTYENTLKLVQYIHSSPNPM-----AQLLPGRTVLDTKPAPSMAIF 692
Query: 462 SSRGPNKVQPSILKPDVTAPGVNILAAY 489
SSRGPN + P+ILKPD+TAPGV+I AA+
Sbjct: 693 SSRGPNIIDPNILKPDITAPGVDIFAAW 776
>TC82946 weakly similar to GP|20160863|dbj|BAB89802. putative
subtilisin-like protease {Oryza sativa (japonica
cultivar-group)}, partial (21%)
Length = 921
Score = 95.5 bits (236), Expect(2) = 9e-41
Identities = 53/133 (39%), Positives = 72/133 (53%)
Frame = +1
Query: 135 RKLIGARFFNKAYQKRNGKLPRSQQTARDFVGHGTHTLSTAGGNFVPGASIFNIGNGTIK 194
R LIGA+FFNK Q + + + RD GHGTHT STA GN V AS F GT
Sbjct: 547 RNLIGAKFFNKGLQSKYPNITLGLNSTRDSHGHGTHTSSTAAGNRVDDASFFGYAPGTAS 726
Query: 195 GGSPRARVATYKVCWSLTDATSCFGADVLSAIDQAIDDGVDIISVSAGGPSSTNSEEIFT 254
G + +RVA YK W + +DV++AID AI DGVD++S+S G N ++
Sbjct: 727 GIASNSRVAMYKAIWDV----GVLSSDVIAAIDAAISDGVDVLSLSFG----INDVPLYD 882
Query: 255 DEISIGAFHALAR 267
++I F + +
Sbjct: 883 XPVAIATFAXMEK 921
Score = 90.1 bits (222), Expect(2) = 9e-41
Identities = 53/134 (39%), Positives = 71/134 (52%)
Frame = +3
Query: 3 LGSILGSKENAKEAIIYSYNKQINGFAAMLEEEEAAQIAKNPKVVSVFLSKEHKLHTTRS 62
L S+ SK +IY+Y +NGF+A L +E + +P +S KL TT S
Sbjct: 186 LNSLFSSK------LIYTYTHVMNGFSANLSPKEHESLKTSPGYISSIKDSHMKLDTTHS 347
Query: 63 WEFLGLRGNDINSAWQKGRFGENTIIGNIDTGVWPESKSFSDRGIGPIPAKWRGGNICQL 122
+FLGL N AW FG + I+G IDTG+WPES+SF D + IP+KW G C+
Sbjct: 348 PQFLGLNPN--KGAWHDSNFGNDVIVGLIDTGIWPESESFKDNLMSEIPSKWNGQ--CEN 515
Query: 123 DKLNTSKKVPCNRK 136
L S CN+K
Sbjct: 516 SILFNSSL--CNKK 551
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.315 0.132 0.386
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 15,051,461
Number of Sequences: 36976
Number of extensions: 195160
Number of successful extensions: 1053
Number of sequences better than 10.0: 90
Number of HSP's better than 10.0 without gapping: 933
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 944
length of query: 547
length of database: 9,014,727
effective HSP length: 101
effective length of query: 446
effective length of database: 5,280,151
effective search space: 2354947346
effective search space used: 2354947346
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 61 (28.1 bits)
Medicago: description of AC133779.5


