

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC133779.11 - phase: 0
(1067 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
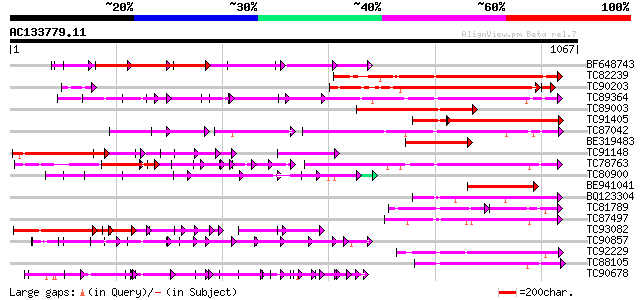
Sequences producing significant alignments: (bits) Value
BF648743 weakly similar to PIR|H84632|H846 probable receptor-lik... 418 e-117
TC82239 weakly similar to GP|16924042|gb|AAL31654.1 Putative pro... 360 2e-99
TC90203 weakly similar to PIR|B86479|B86479 hypothetical protein... 306 4e-84
TC89364 weakly similar to PIR|C96745|C96745 hypothetical protein... 283 3e-76
TC89003 weakly similar to GP|18086327|gb|AAL57627.1 AT4g08850/T3... 276 4e-74
TC91405 weakly similar to GP|16924042|gb|AAL31654.1 Putative pro... 275 7e-74
TC87042 similar to GP|13605827|gb|AAK32899.1 AT5g48380/MJE7_1 {A... 241 8e-64
BE319483 weakly similar to GP|13489172|gb putative protein kinas... 234 1e-61
TC91148 similar to GP|5732036|gb|AAD48937.1| similar to disease ... 234 2e-61
TC78763 somatic embryogenesis receptor kinase 1 [Medicago trunca... 217 2e-56
TC80900 weakly similar to PIR|G84652|G84652 probable receptor-li... 207 1e-53
BE941041 similar to GP|14596041|gb Unknown protein {Arabidopsis ... 204 1e-52
BQ123304 similar to GP|22093756|d putative receptor protein kina... 189 4e-48
TC81789 similar to PIR|B86440|B86440 probable protein kinase [im... 121 1e-47
TC87497 similar to GP|9759480|dbj|BAB10485.1 serine/threonine pr... 183 4e-46
TC93082 similar to GP|1184077|gb|AAC15780.1| Cf-2.2 {Lycopersico... 180 2e-45
TC90857 weakly similar to GP|16930691|gb|AAL32011.1 AT4g26540/M3... 180 3e-45
TC92229 similar to PIR|B86369|B86369 hypothetical protein AAC980... 178 9e-45
TC88105 similar to GP|21593085|gb|AAM65034.1 Putative protein ki... 176 3e-44
TC90678 weakly similar to GP|9759550|dbj|BAB11152.1 receptor pro... 135 2e-36
>BF648743 weakly similar to PIR|H84632|H846 probable receptor-like protein
kinase [imported] - Arabidopsis thaliana, partial (9%)
Length = 648
Score = 418 bits (1075), Expect = e-117
Identities = 212/216 (98%), Positives = 212/216 (98%)
Frame = +1
Query: 162 NLTNLSYLILGGNNWSGGPIPPEIGKLNNLLHLAIQKSNLVGSIPQEIGFLTNLAYIDLS 221
NLTNLSYLILGGNNWSGGPIPPEIGKLNNLLHLAIQKSNLVGSIPQEIGFLTNLAYIDLS
Sbjct: 1 NLTNLSYLILGGNNWSGGPIPPEIGKLNNLLHLAIQKSNLVGSIPQEIGFLTNLAYIDLS 180
Query: 222 KNSLSGGIPETIGNLSKLDTLVLSNNTKMSGPIPHSLWNMSSLTVLYFDNIGLSGSIPDS 281
KNSLSGGIPETIGNLSKLDTLVLSNNTKMSGPIPHSLWNMSSLTVLYFDNIGLSGSIPDS
Sbjct: 181 KNSLSGGIPETIGNLSKLDTLVLSNNTKMSGPIPHSLWNMSSLTVLYFDNIGLSGSIPDS 360
Query: 282 IQNLVNLKELALDINHLSGSIPSTIGDLKNLIKLYLGSNNLSGPIPASIGNLINLQVLSV 341
IQNLVNLKELALDINHLSGSIPSTIGDLKNLIKLYLGSNNLSGPIPASIGNLINLQVLSV
Sbjct: 361 IQNLVNLKELALDINHLSGSIPSTIGDLKNLIKLYLGSNNLSGPIPASIGNLINLQVLSV 540
Query: 342 QENNLTGTIPASIGNLKWLTVFEVATNKLHGRIPNG 377
QENNLTGTIPASIGNLKWLTVFEV TNK RIPNG
Sbjct: 541 QENNLTGTIPASIGNLKWLTVFEVXTNKPSWRIPNG 648
Score = 139 bits (351), Expect = 4e-33
Identities = 86/204 (42%), Positives = 115/204 (56%)
Frame = +1
Query: 101 NNSFYGTIPAQIGNLSNISILTFKNNYFDGSIPQEMCTLTGLQFLDISFCKLNGAIPKSI 160
NN G IP +IG L+N+ L + + GSIPQE+ LT L ++D+S L+G IP++I
Sbjct: 37 NNWSGGPIPPEIGKLNNLLHLAIQKSNLVGSIPQEIGFLTNLAYIDLSKNSLSGGIPETI 216
Query: 161 GNLTNLSYLILGGNNWSGGPIPPEIGKLNNLLHLAIQKSNLVGSIPQEIGFLTNLAYIDL 220
GNL+ L L+L N GPIP + +++L L L GSIP I L NL + L
Sbjct: 217 GNLSKLDTLVLSNNTKMSGPIPHSLWNMSSLTVLYFDNIGLSGSIPDSIQNLVNLKELAL 396
Query: 221 SKNSLSGGIPETIGNLSKLDTLVLSNNTKMSGPIPHSLWNMSSLTVLYFDNIGLSGSIPD 280
N LSG IP TIG+L L L L +N +SGPIP S+ N+ +L VL L+G+IP
Sbjct: 397 DINHLSGSIPSTIGDLKNLIKLYLGSN-NLSGPIPASIGNLINLQVLSVQENNLTGTIPA 573
Query: 281 SIQNLVNLKELALDINHLSGSIPS 304
SI NL L + N S IP+
Sbjct: 574 SIGNLKWLTVFEVXTNKPSWRIPN 645
Score = 120 bits (301), Expect = 3e-27
Identities = 73/214 (34%), Positives = 114/214 (53%), Gaps = 2/214 (0%)
Frame = +1
Query: 308 DLKNLIKLYLGSNNLSG-PIPASIGNLINLQVLSVQENNLTGTIPASIGNLKWLTVFEVA 366
+L NL L LG NN SG PIP IG L NL L++Q++NL G+IP IG L L +++
Sbjct: 1 NLTNLSYLILGGNNWSGGPIPPEIGKLNNLLHLAIQKSNLVGSIPQEIGFLTNLAYIDLS 180
Query: 367 TNKLHGRIPNGLYNITNWISFVVSENDFV-GHLPSQICSGGSLRLLNADHNRFTGPIPTS 425
N L G IP + N++ + V+S N + G +P + + SL +L D+ +G IP S
Sbjct: 181 KNSLSGGIPETIGNLSKLDTLVLSNNTKMSGPIPHSLWNMSSLTVLYFDNIGLSGSIPDS 360
Query: 426 LKTCSSIERITLEVNQIEGDIAQDFGVYPKLQYLDLSDNKFHGQISPNWGKSLNLQTFII 485
++ +++ + L++N + G I G L L L N G I + G +NLQ +
Sbjct: 361 IQNLVNLKELALDINHLSGSIPSTIGDLKNLIKLYLGSNNLSGPIPASIGNLINLQVLSV 540
Query: 486 SNNNISGVIPLDFIGLTKLGVLHLSSNQLTGKLP 519
NN++G IP L L V + +N+ + ++P
Sbjct: 541 QENNLTGTIPASIGNLKWLTVFEVXTNKPSWRIP 642
Score = 107 bits (267), Expect = 2e-23
Identities = 68/210 (32%), Positives = 108/210 (51%), Gaps = 3/210 (1%)
Frame = +1
Query: 410 LLNADHNRFTGPIPTSLKTCSSIERITLEVNQIEGDIAQDFGVYPKLQYLDLSDNKFHGQ 469
L+ +N GPIP + +++ + ++ + + G I Q+ G L Y+DLS N G
Sbjct: 22 LILGGNNWSGGPIPPEIGKLNNLLHLAIQKSNLVGSIPQEIGFLTNLAYIDLSKNSLSGG 201
Query: 470 ISPNWGKSLNLQTFIISNNN-ISGVIPLDFIGLTKLGVLHLSSNQLTGKLPMEVLGGMKS 528
I G L T ++SNN +SG IP ++ L VL+ + L+G +P + + + +
Sbjct: 202 IPETIGNLSKLDTLVLSNNTKMSGPIPHSLWNMSSLTVLYFDNIGLSGSIP-DSIQNLVN 378
Query: 529 LFDLKISNNHFSDNIPSEIGLLQRLQELDLGGNELSGKIPKELVELPNLRMLNLSRNKIE 588
L +L + NH S +IPS IG L+ L +L LG N LSG IP + L NL++L++ N +
Sbjct: 379 LKELALDINHLSGSIPSTIGDLKNLIKLYLGSNNLSGPIPASIGNLINLQVLSVQENNLT 558
Query: 589 GIIPIKFDS--GLESLDLSGNFLKGNIPTG 616
G IP + L ++ N IP G
Sbjct: 559 GTIPASIGNLKWLTVFEVXTNKPSWRIPNG 648
Score = 90.5 bits (223), Expect = 3e-18
Identities = 71/190 (37%), Positives = 97/190 (50%), Gaps = 7/190 (3%)
Frame = +1
Query: 501 LTKLGVLHLSSNQLTG-KLPMEVLGGMKSLFDLKISNNHFSDNIPSEIGLLQRLQELDLG 559
LT L L L N +G +P E+ G + +L L I ++ +IP EIG L L +DL
Sbjct: 4 LTNLSYLILGGNNWSGGPIPPEI-GKLNNLLHLAIQKSNLVGSIPQEIGFLTNLAYIDLS 180
Query: 560 GNELSGKIPKELVELPNLRMLNLSRN-KIEGIIPIKF--DSGLESLDLSGNFLKGNIPTG 616
N LSG IP+ + L L L LS N K+ G IP S L L L G+IP
Sbjct: 181 KNSLSGGIPETIGNLSKLDTLVLSNNTKMSGPIPHSLWNMSSLTVLYFDNIGLSGSIPDS 360
Query: 617 LADLVRLSKLNLSHNMLSGTIPQNFG--RNLVFVNISDNQLEGPLP-KIPAFLSASFESL 673
+ +LV L +L L N LSG+IP G +NL+ + + N L GP+P I ++ S+
Sbjct: 361 IQNLVNLKELALDINHLSGSIPSTIGDLKNLIKLYLGSNNLSGPIPASIGNLINLQVLSV 540
Query: 674 KNNNHLCGNI 683
+ NN L G I
Sbjct: 541 QENN-LTGTI 567
Score = 60.5 bits (145), Expect = 3e-09
Identities = 49/147 (33%), Positives = 63/147 (42%)
Frame = +1
Query: 84 HSLTFSSFPNLLMIDIRNNSFYGTIPAQIGNLSNISILTFKNNYFDGSIPQEMCTLTGLQ 143
HSL S +L D N G+IP I NL N+ L N+ GSIP + L L
Sbjct: 283 HSLWNMSSLTVLYFD--NIGLSGSIPDSIQNLVNLKELALDINHLSGSIPSTIGDLKNLI 456
Query: 144 FLDISFCKLNGAIPKSIGNLTNLSYLILGGNNWSGGPIPPEIGKLNNLLHLAIQKSNLVG 203
L + L+G IP SIGNL NL L++Q++NL G
Sbjct: 457 KLYLGSNNLSGPIPASIGNLINLQV-------------------------LSVQENNLTG 561
Query: 204 SIPQEIGFLTNLAYIDLSKNSLSGGIP 230
+IP IG L L ++ N S IP
Sbjct: 562 TIPASIGNLKWLTVFEVXTNKPSWRIP 642
Score = 46.6 bits (109), Expect = 5e-05
Identities = 26/79 (32%), Positives = 42/79 (52%)
Frame = +1
Query: 79 LKGTLHSLTFSSFPNLLMIDIRNNSFYGTIPAQIGNLSNISILTFKNNYFDGSIPQEMCT 138
L G++ S T NL+ + + +N+ G IPA IGNL N+ +L+ + N G+IP +
Sbjct: 409 LSGSIPS-TIGDLKNLIKLYLGSNNLSGPIPASIGNLINLQVLSVQENNLTGTIPASIGN 585
Query: 139 LTGLQFLDISFCKLNGAIP 157
L L ++ K + IP
Sbjct: 586 LKWLTVFEVXTNKPSWRIP 642
>TC82239 weakly similar to GP|16924042|gb|AAL31654.1 Putative protein kinase
{Oryza sativa}, partial (14%)
Length = 1476
Score = 360 bits (924), Expect = 2e-99
Identities = 196/438 (44%), Positives = 278/438 (62%), Gaps = 7/438 (1%)
Frame = +2
Query: 609 LKGNIPTGLADLVRLSKLNLSHNMLSGTIPQNFGRNLVFVNISDNQLEGPLPKIPAFLSA 668
L G +P ++ ++ LS LNLS+N L G +P++ +F S + L+
Sbjct: 5 LSGKVPNQISGMLSLSSLNLSYNHLEGNVPKSG----IFKLNSSHALD------------ 136
Query: 669 SFESLKNNNHLCGNIRGLDPCATSHSR------KRKNVLRPVFIALG-AVILVLCVVGAL 721
L NN LCG+ +GL PC S S +K V+ P+ +LG A+ L L +VG +
Sbjct: 137 ----LSNNQDLCGSFKGLIPCNVSSSEPSDGGSNKKKVVIPIVASLGGALFLSLVIVGVI 304
Query: 722 MYIMCGRKKPNEESQTEEVQRGVLFSIWSHDGKMMFENIIEATANFDDKYLVGVGSQGNV 781
+ +C +KK + + + FSIW +G++++ +IIEAT NFD+KY +G G+ GNV
Sbjct: 305 L--LCYKKK-SRTLRKSSFKMPNPFSIWYFNGRVVYSDIIEATNNFDNKYCIGEGAFGNV 475
Query: 782 YKAELSEGLVVAVKKLHLVTDEEMSCFSSKSFMSEIETLTGIKHRNIIKLHGFCSHSKFS 841
YKAEL G + AVKKL +E + S K+F SE+E +T +HRNI+KL+GFC +
Sbjct: 476 YKAELKGGQIFAVKKLKC-DEENLDTESIKTFESEVEAMTETRHRNIVKLYGFCCEGMHT 652
Query: 842 FLVYKFLEGGSLDQILNNDTQAVAFDWEKRVNVVKGVANALSYLHHDCSPPIIHRDISSK 901
FLVY++++ GSL+ +L +D +A+ DW KR +VKGVA+ALSY+HHDCSP +IHRDISSK
Sbjct: 653 FLVYEYMDRGSLEDMLIDDKRALELDWSKRFEIVKGVASALSYMHHDCSPALIHRDISSK 832
Query: 902 NVLLNLDYEAHVSDFGTAKFLKPGLHSWTQFAGTFGYAAPELAQTMEVNEKCDVYSFGVL 961
NVLL+ + EAHVSDFGTA+FLKP WT FAGT+GYAAPELA TM V EKCDV+SFGVL
Sbjct: 833 NVLLSKNLEAHVSDFGTARFLKPNSPIWTSFAGTYGYAAPELAYTMAVTEKCDVFSFGVL 1012
Query: 962 ALETIMGKHPGDLISLFLSPSTRPMANNMLLTDVLDQRPQQVMEPIDEEVILIARLAFAC 1021
A E + GKHP DL+S + + + + +L L P+ ++ +E+ L+A LA +C
Sbjct: 1013 AFEILTGKHPSDLVSYIQTSNDQKIDFKEILDPRLPSPPKNIL----KELALVANLALSC 1180
Query: 1022 LSQNPRLRPSMGQVCKML 1039
L +P+ RP+M V + L
Sbjct: 1181 LHTHPQSRPTMRSVAQFL 1234
Score = 36.2 bits (82), Expect = 0.069
Identities = 22/65 (33%), Positives = 35/65 (53%), Gaps = 6/65 (9%)
Frame = +2
Query: 274 LSGSIPDSIQNLVNLKELALDINHLSGSIP-STIGDLKNLIKLYLGSN-----NLSGPIP 327
LSG +P+ I +++L L L NHL G++P S I L + L L +N + G IP
Sbjct: 5 LSGKVPNQISGMLSLSSLNLSYNHLEGNVPKSGIFKLNSSHALDLSNNQDLCGSFKGLIP 184
Query: 328 ASIGN 332
++ +
Sbjct: 185 CNVSS 199
Score = 34.7 bits (78), Expect = 0.20
Identities = 21/50 (42%), Positives = 28/50 (56%), Gaps = 1/50 (2%)
Frame = +2
Query: 514 LTGKLPMEVLGGMKSLFDLKISNNHFSDNIP-SEIGLLQRLQELDLGGNE 562
L+GK+P ++ GM SL L +S NH N+P S I L LDL N+
Sbjct: 5 LSGKVPNQI-SGMLSLSSLNLSYNHLEGNVPKSGIFKLNSSHALDLSNNQ 151
>TC90203 weakly similar to PIR|B86479|B86479 hypothetical protein AAF79881.1
[imported] - Arabidopsis thaliana, partial (12%)
Length = 1303
Score = 306 bits (784), Expect(2) = 4e-84
Identities = 177/403 (43%), Positives = 253/403 (61%), Gaps = 5/403 (1%)
Frame = +3
Query: 602 LDLSGNFLKGNIPTGLADLVRLSKLNLSHNMLSGTIPQNFGRNLVFVNISDNQLEGPLPK 661
+DLS N + G IP+ L LS LNL +N L+G PQ+ N+ +V+IS N L+GPLP
Sbjct: 57 VDLSYNLIGGEIPSQLR---YLSILNLRNNNLTGVFPQSLC-NVNYVDISFNHLKGPLPN 224
Query: 662 IPAFLSASFESLKNNNHLCGNIRGLDPCATSHSRKRKNVLRPVFIALGAVILVLCVVGAL 721
+ + ++ N+ DP + S N+ V I L +++++ A
Sbjct: 225 C---IHNGYNTIIWND---------DPYINNRSN---NINYDVVIVLPILLILIL---AF 350
Query: 722 MYIMCGRKKPNEE-----SQTEEVQRGVLFSIWSHDGKMMFENIIEATANFDDKYLVGVG 776
++C + + N + T + G LF IW+ DGK+ ++II+AT +FD +Y +G G
Sbjct: 351 SLLICFKLRQNSTKIKLANTTISTKNGDLFCIWNFDGKIAHDDIIKATEDFDIRYCIGTG 530
Query: 777 SQGNVYKAELSEGLVVAVKKLHLVTDEEMSCFSSKSFMSEIETLTGIKHRNIIKLHGFCS 836
+ G+VYKA+L G VVA+KKLH + E+ F SF +E+ L+ IKHR+I+KL+GFC
Sbjct: 531 AYGSVYKAQLPCGKVVAIKKLHGY-EAEVPSFDD-SFRNEVRILSEIKHRHIVKLYGFCL 704
Query: 837 HSKFSFLVYKFLEGGSLDQILNNDTQAVAFDWEKRVNVVKGVANALSYLHHDCSPPIIHR 896
H + FL+Y+++E GSL +L +D + V F W KRVN +KGVA A SYLHHDC+ PI+HR
Sbjct: 705 HKRIMFLIYQYMEKGSLFSVLYDDVKVVEFKWRKRVNTIKGVAFAFSYLHHDCTAPIVHR 884
Query: 897 DISSKNVLLNLDYEAHVSDFGTAKFLKPGLHSWTQFAGTFGYAAPELAQTMEVNEKCDVY 956
D+S+ N+LLN +++A V DFG A+ L+ + T AGT GY APELA TM VNEKCDVY
Sbjct: 885 DVSTSNILLNSEWQASVCDFGIARLLQYDSSNRTIVAGTIGYIAPELAYTMAVNEKCDVY 1064
Query: 957 SFGVLALETIMGKHPGDLISLFLSPSTRPMANNMLLTDVLDQR 999
SFGV+ALET++G+HPGDL+S S ST+ ++ L VLD R
Sbjct: 1065SFGVVALETLVGRHPGDLLSSLQSTSTQ----SLKLCQVLDHR 1181
Score = 43.9 bits (102), Expect = 3e-04
Identities = 25/66 (37%), Positives = 35/66 (52%)
Frame = +3
Query: 97 IDIRNNSFYGTIPAQIGNLSNISILTFKNNYFDGSIPQEMCTLTGLQFLDISFCKLNGAI 156
+D+ N G IP+Q L +SIL +NN G PQ +C + ++DISF L G +
Sbjct: 57 VDLSYNLIGGEIPSQ---LRYLSILNLRNNNLTGVFPQSLC---NVNYVDISFNHLKGPL 218
Query: 157 PKSIGN 162
P I N
Sbjct: 219 PNCIHN 236
Score = 25.0 bits (53), Expect(2) = 4e-84
Identities = 11/25 (44%), Positives = 17/25 (68%)
Frame = +1
Query: 1002 QVMEPIDEEVILIARLAFACLSQNP 1026
Q+M + ++I A +AFACL+ NP
Sbjct: 1192 QIMT*LYRDIIHAAVVAFACLNVNP 1266
>TC89364 weakly similar to PIR|C96745|C96745 hypothetical protein T9N14.3
[imported] - Arabidopsis thaliana, partial (45%)
Length = 1903
Score = 283 bits (723), Expect = 3e-76
Identities = 184/555 (33%), Positives = 284/555 (51%), Gaps = 21/555 (3%)
Frame = +1
Query: 506 VLHLSSNQLTGKLPMEVLGGMKSLFDLKISNNHFSDNIPSEIGLLQRLQELDLGGNELSG 565
++ L N +G++ E+ G +L ++ + NN FS +PSEIG L L++L L N SG
Sbjct: 76 IIDLGFNNFSGEVSSEI-GYSTNLSEIVLMNNKFSGKVPSEIGKLVNLEKLYLSNNNFSG 252
Query: 566 KIPKELVELPNLRMLNLSRNKIEGIIPIKFD--SGLESLDLSGNFLKGNIPTGLADLVRL 623
IP+E+ L L L+L N + G+IP + S L L+L+ N L GNIP ++ + L
Sbjct: 253 DIPREIGLLKQLSTLHLEENSLTGVIPKELGHCSRLVDLNLALNSLSGNIPNSVSLMSSL 432
Query: 624 SKLNLSHNMLSGTIPQNFGR-NLVFVNISDNQLEGPLPKIPAFLSASFESLKNNNHLC-- 680
+ LNLS N L+GTIP N + L V+ S N L G +P + ++ N LC
Sbjct: 433 NSLNLSRNKLTGTIPDNLEKMKLSSVDFSQNSLSGGIP-FGILIIGGEKAFVGNKELCVE 609
Query: 681 -----GNIRGLDPCATSHSRKRKNVLRPVFIALGAVILVLCVVGALMYIMCGRKKPNEES 735
L C H +R V + L + ++ + M RK+ N +
Sbjct: 610 QIPKTSMNSDLKICDKDHGHRR-GVFAYKYFLLFFIAVIFAAAIVIHRCMKNRKEKNLQK 786
Query: 736 QTEEVQRGVLFSIWSHDGKMMFENIIEATANFDDKYLVGVGSQGNVYKAELSE-GLVVAV 794
+E + W + + ++ D L+G G G VY+ +L + G+VVAV
Sbjct: 787 GEKEASQK-----WKQASFHQVDIDADEVSHLGDDNLIGYGGTGKVYRVKLKKTGIVVAV 951
Query: 795 KKLHLVTDEEMSCFSSKSFMSEIETLTGIKHRNIIKLHGFCSHSKFSFLVYKFLEGGSLD 854
K+L + K +E+E L I+H+NI+KL+ + LV++++ G+L
Sbjct: 952 KQLE-------KGYGVKILAAEMEILAKIRHKNILKLYACFLKGGSNLLVFEYMPNGNLF 1110
Query: 855 QILNNDT--QAVAFDWEKRVNVVKGVANALSYLHHDCSPPIIHRDISSKNVLLNLDYEAH 912
Q L+ + + V FDW +R + G A + YLHHDCSPP+IHRDI S N+LL+ +YEA
Sbjct: 1111 QALHREVKDEMVTFDWNQRYKIALGGAKGICYLHHDCSPPVIHRDIKSSNILLDANYEAK 1290
Query: 913 VSDFGTAKFLKPGLHSWTQFAGTFGYAAPELAQTMEVNEKCDVYSFGVLALETIMGKHP- 971
++DFG A+F + ++ FAGT GY APELA T E+ EK DVYSFGV+ LE + G+ P
Sbjct: 1291 IADFGVARFAEKSQMGYSVFAGTHGYIAPELAYTTEITEKSDVYSFGVVLLELVSGREPV 1470
Query: 972 -------GDLISLFLSPSTRPMANNMLLTDVLDQRPQQVMEPIDEEVILIARLAFACLSQ 1024
D++ +S + + + +LD R E++I + ++ C ++
Sbjct: 1471 EEEYGEAKDIVHWVMS----NLNDRESVLSILDGRVAST--HCVEDMIKVLKIGIKCTTK 1632
Query: 1025 NPRLRPSMGQVCKML 1039
P LRP+M V KML
Sbjct: 1633 LPTLRPTMRDVVKML 1677
Score = 112 bits (279), Expect = 1e-24
Identities = 74/209 (35%), Positives = 107/209 (50%)
Frame = +1
Query: 212 LTNLAYIDLSKNSLSGGIPETIGNLSKLDTLVLSNNTKMSGPIPHSLWNMSSLTVLYFDN 271
L N IDL N+ SG + IG + L +VL NN K SG +P + + +L LY N
Sbjct: 61 LPNAKIIDLGFNNFSGEVSSEIGYSTNLSEIVLMNN-KFSGKVPSEIGKLVNLEKLYLSN 237
Query: 272 IGLSGSIPDSIQNLVNLKELALDINHLSGSIPSTIGDLKNLIKLYLGSNNLSGPIPASIG 331
SG IP I L L L L+ N L+G IP +G L+ L L N+LSG IP S+
Sbjct: 238 NNFSGDIPREIGLLKQLSTLHLEENSLTGVIPKELGHCSRLVDLNLALNSLSGNIPNSVS 417
Query: 332 NLINLQVLSVQENNLTGTIPASIGNLKWLTVFEVATNKLHGRIPNGLYNITNWISFVVSE 391
+ +L L++ N LTGTIP ++ +K L+ + + N L G IP G+ I +FV ++
Sbjct: 418 LMSSLNSLNLSRNKLTGTIPDNLEKMK-LSSVDFSQNSLSGGIPFGILIIGGEKAFVGNK 594
Query: 392 NDFVGHLPSQICSGGSLRLLNADHNRFTG 420
V +P + L++ + DH G
Sbjct: 595 ELCVEQIP-KTSMNSDLKICDKDHGHRRG 678
Score = 110 bits (276), Expect = 2e-24
Identities = 67/166 (40%), Positives = 100/166 (59%)
Frame = +1
Query: 138 TLTGLQFLDISFCKLNGAIPKSIGNLTNLSYLILGGNNWSGGPIPPEIGKLNNLLHLAIQ 197
+L + +D+ F +G + IG TNLS ++L N +SG +P EIGKL NL L +
Sbjct: 58 SLPNAKIIDLGFNNFSGEVSSEIGYSTNLSEIVLMNNKFSG-KVPSEIGKLVNLEKLYLS 234
Query: 198 KSNLVGSIPQEIGFLTNLAYIDLSKNSLSGGIPETIGNLSKLDTLVLSNNTKMSGPIPHS 257
+N G IP+EIG L L+ + L +NSL+G IP+ +G+ S+L L L+ N+ +SG IP+S
Sbjct: 235 NNNFSGDIPREIGLLKQLSTLHLEENSLTGVIPKELGHCSRLVDLNLALNS-LSGNIPNS 411
Query: 258 LWNMSSLTVLYFDNIGLSGSIPDSIQNLVNLKELALDINHLSGSIP 303
+ MSSL L L+G+IPD+++ + L + N LSG IP
Sbjct: 412 VSLMSSLNSLNLSRNKLTGTIPDNLEKM-KLSSVDFSQNSLSGGIP 546
Score = 110 bits (274), Expect = 4e-24
Identities = 72/193 (37%), Positives = 104/193 (53%)
Frame = +1
Query: 90 SFPNLLMIDIRNNSFYGTIPAQIGNLSNISILTFKNNYFDGSIPQEMCTLTGLQFLDISF 149
S PN +ID+ N+F G + ++IG +N+S + NN F G +P E
Sbjct: 58 SLPNAKIIDLGFNNFSGEVSSEIGYSTNLSEIVLMNNKFSGKVPSE-------------- 195
Query: 150 CKLNGAIPKSIGNLTNLSYLILGGNNWSGGPIPPEIGKLNNLLHLAIQKSNLVGSIPQEI 209
IG L NL L L NN+S G IP EIG L L L +++++L G IP+E+
Sbjct: 196 ----------IGKLVNLEKLYLSNNNFS-GDIPREIGLLKQLSTLHLEENSLTGVIPKEL 342
Query: 210 GFLTNLAYIDLSKNSLSGGIPETIGNLSKLDTLVLSNNTKMSGPIPHSLWNMSSLTVLYF 269
G + L ++L+ NSLSG IP ++ +S L++L LS N K++G IP +L M L+ + F
Sbjct: 343 GHCSRLVDLNLALNSLSGNIPNSVSLMSSLNSLNLSRN-KLTGTIPDNLEKM-KLSSVDF 516
Query: 270 DNIGLSGSIPDSI 282
LSG IP I
Sbjct: 517 SQNSLSGGIPFGI 555
Score = 102 bits (253), Expect = 1e-21
Identities = 59/166 (35%), Positives = 85/166 (50%)
Frame = +1
Query: 258 LWNMSSLTVLYFDNIGLSGSIPDSIQNLVNLKELALDINHLSGSIPSTIGDLKNLIKLYL 317
+W++ + ++ SG + I NL E+ L N SG +PS IG L NL KLYL
Sbjct: 52 VWSLPNAKIIDLGFNNFSGEVSSEIGYSTNLSEIVLMNNKFSGKVPSEIGKLVNLEKLYL 231
Query: 318 GSNNLSGPIPASIGNLINLQVLSVQENNLTGTIPASIGNLKWLTVFEVATNKLHGRIPNG 377
+NN SG IP IG L L L ++EN+LTG IP +G+ L +A N L G IPN
Sbjct: 232 SNNNFSGDIPREIGLLKQLSTLHLEENSLTGVIPKELGHCSRLVDLNLALNSLSGNIPNS 411
Query: 378 LYNITNWISFVVSENDFVGHLPSQICSGGSLRLLNADHNRFTGPIP 423
+ +++ S +S N G +P + L ++ N +G IP
Sbjct: 412 VSLMSSLNSLNLSRNKLTGTIPDNL-EKMKLSSVDFSQNSLSGGIP 546
Score = 76.3 bits (186), Expect = 6e-14
Identities = 61/220 (27%), Positives = 109/220 (48%), Gaps = 1/220 (0%)
Frame = +1
Query: 377 GLYNITNWISFVVSENDFVGHLPSQICSGGSLRLLNADHNRFTGPIPTSLKTCSSIERIT 436
G++++ N + N+F G + S+I +L + +N+F+G +P+ + ++E++
Sbjct: 49 GVWSLPNAKIIDLGFNNFSGEVSSEIGYSTNLSEIVLMNNKFSGKVPSEIGKLVNLEKLY 228
Query: 437 LEVNQIEGDIAQDFGVYPKLQYLDLSDNKFHGQISPNWGKSLNLQTFIISNNNISGVIPL 496
L N GDI ++ G+ +L L L +N G I G L ++ N++SG IP
Sbjct: 229 LSNNNFSGDIPREIGLLKQLSTLHLEENSLTGVIPKELGHCSRLVDLNLALNSLSGNIPN 408
Query: 497 DFIGLTKLGVLHLSSNQLTGKLPMEVLGGMKSLFDLKISNNHFSDNIPSEIGLLQRLQEL 556
++ L L+LS N+LTG +P + L MK L + S N S IP I L+ ++
Sbjct: 409 SVSLMSSLNSLNLSRNKLTGTIP-DNLEKMK-LSSVDFSQNSLSGGIPFGI-LIIGGEKA 579
Query: 557 DLGGNELS-GKIPKELVELPNLRMLNLSRNKIEGIIPIKF 595
+G EL +IPK + +L++ + G+ K+
Sbjct: 580 FVGNKELCVEQIPKTSMN-SDLKICDKDHGHRRGVFAYKY 696
Score = 72.8 bits (177), Expect = 7e-13
Identities = 48/169 (28%), Positives = 81/169 (47%), Gaps = 2/169 (1%)
Frame = +1
Query: 362 VFEVATNKLHGRIPNGLYNITNWISFVVSENDFVGHLPSQICSGGSLRLLNADHNRFTGP 421
+ ++ N G + + + TN V+ N F G +PS+I +L L +N F+G
Sbjct: 76 IIDLGFNNFSGEVSSEIGYSTNLSEIVLMNNKFSGKVPSEIGKLVNLEKLYLSNNNFSGD 255
Query: 422 IPTSLKTCSSIERITLEVNQIEGDIAQDFGVYPKLQYLDLSDNKFHGQISPNWGKSLNLQ 481
IP + + + LE N + G I ++ G +L L+L+ N G I + +L
Sbjct: 256 IPREIGLLKQLSTLHLEENSLTGVIPKELGHCSRLVDLNLALNSLSGNIPNSVSLMSSLN 435
Query: 482 TFIISNNNISGVIPLDFIGLTKLGVLHLSSNQLTGKLPMEVL--GGMKS 528
+ +S N ++G IP D + KL + S N L+G +P +L GG K+
Sbjct: 436 SLNLSRNKLTGTIP-DNLEKMKLSSVDFSQNSLSGGIPFGILIIGGEKA 579
>TC89003 weakly similar to GP|18086327|gb|AAL57627.1 AT4g08850/T32A17_160
{Arabidopsis thaliana}, partial (6%)
Length = 688
Score = 276 bits (705), Expect = 4e-74
Identities = 144/230 (62%), Positives = 174/230 (75%), Gaps = 3/230 (1%)
Frame = +3
Query: 653 NQLEGPLPKIPAFLSASFESLKNNNHLCGNIRGLDPCATSHSRKRKNVLRPV-FIALGAV 711
NQLEG LP AFL A ESLKNN LCGN GL C TSHS+KR +L V F+ LGA+
Sbjct: 3 NQLEGRLPNNQAFLKAPIESLKNNKGLCGNHTGLMLCPTSHSKKRHEILLLVLFVILGAL 182
Query: 712 ILVLCVVGALMYIMC--GRKKPNEESQTEEVQRGVLFSIWSHDGKMMFENIIEATANFDD 769
+LV +G MYI+ RK N++ + E Q +FSIWSHDGKMMFENIIEAT NFDD
Sbjct: 183 VLVFSGLGISMYIIYRRARKTKNKDKDSNEAQAEEVFSIWSHDGKMMFENIIEATNNFDD 362
Query: 770 KYLVGVGSQGNVYKAELSEGLVVAVKKLHLVTDEEMSCFSSKSFMSEIETLTGIKHRNII 829
+YL+GVG +G+VYKA+LS +VVAVKKLH D E S + K+F +EI+ LT I+HRNII
Sbjct: 363 EYLIGVGGEGSVYKAKLSADMVVAVKKLHSRIDGERS--NIKAFENEIQALTEIRHRNII 536
Query: 830 KLHGFCSHSKFSFLVYKFLEGGSLDQILNNDTQAVAFDWEKRVNVVKGVA 879
KL+G+C HS+FSFLVYKFLEGG+L Q+LNNDTQA+AFDWEKRVN+ +GVA
Sbjct: 537 KLYGYCRHSRFSFLVYKFLEGGTLTQMLNNDTQAIAFDWEKRVNIXRGVA 686
>TC91405 weakly similar to GP|16924042|gb|AAL31654.1 Putative protein kinase
{Oryza sativa}, partial (16%)
Length = 1000
Score = 275 bits (703), Expect = 7e-74
Identities = 135/222 (60%), Positives = 168/222 (74%), Gaps = 4/222 (1%)
Frame = +1
Query: 824 KHRNIIKLHGFCSHSKFSFLVYKFLEGGSLDQILNNDTQAVAFDWEKRVNVVKGVANALS 883
+HRNI+KL+GFCSH SFLVY+FLE GSLD IL ++ QA FDW KRVN++K +ANAL
Sbjct: 193 RHRNIVKLYGFCSHQLHSFLVYEFLEKGSLDNILKDNEQAGEFDWNKRVNIIKDIANALF 372
Query: 884 YLHHDCSPPIIHRDISSKNVLLNLDYEAHVSDFGTAKFLKPGLHSWTQFAGTFGYAAPEL 943
YLHHDCSPPI+HRDISSKNV+L+L+Y AHVSDFGT+KFL P + T FAGTFGYAAPEL
Sbjct: 373 YLHHDCSPPIVHRDISSKNVILDLEYVAHVSDFGTSKFLNPNSSNMTSFAGTFGYAAPEL 552
Query: 944 AQTMEVNEKCDVYSFGVLALETIMGKHPGDLIS-LFLSPS---TRPMANNMLLTDVLDQR 999
A TMEVN+KCDVYSFG+L LE + GKHPGD+++ L+ PS T + M L D LDQR
Sbjct: 553 AYTMEVNKKCDVYSFGILTLEILFGKHPGDIVTYLWQQPSQSVTDLRLDTMPLIDKLDQR 732
Query: 1000 PQQVMEPIDEEVILIARLAFACLSQNPRLRPSMGQVCKMLAI 1041
+ I +EV + R+A ACL+++P RP+M QVC+ +
Sbjct: 733 LPHPTKTIVQEVASMIRIAVACLTESPLSRPTMEQVCRQFVM 858
Score = 77.4 bits (189), Expect = 3e-14
Identities = 43/76 (56%), Positives = 55/76 (71%), Gaps = 3/76 (3%)
Frame = +3
Query: 759 NIIEATANFDDKYLVGVGSQGNVYKAELSEGLVVAVKKLHLVTDEEMSCFSSKSFMSEIE 818
NIIEAT +FD+K+L+GVG GNVYKAEL G VVAVKKLH + +EEM + K+F +EI
Sbjct: 3 NIIEATEDFDNKHLIGVGGHGNVYKAELPTGQVVAVKKLHSLQNEEMP--NRKAFTNEIH 176
Query: 819 TLTGI---KHRNIIKL 831
LT I +H +I+L
Sbjct: 177 ALTEIPTSQHCEVIRL 224
>TC87042 similar to GP|13605827|gb|AAK32899.1 AT5g48380/MJE7_1 {Arabidopsis
thaliana}, partial (51%)
Length = 2206
Score = 241 bits (616), Expect = 8e-64
Identities = 173/521 (33%), Positives = 262/521 (50%), Gaps = 30/521 (5%)
Frame = +2
Query: 552 RLQELDLGGNELSGKIPKELVELPNLRMLNLSRNKIEGIIPIKFDSGLE---SLDLSGNF 608
++ L L L G+ P+ +V ++ L+LS N + G IP + L+ SLDLS N
Sbjct: 383 KVLNLKLSNMGLKGQFPRGIVNCSSMTGLDLSVNDLSGTIPGDISTLLKFVTSLDLSSNE 562
Query: 609 LKGNIPTGLADLVRLSKLNLSHNMLSGTIPQNFGR--NLVFVNISDNQLEGPLPKIPAFL 666
G IP LA+ L+ L LS N L+G IP G + ++S+N L G +P A
Sbjct: 563 FSGEIPVSLANCTYLNVLKLSQNQLTGQIPLLLGTLDRIKTFDVSNNLLTGQVPNFTAGG 742
Query: 667 SASFESLKNNNHLCGNIRGLDPC-ATSHSRKRKNVLRPVFIALGAVILVLCVVGALMYIM 725
+ NN LCG L C AT+ S+ V+ A+GAV L +G M+
Sbjct: 743 KVDV-NYANNQGLCGQ-PSLGVCKATASSKSNTAVIAGA--AVGAVTLAALGLGVFMFFF 910
Query: 726 CGR---KKPNEESQTEEVQR------GVLFSIWSHD-GKMMFENIIEATANFDDKYLVGV 775
R +K E+ + + R G+ S++ KM ++++AT NF + ++G
Sbjct: 911 VRRSAYRKKEEDPEGNKWARSLKGTKGIKVSLFEKSISKMKLSDLMKATNNFSNINIIGT 1090
Query: 776 GSQGNVYKAELSEGLVVAVKKLHLVTDEEMSCFSSKSFMSEIETLTGIKHRNIIKLHGFC 835
G G VYKA L +G VK+L + S S K FMSE+ TL +KHRN++ L GFC
Sbjct: 1091 GRTGTVYKATLEDGTAFMVKRL------QESQHSEKEFMSEMATLGTVKHRNLVPLLGFC 1252
Query: 836 SHSKFSFLVYKFLEGGSLDQILNNDTQAVAFDWEKRVNVVKGVANALSYLHHDCSPPIIH 895
K LV+K + G L L+ DW R+ + G A ++LHH C+P IIH
Sbjct: 1253 VAKKERLLVFKNMPNGMLHDQLHPAAGECTLDWPSRLKIAIGAAKGFAWLHHSCNPRIIH 1432
Query: 896 RDISSKNVLLNLDYEAHVSDFGTAKFLKPGLHSWTQFA----GTFGYAAPELAQTMEVNE 951
R+ISSK +LL+ D+E +SDFG A+ + P + F G FGY APE +T+
Sbjct: 1433 RNISSKCILLDADFEPKISDFGLARLMNPLDTHLSTFVNGEFGDFGYVAPEYTKTLVATP 1612
Query: 952 KCDVYSFGVLALETIMGKHPGDLISLFLSPST---------RPMANNMLLTDVLDQRPQQ 1002
K DV+SFG + LE + G+ P ++ +P T +++N L D +D+
Sbjct: 1613 KGDVFSFGTVLLELVTGERPANVAK---APETFKGNLVEWITELSSNSKLHDAIDE--SL 1777
Query: 1003 VMEPIDEEVILIARLAFACLSQNPRLRPSMGQVCKML-AIG 1042
+ + D E+ ++A C+++ P+ RP+M +V + L AIG
Sbjct: 1778 LNKGDDNELFQFLKVACNCVTEVPKERPTMFEVYQFLRAIG 1900
Score = 84.7 bits (208), Expect = 2e-16
Identities = 47/111 (42%), Positives = 62/111 (55%), Gaps = 1/111 (0%)
Frame = +2
Query: 267 LYFDNIGLSGSIPDSIQNLVNLKELALDINHLSGSIPSTIGDL-KNLIKLYLGSNNLSGP 325
L N+GL G P I N ++ L L +N LSG+IP I L K + L L SN SG
Sbjct: 395 LKLSNMGLKGQFPRGIVNCSSMTGLDLSVNDLSGTIPGDISTLLKFVTSLDLSSNEFSGE 574
Query: 326 IPASIGNLINLQVLSVQENNLTGTIPASIGNLKWLTVFEVATNKLHGRIPN 376
IP S+ N L VL + +N LTG IP +G L + F+V+ N L G++PN
Sbjct: 575 IPVSLANCTYLNVLKLSQNQLTGQIPLLLGTLDRIKTFDVSNNLLTGQVPN 727
Score = 63.2 bits (152), Expect = 5e-10
Identities = 40/116 (34%), Positives = 60/116 (51%)
Frame = +2
Query: 189 NNLLHLAIQKSNLVGSIPQEIGFLTNLAYIDLSKNSLSGGIPETIGNLSKLDTLVLSNNT 248
N +L+L + L G P+ I +++ +DLS N LSG IP I L K T + ++
Sbjct: 380 NKVLNLKLSNMGLKGQFPRGIVNCSSMTGLDLSVNDLSGTIPGDISTLLKFVTSLDLSSN 559
Query: 249 KMSGPIPHSLWNMSSLTVLYFDNIGLSGSIPDSIQNLVNLKELALDINHLSGSIPS 304
+ SG IP SL N + L VL L+G IP + L +K + N L+G +P+
Sbjct: 560 EFSGEIPVSLANCTYLNVLKLSQNQLTGQIPLLLGTLDRIKTFDVSNNLLTGQVPN 727
Score = 62.4 bits (150), Expect = 9e-10
Identities = 45/163 (27%), Positives = 71/163 (42%), Gaps = 11/163 (6%)
Frame = +2
Query: 386 SFVVSENDFVGHLPSQICSGGSLRLLNADHNR----------FTGPIPTSLKTCSSIERI 435
+++ S +F IC + + D N+ G P + CSS+ +
Sbjct: 290 NYLTSSWNFNNKTEGFICRFNGVECWHPDENKVLNLKLSNMGLKGQFPRGIVNCSSMTGL 469
Query: 436 TLEVNQIEGDIAQDFGVYPK-LQYLDLSDNKFHGQISPNWGKSLNLQTFIISNNNISGVI 494
L VN + G I D K + LDLS N+F G+I + L +S N ++G I
Sbjct: 470 DLSVNDLSGTIPGDISTLLKFVTSLDLSSNEFSGEIPVSLANCTYLNVLKLSQNQLTGQI 649
Query: 495 PLDFIGLTKLGVLHLSSNQLTGKLPMEVLGGMKSLFDLKISNN 537
PL L ++ +S+N LTG++P GG D+ +NN
Sbjct: 650 PLLLGTLDRIKTFDVSNNLLTGQVPNFTAGGK---VDVNYANN 769
>BE319483 weakly similar to GP|13489172|gb putative protein kinase {Oryza
sativa}, partial (5%)
Length = 381
Score = 234 bits (597), Expect = 1e-61
Identities = 112/126 (88%), Positives = 121/126 (95%)
Frame = +3
Query: 745 LFSIWSHDGKMMFENIIEATANFDDKYLVGVGSQGNVYKAELSEGLVVAVKKLHLVTDEE 804
LFSIWSHDGKMMFENIIEAT NFDDKYL+GVGSQGNVYKAELS G+VVAVKKLH++TDEE
Sbjct: 3 LFSIWSHDGKMMFENIIEATENFDDKYLIGVGSQGNVYKAELSSGMVVAVKKLHIITDEE 182
Query: 805 MSCFSSKSFMSEIETLTGIKHRNIIKLHGFCSHSKFSFLVYKFLEGGSLDQILNNDTQAV 864
+S FSSKSFMSEIETL+GI+HRNIIKLHGFCSHSKFSFLVYKFLEGGSL Q+LN+DTQA
Sbjct: 183 ISHFSSKSFMSEIETLSGIRHRNIIKLHGFCSHSKFSFLVYKFLEGGSLGQMLNSDTQAT 362
Query: 865 AFDWEK 870
AFDWEK
Sbjct: 363 AFDWEK 380
>TC91148 similar to GP|5732036|gb|AAD48937.1| similar to disease resistance
proteins; contains similarity ot Pfam family PF00560 -
Leucine Rich, partial (4%)
Length = 630
Score = 234 bits (596), Expect = 2e-61
Identities = 125/189 (66%), Positives = 142/189 (74%), Gaps = 5/189 (2%)
Frame = +3
Query: 5 PTLIMILCVLPTL-----SVAEDSEAKLALLKWKDSFDDQSQTLLSTWKNNTNPCKPKWR 59
P IMI+ ++ + +VAEDSEA+ ALLKWK SFD+QSQ+LLSTWKN TN C KW+
Sbjct: 69 PRFIMIMFIILFMISWPQAVAEDSEAQ-ALLKWKHSFDNQSQSLLSTWKNTTNTCT-KWK 242
Query: 60 GIKCDKSNFISTIGLANLGLKGTLHSLTFSSFPNLLMIDIRNNSFYGTIPAQIGNLSNIS 119
GI CD S ISTI L N GLKGTLHSLTFSSF NL ++I NN FYGTIP QIGN+S I+
Sbjct: 243 GIFCDNSKSISTINLENFGLKGTLHSLTFSSFSNLQTLNIYNNYFYGTIPPQIGNISKIN 422
Query: 120 ILTFKNNYFDGSIPQEMCTLTGLQFLDISFCKLNGAIPKSIGNLTNLSYLILGGNNWSGG 179
L F N DGSIPQEM TL LQ +D SFCKL+GAIP SIGNL+NL YL L GNN+ G
Sbjct: 423 TLNFSLNPIDGSIPQEMFTLKSLQNIDFSFCKLSGAIPNSIGNLSNLLYLDLRGNNFVGT 602
Query: 180 PIPPEIGKL 188
PIPPEIGKL
Sbjct: 603 PIPPEIGKL 629
Score = 64.7 bits (156), Expect = 2e-10
Identities = 47/124 (37%), Positives = 63/124 (49%), Gaps = 3/124 (2%)
Frame = +3
Query: 237 SKLDTLVLSNNTKMSGPIPHSLW--NMSSLTVLYFDNIGLSGSIPDSIQNLVNLKELALD 294
SK + + N + G + HSL + S+L L N G+IP I N+ + L
Sbjct: 261 SKSISTINLENFGLKGTL-HSLTFSSFSNLQTLNIYNNYFYGTIPPQIGNISKINTLNFS 437
Query: 295 INHLSGSIPSTIGDLKNLIKLYLGSNNLSGPIPASIGNLINLQVLSVQENNLTGT-IPAS 353
+N + GSIP + LK+L + LSG IP SIGNL NL L ++ NN GT IP
Sbjct: 438 LNPIDGSIPQEMFTLKSLQNIDFSFCKLSGAIPNSIGNLSNLLYLDLRGNNFVGTPIPPE 617
Query: 354 IGNL 357
IG L
Sbjct: 618 IGKL 629
Score = 64.3 bits (155), Expect = 2e-10
Identities = 41/97 (42%), Positives = 53/97 (54%)
Frame = +3
Query: 159 SIGNLTNLSYLILGGNNWSGGPIPPEIGKLNNLLHLAIQKSNLVGSIPQEIGFLTNLAYI 218
S NL L+ NN+ G IPP+IG ++ + L + + GSIPQE+ L +L I
Sbjct: 333 SFSNLQTLNIY----NNYFYGTIPPQIGNISKINTLNFSLNPIDGSIPQEMFTLKSLQNI 500
Query: 219 DLSKNSLSGGIPETIGNLSKLDTLVLSNNTKMSGPIP 255
D S LSG IP +IGNLS L L L N + PIP
Sbjct: 501 DFSFCKLSGAIPNSIGNLSNLLYLDLRGNNFVGTPIP 611
Score = 57.4 bits (137), Expect = 3e-08
Identities = 36/114 (31%), Positives = 58/114 (50%), Gaps = 1/114 (0%)
Frame = +3
Query: 284 NLVNLKELALDINHLSGSIPS-TIGDLKNLIKLYLGSNNLSGPIPASIGNLINLQVLSVQ 342
N ++ + L+ L G++ S T NL L + +N G IP IGN+ + L+
Sbjct: 258 NSKSISTINLENFGLKGTLHSLTFSSFSNLQTLNIYNNYFYGTIPPQIGNISKINTLNFS 437
Query: 343 ENNLTGTIPASIGNLKWLTVFEVATNKLHGRIPNGLYNITNWISFVVSENDFVG 396
N + G+IP + LK L + + KL G IPN + N++N + + N+FVG
Sbjct: 438 LNPIDGSIPQEMFTLKSLQNIDFSFCKLSGAIPNSIGNLSNLLYLDLRGNNFVG 599
Score = 56.2 bits (134), Expect = 6e-08
Identities = 38/120 (31%), Positives = 59/120 (48%), Gaps = 3/120 (2%)
Frame = +3
Query: 504 LGVLHLSSNQLTGKLPMEVLGGMKSLFDLKISNNHFSDNIPSEIGLLQRLQELDLGGNEL 563
+ ++L + L G L +L L I NN+F IP +IG + ++ L+ N +
Sbjct: 270 ISTINLENFGLKGTLHSLTFSSFSNLQTLNIYNNYFYGTIPPQIGNISKINTLNFSLNPI 449
Query: 564 SGKIPKELVELPNLRMLNLSRNKIEGIIPIKFD--SGLESLDLSG-NFLKGNIPTGLADL 620
G IP+E+ L +L+ ++ S K+ G IP S L LDL G NF+ IP + L
Sbjct: 450 DGSIPQEMFTLKSLQNIDFSFCKLSGAIPNSIGNLSNLLYLDLRGNNFVGTPIPPEIGKL 629
Score = 49.7 bits (117), Expect = 6e-06
Identities = 32/119 (26%), Positives = 57/119 (47%), Gaps = 2/119 (1%)
Frame = +3
Query: 310 KNLIKLYLGSNNLSGPIPA-SIGNLINLQVLSVQENNLTGTIPASIGNLKWLTVFEVATN 368
K++ + L + L G + + + + NLQ L++ N GTIP IGN+ + + N
Sbjct: 264 KSISTINLENFGLKGTLHSLTFSSFSNLQTLNIYNNYFYGTIPPQIGNISKINTLNFSLN 443
Query: 369 KLHGRIPNGLYNITNWISFVVSENDFVGHLPSQICSGGSLRLLNADHNRFTG-PIPTSL 426
+ G IP ++ + + + S G +P+ I + +L L+ N F G PIP +
Sbjct: 444 PIDGSIPQEMFTLKSLQNIDFSFCKLSGAIPNSIGNLSNLLYLDLRGNNFVGTPIPPEI 620
>TC78763 somatic embryogenesis receptor kinase 1 [Medicago truncatula]
Length = 2737
Score = 217 bits (553), Expect = 2e-56
Identities = 170/517 (32%), Positives = 240/517 (45%), Gaps = 33/517 (6%)
Frame = +3
Query: 556 LDLGGNELSGKIPKELVELPNLRMLNLSRNKIEGIIPIKFDS--GLESLDLSGNFLKGNI 613
+DLG LSG + +L +L NL+ L L N I G IP + L SLDL N G I
Sbjct: 648 VDLGNAALSGTLVPQLGQLKNLQYLELYSNNITGPIPSDLGNLTNLVSLDLYLNRFNGPI 827
Query: 614 PTGLADLVRLSKLNLSHNMLSGTIPQNFGR--NLVFVNISDNQLEGPLPKIPAFLSASFE 671
P L L +L L L++N L G IP + L +++S+NQL G +P +F +
Sbjct: 828 PDSLGKLSKLRFLRLNNNSLMGPIPMSLTNISALQVLDLSNNQLSGVVPDNGSFSLFTPI 1007
Query: 672 SLKNNNHLCGNIRGLDPCATS--HSRKRKNVLRPVFIALG-------------AVILVLC 716
S NN +LCG + G PC S S V P A G A +L
Sbjct: 1008 SFANNLNLCGPVTG-HPCPGSPPFSPPPPFVPPPPISAPGSGGATGAIAGGVAAGAALLF 1184
Query: 717 VVGALMYIMCGRKKPNE------ESQTEEVQRGVLFSIWSHDGKMMFENIIEATANFDDK 770
A+ + R+KP E + EV G L + + AT F +K
Sbjct: 1185 AAPAIAFAWWRRRKPQEFFFDVPAEEDPEVHLGQL-------KRFSLRELQVATDTFSNK 1343
Query: 771 YLVGVGSQGNVYKAELSEGLVVAVKKLHLVTDEEMSCFSSKSFMSEIETLTGIKHRNIIK 830
++G G G VYK L++G +VAVK+L EE + F +E+E ++ HRN+++
Sbjct: 1344 NILGRGGFGKVYKGRLADGSLVAVKRLK----EERTPGGELQFQTEVEMISMAVHRNLLR 1511
Query: 831 LHGFCSHSKFSFLVYKFLEGGSLDQILNN-DTQAVAFDWEKRVNVVKGVANALSYLHHDC 889
L GFC LVY ++ GS+ L DW R + G A LSYLH C
Sbjct: 1512 LRGFCMTPTERLLVYPYMANGSVASCLRERPPHQEPLDWPTRKRIALGSARGLSYLHDHC 1691
Query: 890 SPPIIHRDISSKNVLLNLDYEAHVSDFGTAKFLK-PGLHSWTQFAGTFGYAAPELAQTME 948
P IIHRD+ + N+LL+ ++EA V DFG AK + H T GT G+ APE T +
Sbjct: 1692 DPKIIHRDVKAANILLDEEFEAVVGDFGLAKLMDYKDTHVTTAVRGTIGHIAPEYLSTGK 1871
Query: 949 VNEKCDVYSFGVLALETIMGKHPGDLISL------FLSPSTRPMANNMLLTDVLDQRPQQ 1002
+EK DV+ +G++ LE I G+ DL L L + + L ++D P
Sbjct: 1872 SSEKTDVFGYGIMLLELITGQRAFDLARLANDDDVMLLDWVKGLLKEKKLEMLVD--PDL 2045
Query: 1003 VMEPIDEEVILIARLAFACLSQNPRLRPSMGQVCKML 1039
I+ EV + ++A C +P RP M V +ML
Sbjct: 2046 KTNYIEAEVEQLIQVALLCTQGSPMDRPKMSDVVRML 2156
Score = 95.5 bits (236), Expect = 1e-19
Identities = 51/109 (46%), Positives = 72/109 (65%)
Frame = +3
Query: 173 GNNWSGGPIPPEIGKLNNLLHLAIQKSNLVGSIPQEIGFLTNLAYIDLSKNSLSGGIPET 232
GN G + P++G+L NL +L + +N+ G IP ++G LTNL +DL N +G IP++
Sbjct: 657 GNAALSGTLVPQLGQLKNLQYLELYSNNITGPIPSDLGNLTNLVSLDLYLNRFNGPIPDS 836
Query: 233 IGNLSKLDTLVLSNNTKMSGPIPHSLWNMSSLTVLYFDNIGLSGSIPDS 281
+G LSKL L L+NN+ M GPIP SL N+S+L VL N LSG +PD+
Sbjct: 837 LGKLSKLRFLRLNNNSLM-GPIPMSLTNISALQVLDLSNNQLSGVVPDN 980
Score = 90.1 bits (222), Expect = 4e-18
Identities = 54/147 (36%), Positives = 84/147 (56%), Gaps = 1/147 (0%)
Frame = +3
Query: 260 NMSSLTVLYFDNIGLSGSIPDSIQNLVNLKELALDINHLSGSIPSTIGDLKNLIKLYLGS 319
N +S+ + N LSG++ + L NL+ L L N+++G IPS +G+L NL+ L L
Sbjct: 627 NDNSVIRVDLGNAALSGTLVPQLGQLKNLQYLELYSNNITGPIPSDLGNLTNLVSLDLYL 806
Query: 320 NNLSGPIPASIGNLINLQVLSVQENNLTGTIPASIGNLKWLTVFEVATNKLHGRIP-NGL 378
N +GPIP S+G L L+ L + N+L G IP S+ N+ L V +++ N+L G +P NG
Sbjct: 807 NRFNGPIPDSLGKLSKLRFLRLNNNSLMGPIPMSLTNISALQVLDLSNNQLSGVVPDNGS 986
Query: 379 YNITNWISFVVSENDFVGHLPSQICSG 405
+++ ISF + N G + C G
Sbjct: 987 FSLFTPISFANNLN-LCGPVTGHPCPG 1064
Score = 84.0 bits (206), Expect = 3e-16
Identities = 67/245 (27%), Positives = 104/245 (42%)
Frame = +3
Query: 10 ILCVLPTLSVAEDSEAKLALLKWKDSFDDQSQTLLSTWKNNTNPCKPKWRGIKCDKSNFI 69
+L + P V+ + E AL + + D + L S NPC W + C+ N +
Sbjct: 471 LLLLHPLWLVSANMEGD-ALHNLRTNLQDPNNVLQSWDPTLVNPCT--WFHVTCNNDNSV 641
Query: 70 STIGLANLGLKGTLHSLTFSSFPNLLMIDIRNNSFYGTIPAQIGNLSNISILTFKNNYFD 129
+ L N L GTL +P Q+G L N+ L +N
Sbjct: 642 IRVDLGNAALSGTL------------------------VP-QLGQLKNLQYLELYSNNIT 746
Query: 130 GSIPQEMCTLTGLQFLDISFCKLNGAIPKSIGNLTNLSYLILGGNNWSGGPIPPEIGKLN 189
G IP ++ LT L LD+ + NG IP S+G L+ L +L +LN
Sbjct: 747 GPIPSDLGNLTNLVSLDLYLNRFNGPIPDSLGKLSKLRFL-----------------RLN 875
Query: 190 NLLHLAIQKSNLVGSIPQEIGFLTNLAYIDLSKNSLSGGIPETIGNLSKLDTLVLSNNTK 249
N ++L+G IP + ++ L +DLS N LSG +P+ G+ S + +NN
Sbjct: 876 N--------NSLMGPIPMSLTNISALQVLDLSNNQLSGVVPDN-GSFSLFTPISFANNLN 1028
Query: 250 MSGPI 254
+ GP+
Sbjct: 1029LCGPV 1043
Score = 77.8 bits (190), Expect = 2e-14
Identities = 44/122 (36%), Positives = 65/122 (53%)
Frame = +3
Query: 247 NTKMSGPIPHSLWNMSSLTVLYFDNIGLSGSIPDSIQNLVNLKELALDINHLSGSIPSTI 306
N +SG + L + +L L + ++G IP + NL NL L L +N +G IP ++
Sbjct: 660 NAALSGTLVPQLGQLKNLQYLELYSNNITGPIPSDLGNLTNLVSLDLYLNRFNGPIPDSL 839
Query: 307 GDLKNLIKLYLGSNNLSGPIPASIGNLINLQVLSVQENNLTGTIPASIGNLKWLTVFEVA 366
G L L L L +N+L GPIP S+ N+ LQVL + N L+G +P + G+ T A
Sbjct: 840 GKLSKLRFLRLNNNSLMGPIPMSLTNISALQVLDLSNNQLSGVVPDN-GSFSLFTPISFA 1016
Query: 367 TN 368
N
Sbjct: 1017NN 1022
Score = 72.4 bits (176), Expect = 9e-13
Identities = 36/119 (30%), Positives = 69/119 (57%)
Frame = +3
Query: 305 TIGDLKNLIKLYLGSNNLSGPIPASIGNLINLQVLSVQENNLTGTIPASIGNLKWLTVFE 364
T + ++I++ LG+ LSG + +G L NLQ L + NN+TG IP+ +GNL L +
Sbjct: 618 TCNNDNSVIRVDLGNAALSGTLVPQLGQLKNLQYLELYSNNITGPIPSDLGNLTNLVSLD 797
Query: 365 VATNKLHGRIPNGLYNITNWISFVVSENDFVGHLPSQICSGGSLRLLNADHNRFTGPIP 423
+ N+ +G IP+ L ++ ++ N +G +P + + +L++L+ +N+ +G +P
Sbjct: 798 LYLNRFNGPIPDSLGKLSKLRFLRLNNNSLMGPIPMSLTNISALQVLDLSNNQLSGVVP 974
Score = 68.6 bits (166), Expect = 1e-11
Identities = 39/119 (32%), Positives = 63/119 (52%)
Frame = +3
Query: 346 LTGTIPASIGNLKWLTVFEVATNKLHGRIPNGLYNITNWISFVVSENDFVGHLPSQICSG 405
L+GT+ +G LK L E+ +N + G IP+ L N+TN +S + N F G +P +
Sbjct: 669 LSGTLVPQLGQLKNLQYLELYSNNITGPIPSDLGNLTNLVSLDLYLNRFNGPIPDSLGKL 848
Query: 406 GSLRLLNADHNRFTGPIPTSLKTCSSIERITLEVNQIEGDIAQDFGVYPKLQYLDLSDN 464
LR L ++N GPIP SL S+++ + L NQ+ G + D G + + ++N
Sbjct: 849 SKLRFLRLNNNSLMGPIPMSLTNISALQVLDLSNNQLSG-VVPDNGSFSLFTPISFANN 1022
Score = 62.8 bits (151), Expect = 7e-10
Identities = 41/120 (34%), Positives = 63/120 (52%), Gaps = 1/120 (0%)
Frame = +3
Query: 419 TGPIPTSLKTCSSIERITLEVNQIEGDIAQDFGVYPKLQYLDLSDNKFHGQISPNWGKSL 478
+G + L +++ + L N I G I D G L LDL N+F+G I + GK
Sbjct: 672 SGTLVPQLGQLKNLQYLELYSNNITGPIPSDLGNLTNLVSLDLYLNRFNGPIPDSLGKLS 851
Query: 479 NLQTFIISNNNISGVIPLDFIGLTKLGVLHLSSNQLTGKLPMEVLGGMKSLF-DLKISNN 537
L+ ++NN++ G IP+ ++ L VL LS+NQL+G +P G SLF + +NN
Sbjct: 852 KLRFLRLNNNSLMGPIPMSLTNISALQVLDLSNNQLSGVVPD---NGSFSLFTPISFANN 1022
Score = 57.8 bits (138), Expect = 2e-08
Identities = 33/100 (33%), Positives = 51/100 (51%)
Frame = +3
Query: 396 GHLPSQICSGGSLRLLNADHNRFTGPIPTSLKTCSSIERITLEVNQIEGDIAQDFGVYPK 455
G L Q+ +L+ L N TGPIP+ L +++ + L +N+ G I G K
Sbjct: 675 GTLVPQLGQLKNLQYLELYSNNITGPIPSDLGNLTNLVSLDLYLNRFNGPIPDSLGKLSK 854
Query: 456 LQYLDLSDNKFHGQISPNWGKSLNLQTFIISNNNISGVIP 495
L++L L++N G I + LQ +SNN +SGV+P
Sbjct: 855 LRFLRLNNNSLMGPIPMSLTNISALQVLDLSNNQLSGVVP 974
>TC80900 weakly similar to PIR|G84652|G84652 probable receptor-like protein
kinase [imported] - Arabidopsis thaliana, partial (24%)
Length = 1054
Score = 207 bits (528), Expect = 1e-53
Identities = 132/375 (35%), Positives = 199/375 (52%)
Frame = +2
Query: 212 LTNLAYIDLSKNSLSGGIPETIGNLSKLDTLVLSNNTKMSGPIPHSLWNMSSLTVLYFDN 271
+ L +I L N+LS IP+ IGNL L+ L L N
Sbjct: 11 MKRLKWIYLGYNNLSREIPKNIGNLVSLNHLNLVYNN----------------------- 121
Query: 272 IGLSGSIPDSIQNLVNLKELALDINHLSGSIPSTIGDLKNLIKLYLGSNNLSGPIPASIG 331
L+G IP+S+ NL NL+ L L +N L+G IP +I +LKNLI L L N LSG I +
Sbjct: 122 --LTGPIPESLGNLTNLQYLFLYLNKLTGPIPKSIFNLKNLISLDLSDNYLSGEISNLVV 295
Query: 332 NLINLQVLSVQENNLTGTIPASIGNLKWLTVFEVATNKLHGRIPNGLYNITNWISFVVSE 391
NL L++L + NN TG IP +I +L L V ++ +NKL G IP L N +S
Sbjct: 296 NLQKLEILHLFSNNFTGKIPNTITSLPHLQVLQLWSNKLTGEIPQTLGIHNNLTILDLSS 475
Query: 392 NDFVGHLPSQICSGGSLRLLNADHNRFTGPIPTSLKTCSSIERITLEVNQIEGDIAQDFG 451
N+ G +P+ +C+ +L + N G IP L +C ++ER+ L+ N + G + +
Sbjct: 476 NNLTGKIPNSLCASKNLHKIILFSNSLKGEIPKGLTSCKTLERVRLQDNNLSGKLPLEIT 655
Query: 452 VYPKLQYLDLSDNKFHGQISPNWGKSLNLQTFIISNNNISGVIPLDFIGLTKLGVLHLSS 511
P++ LD+S NKF G+I+ +LQ ++NNN SG +P F G K+ L LS
Sbjct: 656 QLPQIYLLDISGNKFSGKINDRKWNMPSLQMLNLANNNFSGDLPNSF-GGNKVEGLDLSQ 832
Query: 512 NQLTGKLPMEVLGGMKSLFDLKISNNHFSDNIPSEIGLLQRLQELDLGGNELSGKIPKEL 571
NQ +G + + + L LK++NN+ P E+ +L LDL N L+G+IP++L
Sbjct: 833 NQFSGYIQIG-FKNLPELVQLKLNNNNLFGKFPEELFQCNKLVSLDLSHNRLNGEIPEKL 1009
Query: 572 VELPNLRMLNLSRNK 586
++P L +L++S N+
Sbjct: 1010AKMPVLGLLDISENQ 1054
Score = 197 bits (501), Expect = 2e-50
Identities = 125/356 (35%), Positives = 191/356 (53%), Gaps = 3/356 (0%)
Frame = +2
Query: 309 LKNLIKLYLGSNNLSGPIPASIGNLINLQVLSVQENNLTGTIPASIGNLKWLTVFEVATN 368
+K L +YLG NNLS IP +IGNL++L L++ NNLTG IP S+GNL L + N
Sbjct: 11 MKRLKWIYLGYNNLSREIPKNIGNLVSLNHLNLVYNNLTGPIPESLGNLTNLQYLFLYLN 190
Query: 369 KLHGRIPNGLYNITNWISFVVSENDFVGHLPSQICSGGSLRLLNADHNRFTGPIPTSLKT 428
KL G IP ++N+ N IS +S+N G + + + + L +L+ N FTG IP ++ +
Sbjct: 191 KLTGPIPKSIFNLKNLISLDLSDNYLSGEISNLVVNLQKLEILHLFSNNFTGKIPNTITS 370
Query: 429 CSSIERITLEVNQIEGDIAQDFGVYPKLQYLDLSDNKFHGQISPNWGKSLNLQTFIISNN 488
++ + L N++ G+I Q G++ L LDLS N G+I + S NL I+ +N
Sbjct: 371 LPHLQVLQLWSNKLTGEIPQTLGIHNNLTILDLSSNNLTGKIPNSLCASKNLHKIILFSN 550
Query: 489 NISGVIPLDFIGLTKLGVLHLSSNQLTGKLPMEVLGGMKSLFDLKISNNHFSDNIPSEIG 548
++ G IP GLT K+L +++ +N+ S +P EI
Sbjct: 551 SLKGEIPK---GLTSC----------------------KTLERVRLQDNNLSGKLPLEIT 655
Query: 549 LLQRLQELDLGGNELSGKIPKELVELPNLRMLNLSRNKIEGIIPIKFDSG-LESLDLSGN 607
L ++ LD+ GN+ SGKI +P+L+MLNL+ N G +P F +E LDLS N
Sbjct: 656 QLPQIYLLDISGNKFSGKINDRKWNMPSLQMLNLANNNFSGDLPNSFGGNKVEGLDLSQN 835
Query: 608 FLKGNIPTGLADLVRLSKLNLSHNMLSGTIPQNFGR--NLVFVNISDNQLEGPLPK 661
G I G +L L +L L++N L G P+ + LV +++S N+L G +P+
Sbjct: 836 QFSGYIQIGFKNLPELVQLKLNNNNLFGKFPEELFQCNKLVSLDLSHNRLNGEIPE 1003
Score = 184 bits (468), Expect = 1e-46
Identities = 126/372 (33%), Positives = 200/372 (52%)
Frame = +2
Query: 142 LQFLDISFCKLNGAIPKSIGNLTNLSYLILGGNNWSGGPIPPEIGKLNNLLHLAIQKSNL 201
L+++ + + L+ IPK+IGNL +L++L L NN + GPIP +G L NL +L + + L
Sbjct: 20 LKWIYLGYNNLSREIPKNIGNLVSLNHLNLVYNNLT-GPIPESLGNLTNLQYLFLYLNKL 196
Query: 202 VGSIPQEIGFLTNLAYIDLSKNSLSGGIPETIGNLSKLDTLVLSNNTKMSGPIPHSLWNM 261
G IP+ I L NL +DLS N LSG I + NL KL+ L L +N +G IP+++ ++
Sbjct: 197 TGPIPKSIFNLKNLISLDLSDNYLSGEISNLVVNLQKLEILHLFSN-NFTGKIPNTITSL 373
Query: 262 SSLTVLYFDNIGLSGSIPDSIQNLVNLKELALDINHLSGSIPSTIGDLKNLIKLYLGSNN 321
L VL + L+G IP ++ NL L L N+L+G IP+++ KNL K+ L SN+
Sbjct: 374 PHLQVLQLWSNKLTGEIPQTLGIHNNLTILDLSSNNLTGKIPNSLCASKNLHKIILFSNS 553
Query: 322 LSGPIPASIGNLINLQVLSVQENNLTGTIPASIGNLKWLTVFEVATNKLHGRIPNGLYNI 381
L G IP + + L+ + +Q+NNL+G +P I L + + +++ NK G+I
Sbjct: 554 LKGEIPKGLTSCKTLERVRLQDNNLSGKLPLEITQLPQIYLLDISGNKFSGKI------- 712
Query: 382 TNWISFVVSENDFVGHLPSQICSGGSLRLLNADHNRFTGPIPTSLKTCSSIERITLEVNQ 441
ND ++P SL++LN +N F+G +P S + +E + L NQ
Sbjct: 713 ----------NDRKWNMP-------SLQMLNLANNNFSGDLPNSFGG-NKVEGLDLSQNQ 838
Query: 442 IEGDIAQDFGVYPKLQYLDLSDNKFHGQISPNWGKSLNLQTFIISNNNISGVIPLDFIGL 501
G I F P+L L L++N G+ + L + +S+N ++G IP +
Sbjct: 839 FSGYIQIGFKNLPELVQLKLNNNNLFGKFPEELFQCNKLVSLDLSHNRLNGEIPEKLAKM 1018
Query: 502 TKLGVLHLSSNQ 513
LG+L +S NQ
Sbjct: 1019PVLGLLDISENQ 1054
Score = 165 bits (418), Expect = 8e-41
Identities = 111/340 (32%), Positives = 168/340 (48%)
Frame = +2
Query: 102 NSFYGTIPAQIGNLSNISILTFKNNYFDGSIPQEMCTLTGLQFLDISFCKLNGAIPKSIG 161
N+ IP IGNL +++ L N G IP+ + LT LQ+L + KL G IPKSI
Sbjct: 44 NNLSREIPKNIGNLVSLNHLNLVYNNLTGPIPESLGNLTNLQYLFLYLNKLTGPIPKSIF 223
Query: 162 NLTNLSYLILGGNNWSGGPIPPEIGKLNNLLHLAIQKSNLVGSIPQEIGFLTNLAYIDLS 221
NL NL L L +N+ G I + L L L + +N G IP I L +L + L
Sbjct: 224 NLKNLISLDL-SDNYLSGEISNLVVNLQKLEILHLFSNNFTGKIPNTITSLPHLQVLQLW 400
Query: 222 KNSLSGGIPETIGNLSKLDTLVLSNNTKMSGPIPHSLWNMSSLTVLYFDNIGLSGSIPDS 281
N L+G IP+T+G + L L LS+N ++G IP+SL +L + + L G IP
Sbjct: 401 SNKLTGEIPQTLGIHNNLTILDLSSN-NLTGKIPNSLCASKNLHKIILFSNSLKGEIPKG 577
Query: 282 IQNLVNLKELALDINHLSGSIPSTIGDLKNLIKLYLGSNNLSGPIPASIGNLINLQVLSV 341
+ + L+ + L N+LSG +P I L + L + N SG I N+ +LQ+L++
Sbjct: 578 LTSCKTLERVRLQDNNLSGKLPLEITQLPQIYLLDISGNKFSGKINDRKWNMPSLQMLNL 757
Query: 342 QENNLTGTIPASIGNLKWLTVFEVATNKLHGRIPNGLYNITNWISFVVSENDFVGHLPSQ 401
NN +G +P S G K + +++ N+ G I G N+ + ++ N+ G P +
Sbjct: 758 ANNNFSGDLPNSFGGNK-VEGLDLSQNQFSGYIQIGFKNLPELVQLKLNNNNLFGKFPEE 934
Query: 402 ICSGGSLRLLNADHNRFTGPIPTSLKTCSSIERITLEVNQ 441
+ L L+ HNR G IP L + + + NQ
Sbjct: 935 LFQCNKLVSLDLSHNRLNGEIPEKLAKMPVLGLLDISENQ 1054
Score = 117 bits (294), Expect = 2e-26
Identities = 88/278 (31%), Positives = 136/278 (48%)
Frame = +2
Query: 67 NFISTIGLANLGLKGTLHSLTFSSFPNLLMIDIRNNSFYGTIPAQIGNLSNISILTFKNN 126
N IS + L++ L G + +L + L ++ + +N+F G IP I +L ++ +L +N
Sbjct: 233 NLIS-LDLSDNYLSGEISNLVVN-LQKLEILHLFSNNFTGKIPNTITSLPHLQVLQLWSN 406
Query: 127 YFDGSIPQEMCTLTGLQFLDISFCKLNGAIPKSIGNLTNLSYLILGGNNWSGGPIPPEIG 186
G IPQ + L LD+S L G IP S+ NL +IL N+ G IP +
Sbjct: 407 KLTGEIPQTLGIHNNLTILDLSSNNLTGKIPNSLCASKNLHKIILFSNSLK-GEIPKGLT 583
Query: 187 KLNNLLHLAIQKSNLVGSIPQEIGFLTNLAYIDLSKNSLSGGIPETIGNLSKLDTLVLSN 246
L + +Q +NL G +P EI L + +D+S N SG I + N+ L L L+N
Sbjct: 584 SCKTLERVRLQDNNLSGKLPLEITQLPQIYLLDISGNKFSGKINDRKWNMPSLQMLNLAN 763
Query: 247 NTKMSGPIPHSLWNMSSLTVLYFDNIGLSGSIPDSIQNLVNLKELALDINHLSGSIPSTI 306
N SG +P+S + + + L SG I +NL L +L L+ N+L G P +
Sbjct: 764 N-NFSGDLPNS-FGGNKVEGLDLSQNQFSGYIQIGFKNLPELVQLKLNNNNLFGKFPEEL 937
Query: 307 GDLKNLIKLYLGSNNLSGPIPASIGNLINLQVLSVQEN 344
L+ L L N L+G IP + + L +L + EN
Sbjct: 938 FQCNKLVSLDLSHNRLNGEIPEKLAKMPVLGLLDISEN 1051
Score = 60.1 bits (144), Expect = 4e-09
Identities = 57/196 (29%), Positives = 77/196 (39%), Gaps = 53/196 (27%)
Frame = +2
Query: 549 LLQRLQELDLGGNELSGKIPKELVELPNLRMLNLSRNKIEGIIPIKFDS----------- 597
L++RL+ + LG N LS +IPK + L +L LNL N + G IP +
Sbjct: 8 LMKRLKWIYLGYNNLSREIPKNIGNLVSLNHLNLVYNNLTGPIPESLGNLTNLQYLFLYL 187
Query: 598 ---------------GLESLDLSGNFLK------------------------GNIPTGLA 618
L SLDLS N+L G IP +
Sbjct: 188 NKLTGPIPKSIFNLKNLISLDLSDNYLSGEISNLVVNLQKLEILHLFSNNFTGKIPNTIT 367
Query: 619 DLVRLSKLNLSHNMLSGTIPQNFG--RNLVFVNISDNQLEGPLPKIPAFLSASFESLKNN 676
L L L L N L+G IPQ G NL +++S N L G +P + + +
Sbjct: 368 SLPHLQVLQLWSNKLTGEIPQTLGIHNNLTILDLSSNNLTGKIPNSLCASKNLHKIILFS 547
Query: 677 NHLCGNI-RGLDPCAT 691
N L G I +GL C T
Sbjct: 548 NSLKGEIPKGLTSCKT 595
>BE941041 similar to GP|14596041|gb Unknown protein {Arabidopsis thaliana},
partial (28%)
Length = 408
Score = 204 bits (519), Expect = 1e-52
Identities = 96/135 (71%), Positives = 111/135 (82%)
Frame = +2
Query: 861 TQAVAFDWEKRVNVVKGVANALSYLHHDCSPPIIHRDISSKNVLLNLDYEAHVSDFGTAK 920
+QA+AF+ EKRVN+V+GVA+ALSY+HHDC PPI+HRDISSKNVLL++ YEA +SDFGTAK
Sbjct: 2 SQAIAFEGEKRVNIVRGVADALSYMHHDCIPPIVHRDISSKNVLLDISYEAQLSDFGTAK 181
Query: 921 FLKPGLHSWTQFAGTFGYAAPELAQTMEVNEKCDVYSFGVLALETIMGKHPGDLISLFLS 980
FLKP SWT FAGT+GYAAPE AQTMEV EKCDVYSFGVL E ++GKHP D IS S
Sbjct: 182 FLKPDSSSWTAFAGTYGYAAPEFAQTMEVTEKCDVYSFGVLCFEILLGKHPADFISSLFS 361
Query: 981 PSTRPMANNMLLTDV 995
ST M N+LL DV
Sbjct: 362 SSTAKMTYNLLLIDV 406
>BQ123304 similar to GP|22093756|d putative receptor protein kinase {Oryza
sativa (japonica cultivar-group)}, partial (19%)
Length = 979
Score = 189 bits (481), Expect = 4e-48
Identities = 112/297 (37%), Positives = 168/297 (55%), Gaps = 15/297 (5%)
Frame = +2
Query: 758 ENIIEATANFDDKYLVGVGSQGNVYKAELSEGLVVAVKKLHLVTDEEMSCFSSKSFMSEI 817
+ I++AT N D++++G G G VY+ EL G VAVKK+ L DE + KSF+ E+
Sbjct: 2 QEIMDATNNLSDEFIIGSGGSGTVYRVELPTGETVAVKKISL-KDEYLL---HKSFIREV 169
Query: 818 ETLTGIKHRNIIKLHGFCSH----SKFSFLVYKFLEGGSLDQILNNDTQAV--AFDWEKR 871
+TL IKHR+++KL G CS+ + + L+Y+F+E GS+ L+ + + + DW+ R
Sbjct: 170 KTLGRIKHRHLVKLVGCCSNRHKGNGCNLLIYEFMENGSVWDWLHGNALKLRRSLDWDTR 349
Query: 872 VNVVKGVANALSYLHHDCSPPIIHRDISSKNVLLNLDYEAHVSDFGTAKFLKPGLHSWTQ 931
+ G+A + YLHHDC P IIHRDI S N+LL+ + +AH+ DFG AK + L S T+
Sbjct: 350 FKIALGLAQGMEYLHHDCVPKIIHRDIKSSNILLDSNMDAHLGDFGLAKAIVENLDSNTE 529
Query: 932 ----FAGTFGYAAPELAQTMEVNEKCDVYSFGVLALETIMGKHPGDLISLFLSPSTR--P 985
FAG++GY APE +++ EK DVYS GV+ +E + GK P D R
Sbjct: 530 STSCFAGSYGYIAPEFGYSLKATEKSDVYSMGVVLMELVSGKLPTDAAFRGSVDMVRWVE 709
Query: 986 MANNMLLT---DVLDQRPQQVMEPIDEEVILIARLAFACLSQNPRLRPSMGQVCKML 1039
M NM T +++D + ++ + + +A C P RPS QVC +L
Sbjct: 710 MRINMKGTEREELVDPELKPLLPYEEFAAFQVLEIAIQCTKTTPMKRPSSRQVCDLL 880
>TC81789 similar to PIR|B86440|B86440 probable protein kinase [imported] -
Arabidopsis thaliana, partial (54%)
Length = 1189
Score = 121 bits (304), Expect(2) = 1e-47
Identities = 65/191 (34%), Positives = 107/191 (55%), Gaps = 1/191 (0%)
Frame = +2
Query: 714 VLCVVGALMYI-MCGRKKPNEESQTEEVQRGVLFSIWSHDGKMMFENIIEATANFDDKYL 772
++C G +Y C K + S +V G ++ D ++II+ +++++
Sbjct: 8 LMCFWGCFLYKKFC---KNDRISLAGDVGPGASIVMFHGDLPYSSKDIIKKLETLNEEHI 178
Query: 773 VGVGSQGNVYKAELSEGLVVAVKKLHLVTDEEMSCFSSKSFMSEIETLTGIKHRNIIKLH 832
+GVG G VYK + +G V A+KK+ + + + F E+ L IKHR ++ L
Sbjct: 179 IGVGGFGTVYKLAMDDGNVFALKKIVKLNEG-----FDRFFERELAILGSIKHRYLVNLR 343
Query: 833 GFCSHSKFSFLVYKFLEGGSLDQILNNDTQAVAFDWEKRVNVVKGVANALSYLHHDCSPP 892
G+C+ L+Y +L GGSLD++L+ ++ + DW+ R+N++ G A L+YLHHDCSP
Sbjct: 344 GYCNSPTSKLLIYDYLPGGSLDEVLHEKSEQL--DWDSRLNIIMGAAKGLAYLHHDCSPR 517
Query: 893 IIHRDISSKNV 903
IIHRDI S N+
Sbjct: 518 IIHRDIKSSNI 550
Score = 88.2 bits (217), Expect(2) = 1e-47
Identities = 57/143 (39%), Positives = 81/143 (55%), Gaps = 7/143 (4%)
Frame = +3
Query: 904 LLNLDYEAHVSDFGTAKFLKPG-LHSWTQFAGTFGYAAPELAQTMEVNEKCDVYSFGVLA 962
LL+ +A VSDFG AK L+ H T AGTFGY APE Q+ EK DVYSFGVL
Sbjct: 552 LLDGKLDARVSDFGLAKLLEDEESHITTIVAGTFGYLAPEYMQSGRATEKTDVYSFGVLT 731
Query: 963 LETIMGKHPGDLISLFLSPSTRPMA-NNMLLTDVLDQRPQQVMEPI-----DEEVILIAR 1016
LE + GK P D + F+ + N L+T + RP+++++P+ E + +
Sbjct: 732 LEVLSGKRPTD--ASFIEKGLNVVGWLNFLIT---ENRPREIVDPLCDGVQVESLDALLS 896
Query: 1017 LAFACLSQNPRLRPSMGQVCKML 1039
+A C+S NP RP+M +V ++L
Sbjct: 897 MAIQCVSSNPEDRPTMHRVVQLL 965
>TC87497 similar to GP|9759480|dbj|BAB10485.1 serine/threonine protein
kinase-like protein {Arabidopsis thaliana}, partial (59%)
Length = 1331
Score = 183 bits (464), Expect = 4e-46
Identities = 129/376 (34%), Positives = 190/376 (50%), Gaps = 42/376 (11%)
Frame = +2
Query: 706 IALGAVILVLCVVGALMYIMCGRKKPNEESQTEEVQRGVLFS-----------------I 748
+A A V+ V+ A + ++C +K + + +++ L +
Sbjct: 68 LAAIASFFVVTVILAAIILICQHRKTTQTRRNNQIRTRPLRNPNHYPTSSSSLPVDASWS 247
Query: 749 WSHDGKMMFENIIEATANFDDKYLVGVGSQGNVYKAELSEGLVVAVKKLHLVTDEEMSCF 808
W + K+ E + AT NF + +VG GS G VYKA LS G VAVKKL + + F
Sbjct: 248 WDPNLKISMEELSRATKNFSNTLIVGDGSFGYVYKASLSTGATVAVKKL---SPDAFQGF 418
Query: 809 SSKSFMSEIETLTGIKHRNIIKLHGFCSHSKFSFLVYKFLEGGSLDQILN---------- 858
+ F +E+ETL+ ++H NI+K+ G+ + LVY+F+E G+LDQ L+
Sbjct: 419 --REFAAEMETLSKLRHHNIVKILGYWASGAERLLVYEFIEKGNLDQWLHESSPTSSSTH 592
Query: 859 -NDTQAVAFD-------WEKRVNVVKGVANALSYLHHDCSPPIIHRDISSKNVLLNLDYE 910
ND +++ D WE RV +++GVA+ L YL H PIIHRDI + NVLL+ ++E
Sbjct: 593 QNDEVSISIDFIRSPLPWETRVKIIRGVAHGLCYL-HGLEKPIIHRDIKASNVLLDSEFE 769
Query: 911 AHVSDFGTAKFL-KPGLHSWTQFAGTFGYAAPELAQTMEV-NEKCDVYSFGVLALETIMG 968
AH++DFG A+ + K H TQ AGT GY PE V N K DVYSFGVL +ET+ G
Sbjct: 770 AHIADFGLARRMDKSHSHVSTQVAGTIGYMPPEYRDGSNVANPKVDVYSFGVLMIETVSG 949
Query: 969 KHPGDLISL-----FLSPSTRPMANNMLLTDVLDQRPQQVMEPIDEEVILIARLAFACLS 1023
P + L L R M ++LD + +E V R+A C
Sbjct: 950 HRPNLAVKLEGNDIGLVNWARKMKERNTELEMLDGNIPREEGLKEESVREYVRIACMCTG 1129
Query: 1024 QNPRLRPSMGQVCKML 1039
+ + RP M +V K+L
Sbjct: 1130 ELQKDRPEMPEVVKLL 1177
>TC93082 similar to GP|1184077|gb|AAC15780.1| Cf-2.2 {Lycopersicon
pimpinellifolium}, partial (3%)
Length = 662
Score = 180 bits (457), Expect(2) = 2e-45
Identities = 91/157 (57%), Positives = 110/157 (69%)
Frame = +2
Query: 8 IMILCVLPTLSVAEDSEAKLALLKWKDSFDDQSQTLLSTWKNNTNPCKPKWRGIKCDKSN 67
+ + + P +E +ALLKWKDSFD+ SQ LLSTW T+PC W GI+CDKS
Sbjct: 92 VRLTIIFPQQVAGFSNEEAVALLKWKDSFDNHSQALLSTWTRTTSPCN--WEGIQCDKSK 265
Query: 68 FISTIGLANLGLKGTLHSLTFSSFPNLLMIDIRNNSFYGTIPAQIGNLSNISILTFKNNY 127
ISTI LAN GLKG LH+L+FSSFPNLL+++I NN+FYGTIP QIGNLS I+ L F N
Sbjct: 266 SISTINLANYGLKGKLHTLSFSSFPNLLILNIFNNNFYGTIPPQIGNLSRINTLNFSKNP 445
Query: 128 FDGSIPQEMCTLTGLQFLDISFCKLNGAIPKSIGNLT 164
GSIP EM TL L+ LD + C+L G IP SIGNL+
Sbjct: 446 IIGSIPIEMWTLRSLKGLDFAQCQLTGEIPNSIGNLS 556
Score = 58.2 bits (139), Expect = 2e-08
Identities = 33/80 (41%), Positives = 49/80 (61%)
Frame = +2
Query: 159 SIGNLTNLSYLILGGNNWSGGPIPPEIGKLNNLLHLAIQKSNLVGSIPQEIGFLTNLAYI 218
S + NL L + NN+ G IPP+IG L+ + L K+ ++GSIP E+ L +L +
Sbjct: 323 SFSSFPNLLILNIFNNNFYG-TIPPQIGNLSRINTLNFSKNPIIGSIPIEMWTLRSLKGL 499
Query: 219 DLSKNSLSGGIPETIGNLSK 238
D ++ L+G IP +IGNLSK
Sbjct: 500 DFAQCQLTGEIPNSIGNLSK 559
Score = 52.8 bits (125), Expect = 7e-07
Identities = 34/96 (35%), Positives = 49/96 (50%), Gaps = 1/96 (1%)
Frame = +2
Query: 263 SLTVLYFDNIGLSGSIPD-SIQNLVNLKELALDINHLSGSIPSTIGDLKNLIKLYLGSNN 321
S++ + N GL G + S + NL L + N+ G+IP IG+L + L N
Sbjct: 266 SISTINLANYGLKGKLHTLSFSSFPNLLILNIFNNNFYGTIPPQIGNLSRINTLNFSKNP 445
Query: 322 LSGPIPASIGNLINLQVLSVQENNLTGTIPASIGNL 357
+ G IP + L +L+ L + LTG IP SIGNL
Sbjct: 446 IIGSIPIEMWTLRSLKGLDFAQCQLTGEIPNSIGNL 553
Score = 51.6 bits (122), Expect = 2e-06
Identities = 27/75 (36%), Positives = 42/75 (56%)
Frame = +2
Query: 311 NLIKLYLGSNNLSGPIPASIGNLINLQVLSVQENNLTGTIPASIGNLKWLTVFEVATNKL 370
NL+ L + +NN G IP IGNL + L+ +N + G+IP + L+ L + A +L
Sbjct: 341 NLLILNIFNNNFYGTIPPQIGNLSRINTLNFSKNPIIGSIPIEMWTLRSLKGLDFAQCQL 520
Query: 371 HGRIPNGLYNITNWI 385
G IPN + N++ I
Sbjct: 521 TGEIPNSIGNLSKTI 565
Score = 50.8 bits (120), Expect = 3e-06
Identities = 29/89 (32%), Positives = 48/89 (53%)
Frame = +2
Query: 504 LGVLHLSSNQLTGKLPMEVLGGMKSLFDLKISNNHFSDNIPSEIGLLQRLQELDLGGNEL 563
+ ++L++ L GKL +L L I NN+F IP +IG L R+ L+ N +
Sbjct: 269 ISTINLANYGLKGKLHTLSFSSFPNLLILNIFNNNFYGTIPPQIGNLSRINTLNFSKNPI 448
Query: 564 SGKIPKELVELPNLRMLNLSRNKIEGIIP 592
G IP E+ L +L+ L+ ++ ++ G IP
Sbjct: 449 IGSIPIEMWTLRSLKGLDFAQCQLTGEIP 535
Score = 48.5 bits (114), Expect = 1e-05
Identities = 30/93 (32%), Positives = 45/93 (48%), Gaps = 1/93 (1%)
Frame = +2
Query: 431 SIERITLEVNQIEGDI-AQDFGVYPKLQYLDLSDNKFHGQISPNWGKSLNLQTFIISNNN 489
SI I L ++G + F +P L L++ +N F+G I P G + T S N
Sbjct: 266 SISTINLANYGLKGKLHTLSFSSFPNLLILNIFNNNFYGTIPPQIGNLSRINTLNFSKNP 445
Query: 490 ISGVIPLDFIGLTKLGVLHLSSNQLTGKLPMEV 522
I G IP++ L L L + QLTG++P +
Sbjct: 446 IIGSIPIEMWTLRSLKGLDFAQCQLTGEIPNSI 544
Score = 46.6 bits (109), Expect = 5e-05
Identities = 27/77 (35%), Positives = 44/77 (57%)
Frame = +2
Query: 190 NLLHLAIQKSNLVGSIPQEIGFLTNLAYIDLSKNSLSGGIPETIGNLSKLDTLVLSNNTK 249
NLL L I +N G+IP +IG L+ + ++ SKN + G IP + L L L + +
Sbjct: 341 NLLILNIFNNNFYGTIPPQIGNLSRINTLNFSKNPIIGSIPIEMWTLRSLKGLDFA-QCQ 517
Query: 250 MSGPIPHSLWNMSSLTV 266
++G IP+S+ N+S +
Sbjct: 518 LTGEIPNSIGNLSKTII 568
Score = 44.7 bits (104), Expect = 2e-04
Identities = 25/94 (26%), Positives = 45/94 (47%), Gaps = 1/94 (1%)
Frame = +2
Query: 310 KNLIKLYLGSNNLSGPIPA-SIGNLINLQVLSVQENNLTGTIPASIGNLKWLTVFEVATN 368
K++ + L + L G + S + NL +L++ NN GTIP IGNL + + N
Sbjct: 263 KSISTINLANYGLKGKLHTLSFSSFPNLLILNIFNNNFYGTIPPQIGNLSRINTLNFSKN 442
Query: 369 KLHGRIPNGLYNITNWISFVVSENDFVGHLPSQI 402
+ G IP ++ + + ++ G +P+ I
Sbjct: 443 PIIGSIPIEMWTLRSLKGLDFAQCQLTGEIPNSI 544
Score = 43.9 bits (102), Expect = 3e-04
Identities = 28/77 (36%), Positives = 38/77 (48%)
Frame = +2
Query: 257 SLWNMSSLTVLYFDNIGLSGSIPDSIQNLVNLKELALDINHLSGSIPSTIGDLKNLIKLY 316
S + +L +L N G+IP I NL + L N + GSIP + L++L L
Sbjct: 323 SFSSFPNLLILNIFNNNFYGTIPPQIGNLSRINTLNFSKNPIIGSIPIEMWTLRSLKGLD 502
Query: 317 LGSNNLSGPIPASIGNL 333
L+G IP SIGNL
Sbjct: 503 FAQCQLTGEIPNSIGNL 553
Score = 21.9 bits (45), Expect(2) = 2e-45
Identities = 9/21 (42%), Positives = 13/21 (61%)
Frame = +1
Query: 175 NWSGGPIPPEIGKLNNLLHLA 195
N+ P IGKLN L+H++
Sbjct: 592 NFLAAIFPLAIGKLNQLVHVS 654
>TC90857 weakly similar to GP|16930691|gb|AAL32011.1 AT4g26540/M3E9_30
{Arabidopsis thaliana}, partial (22%)
Length = 829
Score = 180 bits (456), Expect = 3e-45
Identities = 106/277 (38%), Positives = 149/277 (53%)
Frame = +3
Query: 102 NSFYGTIPAQIGNLSNISILTFKNNYFDGSIPQEMCTLTGLQFLDISFCKLNGAIPKSIG 161
NS G+IP ++GNL N+ L N G+IP E+ L +D S + G+IPK+ G
Sbjct: 3 NSLTGSIPTKLGNLKNLKNLLLWQNNLVGTIPSEIGNCYQLSVIDASMNSITGSIPKTFG 182
Query: 162 NLTNLSYLILGGNNWSGGPIPPEIGKLNNLLHLAIQKSNLVGSIPQEIGFLTNLAYIDLS 221
NLT L L L N SG IP E+G L H+ I + + G+IP E+G L NL + L
Sbjct: 183 NLTLLQELQLSVNQISG-EIPAELGNCQQLTHVEIDNNLITGTIPSELGNLGNLTLLFLW 359
Query: 222 KNSLSGGIPETIGNLSKLDTLVLSNNTKMSGPIPHSLWNMSSLTVLYFDNIGLSGSIPDS 281
N L G IP T+ N L+ + LS N ++GPIP ++ + +L L + LSG IP
Sbjct: 360 HNKLQGNIPSTLSNCQNLEAIDLSQNL-LTGPIPKGIFQLQNLNKLLLLSNNLSGKIPSQ 536
Query: 282 IQNLVNLKELALDINHLSGSIPSTIGDLKNLIKLYLGSNNLSGPIPASIGNLINLQVLSV 341
I N +L + N+++G IPS IG+LKNL L LGSN + G IP I NL L +
Sbjct: 537 IGNCSSLIRFRANNNNITGFIPSQIGNLKNLNFLDLGSNRIEGIIPEKISGCRNLTFLDL 716
Query: 342 QENNLTGTIPASIGNLKWLTVFEVATNKLHGRIPNGL 378
N + GT+P S+ L L + + N + G + + L
Sbjct: 717 HSNYIAGTLPDSLSELVSLQFLDFSDNMIEGALKSSL 827
Score = 177 bits (449), Expect = 2e-44
Identities = 108/275 (39%), Positives = 152/275 (55%)
Frame = +3
Query: 152 LNGAIPKSIGNLTNLSYLILGGNNWSGGPIPPEIGKLNNLLHLAIQKSNLVGSIPQEIGF 211
L G+IP +GNL NL L+L NN G IP EIG L + +++ GSIP+ G
Sbjct: 9 LTGSIPTKLGNLKNLKNLLLWQNNLVG-TIPSEIGNCYQLSVIDASMNSITGSIPKTFGN 185
Query: 212 LTNLAYIDLSKNSLSGGIPETIGNLSKLDTLVLSNNTKMSGPIPHSLWNMSSLTVLYFDN 271
LT L + LS N +SG IP +GN +L + + NN ++G IP L N+ +LT+L+ +
Sbjct: 186 LTLLQELQLSVNQISGEIPAELGNCQQLTHVEIDNNL-ITGTIPSELGNLGNLTLLFLWH 362
Query: 272 IGLSGSIPDSIQNLVNLKELALDINHLSGSIPSTIGDLKNLIKLYLGSNNLSGPIPASIG 331
L G+IP ++ N NL+ + L N L+G IP I L+NL KL L SNNLSG IP+ IG
Sbjct: 363 NKLQGNIPSTLSNCQNLEAIDLSQNLLTGPIPKGIFQLQNLNKLLLLSNNLSGKIPSQIG 542
Query: 332 NLINLQVLSVQENNLTGTIPASIGNLKWLTVFEVATNKLHGRIPNGLYNITNWISFVVSE 391
N +L NN+TG IP+ IGNLK L ++ +N++ G IP + N +
Sbjct: 543 NCSSLIRFRANNNNITGFIPSQIGNLKNLNFLDLGSNRIEGIIPEKISGCRNLTFLDLHS 722
Query: 392 NDFVGHLPSQICSGGSLRLLNADHNRFTGPIPTSL 426
N G LP + SL+ L+ N G + +SL
Sbjct: 723 NYIAGTLPDSLSELVSLQFLDFSDNMIEGALKSSL 827
Score = 177 bits (448), Expect = 3e-44
Identities = 104/292 (35%), Positives = 154/292 (52%)
Frame = +3
Query: 179 GPIPPEIGKLNNLLHLAIQKSNLVGSIPQEIGFLTNLAYIDLSKNSLSGGIPETIGNLSK 238
G IP ++G L NL +L + ++NLVG+IP EIG L+ ID S NS++G IP+T GNL+
Sbjct: 15 GSIPTKLGNLKNLKNLLLWQNNLVGTIPSEIGNCYQLSVIDASMNSITGSIPKTFGNLTL 194
Query: 239 LDTLVLSNNTKMSGPIPHSLWNMSSLTVLYFDNIGLSGSIPDSIQNLVNLKELALDINHL 298
L L LS N +SG IP + N L + +D N +
Sbjct: 195 LQELQLSVNQ-------------------------ISGEIPAELGNCQQLTHVEIDNNLI 299
Query: 299 SGSIPSTIGDLKNLIKLYLGSNNLSGPIPASIGNLINLQVLSVQENNLTGTIPASIGNLK 358
+G+IPS +G+L NL L+L N L G IP+++ N NL+ + + +N LTG IP I L+
Sbjct: 300 TGTIPSELGNLGNLTLLFLWHNKLQGNIPSTLSNCQNLEAIDLSQNLLTGPIPKGIFQLQ 479
Query: 359 WLTVFEVATNKLHGRIPNGLYNITNWISFVVSENDFVGHLPSQICSGGSLRLLNADHNRF 418
L + +N L G+IP+ + N ++ I F + N+ G +PSQI + +L L+ NR
Sbjct: 480 NLNKLLLLSNNLSGKIPSQIGNCSSLIRFRANNNNITGFIPSQIGNLKNLNFLDLGSNRI 659
Query: 419 TGPIPTSLKTCSSIERITLEVNQIEGDIAQDFGVYPKLQYLDLSDNKFHGQI 470
G IP + C ++ + L N I G + LQ+LD SDN G +
Sbjct: 660 EGIIPEKISGCRNLTFLDLHSNYIAGTLPDSLSELVSLQFLDFSDNMIEGAL 815
Score = 166 bits (421), Expect = 3e-41
Identities = 98/276 (35%), Positives = 144/276 (51%)
Frame = +3
Query: 296 NHLSGSIPSTIGDLKNLIKLYLGSNNLSGPIPASIGNLINLQVLSVQENNLTGTIPASIG 355
N L+GSIP+ +G+LKNL L L NNL G IP+ IGN L V+ N++TG+IP + G
Sbjct: 3 NSLTGSIPTKLGNLKNLKNLLLWQNNLVGTIPSEIGNCYQLSVIDASMNSITGSIPKTFG 182
Query: 356 NLKWLTVFEVATNKLHGRIPNGLYNITNWISFVVSENDFVGHLPSQICSGGSLRLLNADH 415
NL L +++ N++ G IP L N + N G +PS++ + G+L LL H
Sbjct: 183 NLTLLQELQLSVNQISGEIPAELGNCQQLTHVEIDNNLITGTIPSELGNLGNLTLLFLWH 362
Query: 416 NRFTGPIPTSLKTCSSIERITLEVNQIEGDIAQDFGVYPKLQYLDLSDNKFHGQISPNWG 475
N+ G IP++L C ++E I L N + G I + L L L N G+I G
Sbjct: 363 NKLQGNIPSTLSNCQNLEAIDLSQNLLTGPIPKGIFQLQNLNKLLLLSNNLSGKIPSQIG 542
Query: 476 KSLNLQTFIISNNNISGVIPLDFIGLTKLGVLHLSSNQLTGKLPMEVLGGMKSLFDLKIS 535
+L F +NNNI+G IP L L L L SN++ G +P E + G ++L L +
Sbjct: 543 NCSSLIRFRANNNNITGFIPSQIGNLKNLNFLDLGSNRIEGIIP-EKISGCRNLTFLDLH 719
Query: 536 NNHFSDNIPSEIGLLQRLQELDLGGNELSGKIPKEL 571
+N+ + +P + L LQ LD N + G + L
Sbjct: 720 SNYIAGTLPDSLSELVSLQFLDFSDNMIEGALKSSL 827
Score = 164 bits (414), Expect = 2e-40
Identities = 100/262 (38%), Positives = 147/262 (55%)
Frame = +3
Query: 93 NLLMIDIRNNSFYGTIPAQIGNLSNISILTFKNNYFDGSIPQEMCTLTGLQFLDISFCKL 152
NL + + N+ GTIP++IGN +S++ N GSIP+ LT LQ L +S ++
Sbjct: 48 NLKNLLLWQNNLVGTIPSEIGNCYQLSVIDASMNSITGSIPKTFGNLTLLQELQLSVNQI 227
Query: 153 NGAIPKSIGNLTNLSYLILGGNNWSGGPIPPEIGKLNNLLHLAIQKSNLVGSIPQEIGFL 212
+G IP +GN L+++ + NN G IP E+G L NL L + + L G+IP +
Sbjct: 228 SGEIPAELGNCQQLTHVEI-DNNLITGTIPSELGNLGNLTLLFLWHNKLQGNIPSTLSNC 404
Query: 213 TNLAYIDLSKNSLSGGIPETIGNLSKLDTLVLSNNTKMSGPIPHSLWNMSSLTVLYFDNI 272
NL IDLS+N L+G IP+ I L L+ L+L +N +SG IP + N SSL +N
Sbjct: 405 QNLEAIDLSQNLLTGPIPKGIFQLQNLNKLLLLSN-NLSGKIPSQIGNCSSLIRFRANNN 581
Query: 273 GLSGSIPDSIQNLVNLKELALDINHLSGSIPSTIGDLKNLIKLYLGSNNLSGPIPASIGN 332
++G IP I NL NL L L N + G IP I +NL L L SN ++G +P S+
Sbjct: 582 NITGFIPSQIGNLKNLNFLDLGSNRIEGIIPEKISGCRNLTFLDLHSNYIAGTLPDSLSE 761
Query: 333 LINLQVLSVQENNLTGTIPASI 354
L++LQ L +N + G + +S+
Sbjct: 762 LVSLQFLDFSDNMIEGALKSSL 827
Score = 157 bits (398), Expect = 2e-38
Identities = 101/300 (33%), Positives = 146/300 (48%), Gaps = 2/300 (0%)
Frame = +3
Query: 320 NNLSGPIPASIGNLINLQVLSVQENNLTGTIPASIGNLKWLTVFEVATNKLHGRIPNGLY 379
N+L+G IP +GNL NL+ L + +NNL GTIP+ IGN L+V + + N + G IP
Sbjct: 3 NSLTGSIPTKLGNLKNLKNLLLWQNNLVGTIPSEIGNCYQLSVIDASMNSITGSIPKTFG 182
Query: 380 NITNWISFVVSENDFVGHLPSQICSGGSLRLLNADHNRFTGPIPTSLKTCSSIERITLEV 439
N+T L+ L N+ +G IP L C + + ++
Sbjct: 183 NLT------------------------LLQELQLSVNQISGEIPAELGNCQQLTHVEIDN 290
Query: 440 NQIEGDIAQDFGVYPKLQYLDLSDNKFHGQISPNWGKSLNLQTFIISNNNISGVIPLDFI 499
N I G I + G L L L NK G I NL+ +S N ++G IP
Sbjct: 291 NLITGTIPSELGNLGNLTLLFLWHNKLQGNIPSTLSNCQNLEAIDLSQNLLTGPIPKGIF 470
Query: 500 GLTKLGVLHLSSNQLTGKLPMEVLGGMKSLFDLKISNNHFSDNIPSEIGLLQRLQELDLG 559
L L L L SN L+GK+P ++ G SL + +NN+ + IPS+IG L+ L LDLG
Sbjct: 471 QLQNLNKLLLLSNNLSGKIPSQI-GNCSSLIRFRANNNNITGFIPSQIGNLKNLNFLDLG 647
Query: 560 GNELSGKIPKELVELPNLRMLNLSRNKIEGIIPIKFDS--GLESLDLSGNFLKGNIPTGL 617
N + G IP+++ NL L+L N I G +P L+ LD S N ++G + + L
Sbjct: 648 SNRIEGIIPEKISGCRNLTFLDLHSNYIAGTLPDSLSELVSLQFLDFSDNMIEGALKSSL 827
Score = 153 bits (387), Expect = 3e-37
Identities = 91/272 (33%), Positives = 143/272 (52%), Gaps = 2/272 (0%)
Frame = +3
Query: 368 NKLHGRIPNGLYNITNWISFVVSENDFVGHLPSQICSGGSLRLLNADHNRFTGPIPTSLK 427
N L G IP L N+ N + ++ +N+ VG +PS+I + L +++A N TG IP +
Sbjct: 3 NSLTGSIPTKLGNLKNLKNLLLWQNNLVGTIPSEIGNCYQLSVIDASMNSITGSIPKTFG 182
Query: 428 TCSSIERITLEVNQIEGDIAQDFGVYPKLQYLDLSDNKFHGQISPNWGKSLNLQTFIISN 487
+ ++ + L VNQI G+I + G +L ++++ +N G I G NL + +
Sbjct: 183 NLTLLQELQLSVNQISGEIPAELGNCQQLTHVEIDNNLITGTIPSELGNLGNLTLLFLWH 362
Query: 488 NNISGVIPLDFIGLTKLGVLHLSSNQLTGKLPMEVLGGMKSLFDLKISNNHFSDNIPSEI 547
N + G IP L + LS N LTG +P + +++L L + +N+ S IPS+I
Sbjct: 363 NKLQGNIPSTLSNCQNLEAIDLSQNLLTGPIPKGIFQ-LQNLNKLLLLSNNLSGKIPSQI 539
Query: 548 GLLQRLQELDLGGNELSGKIPKELVELPNLRMLNLSRNKIEGIIPIKFDS--GLESLDLS 605
G L N ++G IP ++ L NL L+L N+IEGIIP K L LDL
Sbjct: 540 GNCSSLIRFRANNNNITGFIPSQIGNLKNLNFLDLGSNRIEGIIPEKISGCRNLTFLDLH 719
Query: 606 GNFLKGNIPTGLADLVRLSKLNLSHNMLSGTI 637
N++ G +P L++LV L L+ S NM+ G +
Sbjct: 720 SNYIAGTLPDSLSELVSLQFLDFSDNMIEGAL 815
Score = 150 bits (379), Expect = 3e-36
Identities = 100/296 (33%), Positives = 136/296 (45%)
Frame = +3
Query: 223 NSLSGGIPETIGNLSKLDTLVLSNNTKMSGPIPHSLWNMSSLTVLYFDNIGLSGSIPDSI 282
NSL+G IP +GNL L L+L N L G+IP I
Sbjct: 3 NSLTGSIPTKLGNLKNLKNLLLWQNN-------------------------LVGTIPSEI 107
Query: 283 QNLVNLKELALDINHLSGSIPSTIGDLKNLIKLYLGSNNLSGPIPASIGNLINLQVLSVQ 342
N L + +N ++GSIP T G+L L +L L N +SG IPA +GN L + +
Sbjct: 108 GNCYQLSVIDASMNSITGSIPKTFGNLTLLQELQLSVNQISGEIPAELGNCQQLTHVEID 287
Query: 343 ENNLTGTIPASIGNLKWLTVFEVATNKLHGRIPNGLYNITNWISFVVSENDFVGHLPSQI 402
N +TGTIP+ +GNL LT+ + NKL G IP+ L N N + +S+N G +P I
Sbjct: 288 NNLITGTIPSELGNLGNLTLLFLWHNKLQGNIPSTLSNCQNLEAIDLSQNLLTGPIPKGI 467
Query: 403 CSGGSLRLLNADHNRFTGPIPTSLKTCSSIERITLEVNQIEGDIAQDFGVYPKLQYLDLS 462
+L L N +G IP+ + CSS+ R N I G I G L +LDL
Sbjct: 468 FQLQNLNKLLLLSNNLSGKIPSQIGNCSSLIRFRANNNNITGFIPSQIGNLKNLNFLDLG 647
Query: 463 DNKFHGQISPNWGKSLNLQTFIISNNNISGVIPLDFIGLTKLGVLHLSSNQLTGKL 518
N+ G I NL + +N I+G +P L L L S N + G L
Sbjct: 648 SNRIEGIIPEKISGCRNLTFLDLHSNYIAGTLPDSLSELVSLQFLDFSDNMIEGAL 815
Score = 123 bits (309), Expect = 3e-28
Identities = 85/263 (32%), Positives = 135/263 (51%)
Frame = +3
Query: 44 LSTWKNNTNPCKPKWRGIKCDKSNFISTIGLANLGLKGTLHSLTFSSFPNLLMIDIRNNS 103
L W+NN P G C + +S I + + G++ TF + L + + N
Sbjct: 60 LLLWQNNLVGTIPSEIG-NCYQ---LSVIDASMNSITGSIPK-TFGNLTLLQELQLSVNQ 224
Query: 104 FYGTIPAQIGNLSNISILTFKNNYFDGSIPQEMCTLTGLQFLDISFCKLNGAIPKSIGNL 163
G IPA++GN ++ + NN G+IP E+ L L L + KL G IP ++ N
Sbjct: 225 ISGEIPAELGNCQQLTHVEIDNNLITGTIPSELGNLGNLTLLFLWHNKLQGNIPSTLSNC 404
Query: 164 TNLSYLILGGNNWSGGPIPPEIGKLNNLLHLAIQKSNLVGSIPQEIGFLTNLAYIDLSKN 223
NL + L N GPIP I +L NL L + +NL G IP +IG ++L + N
Sbjct: 405 QNLEAIDL-SQNLLTGPIPKGIFQLQNLNKLLLLSNNLSGKIPSQIGNCSSLIRFRANNN 581
Query: 224 SLSGGIPETIGNLSKLDTLVLSNNTKMSGPIPHSLWNMSSLTVLYFDNIGLSGSIPDSIQ 283
+++G IP IGNL L+ L L +N ++ G IP + +LT L + ++G++PDS+
Sbjct: 582 NITGFIPSQIGNLKNLNFLDLGSN-RIEGIIPEKISGCRNLTFLDLHSNYIAGTLPDSLS 758
Query: 284 NLVNLKELALDINHLSGSIPSTI 306
LV+L+ L N + G++ S++
Sbjct: 759 ELVSLQFLDFSDNMIEGALKSSL 827
Score = 123 bits (308), Expect = 4e-28
Identities = 82/249 (32%), Positives = 126/249 (49%), Gaps = 29/249 (11%)
Frame = +3
Query: 464 NKFHGQISPNWGKSLNLQTFIISNNNISGVIPLDFIGLTKLGVLHLSSNQLTGKLPMEVL 523
N G I G NL+ ++ NN+ G IP + +L V+ S N +TG +P +
Sbjct: 3 NSLTGSIPTKLGNLKNLKNLLLWQNNLVGTIPSEIGNCYQLSVIDASMNSITGSIP-KTF 179
Query: 524 GGMKSLFDLKISNNHFSDNIPSEIGLLQRLQELDLGGNELSGKIPKELVELPNLRMLNLS 583
G + L +L++S N S IP+E+G Q+L +++ N ++G IP EL L NL +L L
Sbjct: 180 GNLTLLQELQLSVNQISGEIPAELGNCQQLTHVEIDNNLITGTIPSELGNLGNLTLLFLW 359
Query: 584 RNKIEGIIPIKFDS--GLESLDLSGNFLKGNIPTGLADLVRLSKLNLSHNMLSGTIPQNF 641
NK++G IP + LE++DLS N L G IP G+ L L+KL L N LSG IP
Sbjct: 360 HNKLQGNIPSTLSNCQNLEAIDLSQNLLTGPIPKGIFQLQNLNKLLLLSNNLSGKIPSQI 539
Query: 642 G--------------------------RNLVFVNISDNQLEGPLP-KIPAFLSASFESLK 674
G +NL F+++ N++EG +P KI + +F L
Sbjct: 540 GNCSSLIRFRANNNNITGFIPSQIGNLKNLNFLDLGSNRIEGIIPEKISGCRNLTFLDL- 716
Query: 675 NNNHLCGNI 683
++N++ G +
Sbjct: 717 HSNYIAGTL 743
Score = 64.7 bits (156), Expect = 2e-10
Identities = 48/168 (28%), Positives = 77/168 (45%)
Frame = +3
Query: 42 TLLSTWKNNTNPCKPKWRGIKCDKSNFISTIGLANLGLKGTLHSLTFSSFPNLLMIDIRN 101
TLL W N P C + I L+ L G + F NL + + +
Sbjct: 342 TLLFLWHNKLQGNIPSTLS-NCQN---LEAIDLSQNLLTGPIPKGIFQ-LQNLNKLLLLS 506
Query: 102 NSFYGTIPAQIGNLSNISILTFKNNYFDGSIPQEMCTLTGLQFLDISFCKLNGAIPKSIG 161
N+ G IP+QIGN S++ NN G IP ++ L L FLD+ ++ G IP+ I
Sbjct: 507 NNLSGKIPSQIGNCSSLIRFRANNNNITGFIPSQIGNLKNLNFLDLGSNRIEGIIPEKIS 686
Query: 162 NLTNLSYLILGGNNWSGGPIPPEIGKLNNLLHLAIQKSNLVGSIPQEI 209
NL++L L +N+ G +P + +L +L L + + G++ +
Sbjct: 687 GCRNLTFLDL-HSNYIAGTLPDSLSELVSLQFLDFSDNMIEGALKSSL 827
>TC92229 similar to PIR|B86369|B86369 hypothetical protein AAC98010.1
[imported] - Arabidopsis thaliana, partial (44%)
Length = 1347
Score = 178 bits (452), Expect = 9e-45
Identities = 107/321 (33%), Positives = 171/321 (52%), Gaps = 7/321 (2%)
Frame = +2
Query: 729 KKPNEESQTEEVQRGVLFSIWSHDGKMMFENIIEATANFDDKYLVGVGSQGNVYKAELSE 788
+ P+E + + +LFS ++ I+E T F + ++G G G VYKA + +
Sbjct: 107 RSPSEATPPQMSGGQILFS---------YDQILEITNGFSSENVIGEGGFGRVYKALMPD 259
Query: 789 GLVVAVKKLHLVTDEEMSCFSSKSFMSEIETLTGIKHRNIIKLHGFCSHSKFSFLVYKFL 848
G V A+K L + + + F +E++T++ + HR+++ L G+C + L+Y+F+
Sbjct: 260 GRVGALKLLKAGSGQ-----GEREFRAEVDTISRVHHRHLVSLIGYCIAEQQRVLIYEFV 424
Query: 849 EGGSLDQILNNDTQAVAFDWEKRVNVVKGVANALSYLHHDCSPPIIHRDISSKNVLLNLD 908
G+LDQ L+ ++Q DW KR+ + G A L+YLH C+P IIHRDI S N+LL+
Sbjct: 425 PNGNLDQHLH-ESQWNVLDWPKRMKIAIGAARGLAYLHEGCNPKIIHRDIKSSNILLDDS 601
Query: 909 YEAHVSDFGTAKFL-KPGLHSWTQFAGTFGYAAPELAQTMEVNEKCDVYSFGVLALETIM 967
YEA V+DFG A+ H T+ GTFGY APE A + ++ ++ DV+SFGV+ LE +
Sbjct: 602 YEAQVADFGLARLTDDTNTHVSTRVMGTFGYMAPEYATSGKLTDRSDVFSFGVVLLELVT 781
Query: 968 GKHPGDLISLFLSPSTRPMANNMLLTDVLDQRPQQVMEP------IDEEVILIARLAFAC 1021
G+ P D S A +LL + ++ +P ID E+ + A AC
Sbjct: 782 GRKPVDPTQPVGDESLVEWARPILLRAIETGDFSELADPRLHRQYIDSEMFRMIEAAAAC 961
Query: 1022 LSQNPRLRPSMGQVCKMLAIG 1042
+ + RP M Q+ + L G
Sbjct: 962 IRHSAPKRPRMVQIARALDSG 1024
>TC88105 similar to GP|21593085|gb|AAM65034.1 Putative protein kinase
{Arabidopsis thaliana}, partial (72%)
Length = 2379
Score = 176 bits (447), Expect = 3e-44
Identities = 103/290 (35%), Positives = 154/290 (52%), Gaps = 6/290 (2%)
Frame = +2
Query: 763 ATANFDDKYLVGVGSQGNVYKAELSEGLVVAVKKLHLVTDEEMSCFSSKSFMSEIETLTG 822
AT F ++G G G VY+ +L G VA+KKL + + K F E+E +
Sbjct: 713 ATNKFSKDNIIGEGGYGVVYQGQLINGNPVAIKKLLNNLGQ-----AEKEFRVEVEAIGH 877
Query: 823 IKHRNIIKLHGFCSHSKFSFLVYKFLEGGSLDQILNNDTQAVAF-DWEKRVNVVKGVANA 881
++H+N+++L GFC L+Y+++ G+L+Q L+ + + W+ R+ ++ G A A
Sbjct: 878 VRHKNLVRLLGFCIEGTHRLLIYEYVNNGNLEQWLHGAMRQYGYLTWDARIKILLGTAKA 1057
Query: 882 LSYLHHDCSPPIIHRDISSKNVLLNLDYEAHVSDFGTAKFLKPG-LHSWTQFAGTFGYAA 940
L+YLH P ++HRDI S N+L++ D+ A +SDFG AK L G H T+ GTFGY A
Sbjct: 1058 LAYLHEAIEPKVVHRDIKSSNILIDDDFNAKISDFGLAKLLGAGKSHITTRVMGTFGYVA 1237
Query: 941 PELAQTMEVNEKCDVYSFGVLALETIMGKHPGD----LISLFLSPSTRPMANNMLLTDVL 996
PE A + +NEK DVYSFGVL LE I G+ P D + L + M N +V+
Sbjct: 1238 PEYANSGLLNEKSDVYSFGVLLLEAITGRDPVDYNRSAAEVNLVDWLKMMVGNRHAEEVV 1417
Query: 997 DQRPQQVMEPIDEEVILIARLAFACLSQNPRLRPSMGQVCKMLAIGKSPL 1046
D P P + + A C+ + RP M QV +ML + P+
Sbjct: 1418 D--PNIETRPSTSALKRVLLTALRCVDPDSEKRPKMSQVVRMLESEEYPI 1561
>TC90678 weakly similar to GP|9759550|dbj|BAB11152.1 receptor protein
kinase-like protein {Arabidopsis thaliana}, partial
(48%)
Length = 1392
Score = 135 bits (340), Expect = 8e-32
Identities = 91/292 (31%), Positives = 139/292 (47%), Gaps = 1/292 (0%)
Frame = +2
Query: 229 IPETIGNLSKLDTLVLSNNTKMSGPIPHSLWNMSSLTVLYFDNIGLSGSIPDSIQNLVNL 288
+P GNL+KL+ L LS N P+ L IP+ + L NL
Sbjct: 566 VPNVFGNLTKLEVLDLSMN-------PY-----------------LVSEIPEDVGELGNL 673
Query: 289 KELALDINHLSGSIPSTIGDLKNLIKLYLGSNNLSGPIPAS-IGNLINLQVLSVQENNLT 347
K+L L + G +P ++ L +L L L NNL+G + + + +L+NL V +N L
Sbjct: 674 KQLLLQGSSFQGEVPESLKGLISLTHLDLSENNLTGEVSKTLVSSLMNLVSFDVSQNKLL 853
Query: 348 GTIPASIGNLKWLTVFEVATNKLHGRIPNGLYNITNWISFVVSENDFVGHLPSQICSGGS 407
G+ P + K L + TN+ G IPN + F V N F G P + S
Sbjct: 854 GSFPNGLCKGKGLINLSLHTNRFTGLIPNSTSECKSLERFQVQNNGFSGDFPIVLFSLPK 1033
Query: 408 LRLLNADHNRFTGPIPTSLKTCSSIERITLEVNQIEGDIAQDFGVYPKLQYLDLSDNKFH 467
++L+ ++NRFTG IP S+ +E++ L+ N ++G I G L S N F+
Sbjct: 1034IKLIRGENNRFTGKIPESISEAVQLEQVQLDNNLLDGKIPSGLGFVKSLYRFSASLNHFY 1213
Query: 468 GQISPNWGKSLNLQTFIISNNNISGVIPLDFIGLTKLGVLHLSSNQLTGKLP 519
G++ PN+ S + +S+N++SG IP KL L L+ N LTG +P
Sbjct: 1214GELPPNFCDSPVMSIVNLSHNSLSGSIP-QLKKCKKLVSLSLADNSLTGXIP 1366
Score = 125 bits (315), Expect = 7e-29
Identities = 80/276 (28%), Positives = 138/276 (49%), Gaps = 1/276 (0%)
Frame = +2
Query: 104 FYGTIPAQIGNLSNISILTFKNN-YFDGSIPQEMCTLTGLQFLDISFCKLNGAIPKSIGN 162
F +P GNL+ + +L N Y IP+++ L L+ L + G +P+S+
Sbjct: 554 FLVMVPNVFGNLTKLEVLDLSMNPYLVSEIPEDVGELGNLKQLLLQGSSFQGEVPESLKG 733
Query: 163 LTNLSYLILGGNNWSGGPIPPEIGKLNNLLHLAIQKSNLVGSIPQEIGFLTNLAYIDLSK 222
L +L++L L NN +G + L NL+ + ++ L+GS P + L + L
Sbjct: 734 LISLTHLDLSENNLTGEVSKTLVSSLMNLVSFDVSQNKLLGSFPNGLCKGKGLINLSLHT 913
Query: 223 NSLSGGIPETIGNLSKLDTLVLSNNTKMSGPIPHSLWNMSSLTVLYFDNIGLSGSIPDSI 282
N +G IP + L+ + NN SG P L+++ + ++ +N +G IP+SI
Sbjct: 914 NRFTGLIPNSTSECKSLERFQVQNNG-FSGDFPIVLFSLPKIKLIRGENNRFTGKIPESI 1090
Query: 283 QNLVNLKELALDINHLSGSIPSTIGDLKNLIKLYLGSNNLSGPIPASIGNLINLQVLSVQ 342
V L+++ LD N L G IPS +G +K+L + N+ G +P + + + ++++
Sbjct: 1091SEAVQLEQVQLDNNLLDGKIPSGLGFVKSLYRFSASLNHFYGELPPNFCDSPVMSIVNLS 1270
Query: 343 ENNLTGTIPASIGNLKWLTVFEVATNKLHGRIPNGL 378
N+L+G+IP + K L +A N L G IPN L
Sbjct: 1271HNSLSGSIP-QLKKCKKLVSLSLADNSLTGXIPNSL 1375
Score = 112 bits (279), Expect = 1e-24
Identities = 89/292 (30%), Positives = 134/292 (45%), Gaps = 5/292 (1%)
Frame = +2
Query: 374 IPNGLYNITNWISFVVSENDF-VGHLPSQICSGGSLRLLNADHNRFTGPIPTSLKTCSSI 432
+PN N+T +S N + V +P + G+L+ L + F G +P SLK S
Sbjct: 566 VPNVFGNLTKLEVLDLSMNPYLVSEIPEDVGELGNLKQLLLQGSSFQGEVPESLKGLIS- 742
Query: 433 ERITLEVNQIEGDIAQDFGVYPKLQYLDLSDNKFHGQISPNWGKSL-NLQTFIISNNNIS 491
L +LDLS+N G++S SL NL +F +S N +
Sbjct: 743 -----------------------LTHLDLSENNLTGEVSKTLVSSLMNLVSFDVSQNKLL 853
Query: 492 GVIPLDFIGLTKLGVLHLSSNQLTGKLPMEVLGGMKSLFDLKISNNHFSDNIPSEIGLLQ 551
G P L L L +N+ TG +P KSL ++ NN FS + P + L
Sbjct: 854 GSFPNGLCKGKGLINLSLHTNRFTGLIPNST-SECKSLERFQVQNNGFSGDFPIVLFSLP 1030
Query: 552 RLQELDLGGNELSGKIPKELVELPNLRMLNLSRNKIEGIIP--IKFDSGLESLDLSGNFL 609
+++ + N +GKIP+ + E L + L N ++G IP + F L S N
Sbjct: 1031KIKLIRGENNRFTGKIPESISEAVQLEQVQLDNNLLDGKIPSGLGFVKSLYRFSASLNHF 1210
Query: 610 KGNIPTGLADLVRLSKLNLSHNMLSGTIPQ-NFGRNLVFVNISDNQLEGPLP 660
G +P D +S +NLSHN LSG+IPQ + LV ++++DN L G +P
Sbjct: 1211YGELPPNFCDSPVMSIVNLSHNSLSGSIPQLKKCKKLVSLSLADNSLTGXIP 1366
Score = 109 bits (272), Expect(2) = 2e-36
Identities = 79/275 (28%), Positives = 130/275 (46%), Gaps = 4/275 (1%)
Frame = +2
Query: 350 IPASIGNLKWLTVFEVATNK-LHGRIPNGLYNITNWISFVVSENDFVGHLPSQICSGGSL 408
+P GNL L V +++ N L IP + + N ++ + F G +P + SL
Sbjct: 566 VPNVFGNLTKLEVLDLSMNPYLVSEIPEDVGELGNLKQLLLQGSSFQGEVPESLKGLISL 745
Query: 409 RLLNADHNRFTGPIPTSLKTCSSIERITLEV--NQIEGDIAQDFGVYPKLQYLDLSDNKF 466
L+ N TG + +L + S + ++ +V N++ G L L L N+F
Sbjct: 746 THLDLSENNLTGEVSKTLVS-SLMNLVSFDVSQNKLLGSFPNGLCKGKGLINLSLHTNRF 922
Query: 467 HGQISPNWGKSLNLQTFIISNNNISGVIPLDFIGLTKLGVLHLSSNQLTGKLPMEVLGGM 526
G I + + +L+ F + NN SG P+ L K+ ++ +N+ TGK+P E +
Sbjct: 923 TGLIPNSTSECKSLERFQVQNNGFSGDFPIVLFSLPKIKLIRGENNRFTGKIP-ESISEA 1099
Query: 527 KSLFDLKISNNHFSDNIPSEIGLLQRLQELDLGGNELSGKIPKELVELPNLRMLNLSRNK 586
L +++ NN IPS +G ++ L N G++P + P + ++NLS N
Sbjct: 1100VQLEQVQLDNNLLDGKIPSGLGFVKSLYRFSASLNHFYGELPPNFCDSPVMSIVNLSHNS 1279
Query: 587 IEGIIP-IKFDSGLESLDLSGNFLKGNIPTGLADL 620
+ G IP +K L SL L+ N L G IP LA+L
Sbjct: 1280LSGSIPQLKKCKKLVSLSLADNSLTGXIPNSLAEL 1384
Score = 65.5 bits (158), Expect = 1e-10
Identities = 58/207 (28%), Positives = 91/207 (43%), Gaps = 5/207 (2%)
Frame = +2
Query: 35 SFDDQSQTLLSTWKNNTNPCKPKWRGIKCDKSNFISTIGLANLGLK-----GTLHSLTFS 89
SFD LL ++ N CK K GL NL L G + + T S
Sbjct: 824 SFDVSQNKLLGSFPNGL--CKGK---------------GLINLSLHTNRFTGLIPNST-S 949
Query: 90 SFPNLLMIDIRNNSFYGTIPAQIGNLSNISILTFKNNYFDGSIPQEMCTLTGLQFLDISF 149
+L ++NN F G P + +L I ++ +NN F G IP+ + L+ + +
Sbjct: 950 ECKSLERFQVQNNGFSGDFPIVLFSLPKIKLIRGENNRFTGKIPESISEAVQLEQVQLDN 1129
Query: 150 CKLNGAIPKSIGNLTNLSYLILGGNNWSGGPIPPEIGKLNNLLHLAIQKSNLVGSIPQEI 209
L+G IP +G + +L Y N G +PP + + + ++L GSIPQ +
Sbjct: 1130 NLLDGKIPSGLGFVKSL-YRFSASLNHFYGELPPNFCDSPVMSIVNLSHNSLSGSIPQ-L 1303
Query: 210 GFLTNLAYIDLSKNSLSGGIPETIGNL 236
L + L+ NSL+G IP ++ L
Sbjct: 1304 KKCKKLVSLSLADNSLTGXIPNSLAEL 1384
Score = 65.1 bits (157), Expect = 1e-10
Identities = 49/149 (32%), Positives = 77/149 (50%), Gaps = 3/149 (2%)
Frame = +3
Query: 238 KLDTLVLSNNTKMSGPIPHSLW-NMSSLTVLYFDNIGLSGSIP-DSIQ-NLVNLKELALD 294
++DTL+ +T S W N SS + I S + P DS+ VNL+ L
Sbjct: 90 EVDTLLSFKSTIQDSKKALSTWSNTSSNHFCNWTGISCSSTTPSDSLSVTSVNLQSL--- 260
Query: 295 INHLSGSIPSTIGDLKNLIKLYLGSNNLSGPIPASIGNLINLQVLSVQENNLTGTIPASI 354
+LSG I S+I DL +L L L +N + PIP + +L+ L++ N + GTIP+ I
Sbjct: 261 --NLSGDISSSICDLPSLSYLNLANNIFNQPIPLHLSQCSSLKSLNLSNNLIWGTIPSQI 434
Query: 355 GNLKWLTVFEVATNKLHGRIPNGLYNITN 383
L+V +++ N + G IP+ L ++ N
Sbjct: 435 SQFVSLSVLDLSRNHIEGNIPDSLGSLKN 521
Score = 62.8 bits (151), Expect = 7e-10
Identities = 42/118 (35%), Positives = 61/118 (51%), Gaps = 10/118 (8%)
Frame = +3
Query: 535 SNNHF--------SDNIPSEIGLLQRLQELDLGGNELSGKIPKELVELPNLRMLNLSRNK 586
S+NHF S PS+ + ++L LSG I + +LP+L LNL+ N
Sbjct: 165 SSNHFCNWTGISCSSTTPSDS---LSVTSVNLQSLNLSGDISSSICDLPSLSYLNLANNI 335
Query: 587 IEGIIPIKFD--SGLESLDLSGNFLKGNIPTGLADLVRLSKLNLSHNMLSGTIPQNFG 642
IP+ S L+SL+LS N + G IP+ ++ V LS L+LS N + G IP + G
Sbjct: 336 FNQPIPLHLSQCSSLKSLNLSNNLIWGTIPSQISQFVSLSVLDLSRNHIEGNIPDSLG 509
Score = 62.8 bits (151), Expect(2) = 2e-36
Identities = 39/95 (41%), Positives = 53/95 (55%)
Frame = +3
Query: 218 IDLSKNSLSGGIPETIGNLSKLDTLVLSNNTKMSGPIPHSLWNMSSLTVLYFDNIGLSGS 277
++L +LSG I +I +L L L L+NN + PIP L SSL L N + G+
Sbjct: 243 VNLQSLNLSGDISSSICDLPSLSYLNLANNI-FNQPIPLHLSQCSSLKSLNLSNNLIWGT 419
Query: 278 IPDSIQNLVNLKELALDINHLSGSIPSTIGDLKNL 312
IP I V+L L L NH+ G+IP ++G LKNL
Sbjct: 420 IPSQISQFVSLSVLDLSRNHIEGNIPDSLGSLKNL 524
Score = 60.8 bits (146), Expect = 3e-09
Identities = 35/107 (32%), Positives = 62/107 (57%)
Frame = +3
Query: 133 PQEMCTLTGLQFLDISFCKLNGAIPKSIGNLTNLSYLILGGNNWSGGPIPPEIGKLNNLL 192
P + ++T + ++ L+G I SI +L +LSYL L NN PIP + + ++L
Sbjct: 216 PSDSLSVTSVNLQSLN---LSGDISSSICDLPSLSYLNLA-NNIFNQPIPLHLSQCSSLK 383
Query: 193 HLAIQKSNLVGSIPQEIGFLTNLAYIDLSKNSLSGGIPETIGNLSKL 239
L + + + G+IP +I +L+ +DLS+N + G IP+++G+L L
Sbjct: 384 SLNLSNNLIWGTIPSQISQFVSLSVLDLSRNHIEGNIPDSLGSLKNL 524
Score = 58.5 bits (140), Expect = 1e-08
Identities = 38/142 (26%), Positives = 64/142 (44%), Gaps = 28/142 (19%)
Frame = +3
Query: 29 LLKWKDSFDDQSQTLLSTWKNNTNPCKPKWRGIKCDKSN-----FISTIGLANLGLKGTL 83
LL +K + D S+ LSTW N ++ W GI C + ++++ L +L L G +
Sbjct: 102 LLSFKSTIQD-SKKALSTWSNTSSNHFCNWTGISCSSTTPSDSLSVTSVNLQSLNLSGDI 278
Query: 84 HS-----------------------LTFSSFPNLLMIDIRNNSFYGTIPAQIGNLSNISI 120
S L S +L +++ NN +GTIP+QI ++S+
Sbjct: 279 SSSICDLPSLSYLNLANNIFNQPIPLHLSQCSSLKSLNLSNNLIWGTIPSQISQFVSLSV 458
Query: 121 LTFKNNYFDGSIPQEMCTLTGL 142
L N+ +G+IP + +L L
Sbjct: 459 LDLSRNHIEGNIPDSLGSLKNL 524
Score = 56.2 bits (134), Expect = 6e-08
Identities = 33/85 (38%), Positives = 40/85 (46%)
Frame = +3
Query: 396 GHLPSQICSGGSLRLLNADHNRFTGPIPTSLKTCSSIERITLEVNQIEGDIAQDFGVYPK 455
G + S IC SL LN +N F PIP L CSS++ + L N I G I +
Sbjct: 270 GDISSSICDLPSLSYLNLANNIFNQPIPLHLSQCSSLKSLNLSNNLIWGTIPSQISQFVS 449
Query: 456 LQYLDLSDNKFHGQISPNWGKSLNL 480
L LDLS N G I + G NL
Sbjct: 450 LSVLDLSRNHIEGNIPDSLGSLKNL 524
Score = 54.3 bits (129), Expect = 2e-07
Identities = 34/102 (33%), Positives = 55/102 (53%)
Frame = +3
Query: 491 SGVIPLDFIGLTKLGVLHLSSNQLTGKLPMEVLGGMKSLFDLKISNNHFSDNIPSEIGLL 550
S P D + +T + +L S L+G + + + SL L ++NN F+ IP +
Sbjct: 204 SSTTPSDSLSVTSV---NLQSLNLSGDISSSICD-LPSLSYLNLANNIFNQPIPLHLSQC 371
Query: 551 QRLQELDLGGNELSGKIPKELVELPNLRMLNLSRNKIEGIIP 592
L+ L+L N + G IP ++ + +L +L+LSRN IEG IP
Sbjct: 372 SSLKSLNLSNNLIWGTIPSQISQFVSLSVLDLSRNHIEGNIP 497
Score = 53.5 bits (127), Expect = 4e-07
Identities = 36/98 (36%), Positives = 50/98 (50%), Gaps = 2/98 (2%)
Frame = +3
Query: 528 SLFDLKISNNHFSDNIPSEIGLLQRLQELDLGGNELSGKIPKELVELPNLRMLNLSRNKI 587
S+ + + + + S +I S I L L L+L N + IP L + +L+ LNLS N I
Sbjct: 231 SVTSVNLQSLNLSGDISSSICDLPSLSYLNLANNIFNQPIPLHLSQCSSLKSLNLSNNLI 410
Query: 588 EGIIPIKFDS--GLESLDLSGNFLKGNIPTGLADLVRL 623
G IP + L LDLS N ++GNIP L L L
Sbjct: 411 WGTIPSQISQFVSLSVLDLSRNHIEGNIPDSLGSLKNL 524
Score = 52.8 bits (125), Expect = 7e-07
Identities = 35/101 (34%), Positives = 49/101 (47%)
Frame = +3
Query: 477 SLNLQTFIISNNNISGVIPLDFIGLTKLGVLHLSSNQLTGKLPMEVLGGMKSLFDLKISN 536
SL++ + + + N+SG I L L L+L++N +P+ L SL L +SN
Sbjct: 225 SLSVTSVNLQSLNLSGDISSSICDLPSLSYLNLANNIFNQPIPLH-LSQCSSLKSLNLSN 401
Query: 537 NHFSDNIPSEIGLLQRLQELDLGGNELSGKIPKELVELPNL 577
N IPS+I L LDL N + G IP L L NL
Sbjct: 402 NLIWGTIPSQISQFVSLSVLDLSRNHIEGNIPDSLGSLKNL 524
Score = 47.0 bits (110), Expect = 4e-05
Identities = 35/102 (34%), Positives = 51/102 (49%), Gaps = 2/102 (1%)
Frame = +3
Query: 576 NLRMLNLSRNKIEGIIPIKFDSGLESLDLSGNFLKGNIPTGLADLVRLSKLNLSHNMLSG 635
NL+ LNLS + I + L L+L+ N IP L+ L LNLS+N++ G
Sbjct: 246 NLQSLNLSGDISSSICDLP---SLSYLNLANNIFNQPIPLHLSQCSSLKSLNLSNNLIWG 416
Query: 636 TIPQNFGR--NLVFVNISDNQLEGPLPKIPAFLSASFESLKN 675
TIP + +L +++S N +EG +P S SLKN
Sbjct: 417 TIPSQISQFVSLSVLDLSRNHIEGNIPD-------SLGSLKN 521
Score = 46.6 bits (109), Expect = 5e-05
Identities = 34/123 (27%), Positives = 53/123 (42%)
Frame = +3
Query: 431 SIERITLEVNQIEGDIAQDFGVYPKLQYLDLSDNKFHGQISPNWGKSLNLQTFIISNNNI 490
S+ + L+ + GDI+ P L YL+L++N F+ I
Sbjct: 231 SVTSVNLQSLNLSGDISSSICDLPSLSYLNLANNIFNQPI-------------------- 350
Query: 491 SGVIPLDFIGLTKLGVLHLSSNQLTGKLPMEVLGGMKSLFDLKISNNHFSDNIPSEIGLL 550
PL + L L+LS+N + G +P ++ SL L +S NH NIP +G L
Sbjct: 351 ----PLHLSQCSSLKSLNLSNNLIWGTIPSQI-SQFVSLSVLDLSRNHIEGNIPDSLGSL 515
Query: 551 QRL 553
+ L
Sbjct: 516 KNL 524
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.319 0.138 0.404
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 33,706,968
Number of Sequences: 36976
Number of extensions: 490301
Number of successful extensions: 6922
Number of sequences better than 10.0: 854
Number of HSP's better than 10.0 without gapping: 3266
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 4959
length of query: 1067
length of database: 9,014,727
effective HSP length: 106
effective length of query: 961
effective length of database: 5,095,271
effective search space: 4896555431
effective search space used: 4896555431
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 63 (28.9 bits)
Medicago: description of AC133779.11


