

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC133779.1 + phase: 0 /pseudo
(611 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
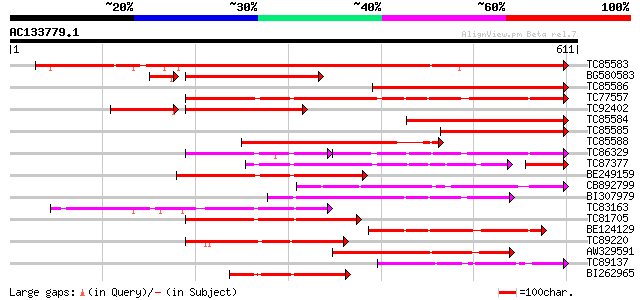
Score E
Sequences producing significant alignments: (bits) Value
TC85583 similar to PIR|T51335|T51335 subtilisin-like proteinase ... 680 0.0
BG580583 weakly similar to GP|9757901|dbj serine protease-like p... 298 2e-91
TC85586 similar to GP|20521377|dbj|BAB91889. putative subtilisin... 311 5e-85
TC77557 similar to PIR|JC7519|JC7519 subtilisin-like serine prot... 303 1e-82
TC92402 similar to GP|20198252|gb|AAM15483.1 subtilisin-like ser... 213 4e-77
TC85584 weakly similar to PIR|A84473|A84473 probable serine prot... 275 3e-74
TC85585 similar to GP|22773236|gb|AAN06842.1 Putatvie subtilisin... 273 1e-73
TC85588 similar to GP|20198169|gb|AAM15440.1 subtilisin-like ser... 270 2e-72
TC86329 similar to PIR|T07171|T07171 subtilisin-like proteinase ... 166 4e-70
TC87377 weakly similar to PIR|T07171|T07171 subtilisin-like prot... 186 2e-59
BE249159 weakly similar to GP|20198252|gb subtilisin-like serine... 213 2e-55
CB892799 weakly similar to PIR|T05768|T05 subtilisin-like protei... 192 3e-49
BI307979 weakly similar to GP|20521377|db putative subtilisin-li... 170 1e-42
TC83163 weakly similar to GP|6721520|dbj|BAA89562.1 EST AU029428... 160 2e-39
TC81705 weakly similar to GP|6721520|dbj|BAA89562.1 EST AU029428... 158 5e-39
BE124129 weakly similar to GP|9757901|dbj serine protease-like p... 151 8e-37
TC89220 similar to PIR|T05768|T05768 subtilisin-like proteinase ... 147 9e-36
AW329591 weakly similar to GP|15010692|gb| At2g04160/T16B23.1 {A... 141 6e-34
TC89137 similar to GP|4200338|emb|CAA76726.1 P69C protein {Lycop... 136 2e-32
BI262965 similar to PIR|A84473|A84 probable serine proteinase [i... 129 3e-30
>TC85583 similar to PIR|T51335|T51335 subtilisin-like proteinase AIR3
auxin-induced [imported] - Arabidopsis thaliana
(fragment), partial (40%)
Length = 2691
Score = 680 bits (1754), Expect = 0.0
Identities = 381/624 (61%), Positives = 439/624 (70%), Gaps = 50/624 (8%)
Frame = +2
Query: 29 NFISFWFLVSYLFL------------QCYIVYLGAHVHGPTPSSVDLETATYSHYDLLGS 76
+F S +V+ LFL +CYIVYLGAH HGP P+S++LE AT SHYDLL S
Sbjct: 38 SFFSLHTMVTLLFLFMFLLETVHGTKKCYIVYLGAHSHGPRPTSLELEIATNSHYDLLSS 217
Query: 77 ILGSHEEAEEAIIYSYNKQINGFAAILEEEEAAQLANCTQHVLGNSLD*VQMMSTQ---- 132
LGS E+A+EAIIYSYNK INGFAA+LE+EEAA +A ++V+ L + T
Sbjct: 218 TLGSREKAKEAIIYSYNKHINGFAALLEDEEAADIAK-KRNVVSVFLSKPHKLHTTRSWE 394
Query: 133 -LGKREGLVKIPL*LTLIQV--FGRNRRVLTTEE*V-----QFH*DGVGILLRKFH---- 180
LG R T Q FG N + + V F+ G G + K+
Sbjct: 395 FLGLRRNAKN-----TAWQKGKFGENTIIANIDTGVWPESKSFNDKGYGPVPSKWRGGKA 559
Query: 181 ------VTVYLNGLPRKLIGARFFNKAYEAFHGKLPSSQQTARDFVGPGTHTLSTAGGNF 234
N RKLIGARFF+ AYEA++ KLPS Q+TARDF+G GTHTLSTAGGNF
Sbjct: 560 CEISKFSKYKKNPCNRKLIGARFFSNAYEAYNDKLPSWQRTARDFLGHGTHTLSTAGGNF 739
Query: 235 VQNATIFGIGNGTIKGGSPRSRVATYKACWSLTDVVDCFGADVLAAIDQAIYDGADLISV 294
V +A++F IGNGT+KGGSPR+RVATYK CWSL D+ DCFGADVLAAIDQAI DG D+IS+
Sbjct: 740 VPDASVFAIGNGTVKGGSPRARVATYKVCWSLLDLEDCFGADVLAAIDQAISDGVDIISL 919
Query: 295 SAGGKPNTNPEVIFTDEISIGAFHALARNILLVASAGNEGPTPGSVTNVAPWVFTVAAST 354
S G PE IFTDE+SIGAFHAL+RNILLVASAGNEGPT GSV NVAPWVFT+AAST
Sbjct: 920 SLAGHSLVYPEDIFTDEVSIGAFHALSRNILLVASAGNEGPTGGSVVNVAPWVFTIAAST 1099
Query: 355 LDRDFSSVMTINNKTLTGASLFVNLPPNQDFLIIISTDAKFANVTDVDAQFCRPGTLDPS 414
LDRDFSS +TI N+T+ GASLFVNLPPNQ F +I+STD K AN T+ DAQFC+PGTLDPS
Sbjct: 1100LDRDFSSTITIGNQTIRGASLFVNLPPNQAFPLIVSTDGKLANATNHDAQFCKPGTLDPS 1279
Query: 415 KVNGKVVACDREGKINSIAEGQEALSAGAVGVIMRNQPEVDGKTLLAEPHVVSTINY-YD 473
KV GK+V C REG I S+AEGQEALSAGA G+++ NQP+ GKT LAEPH +S + +
Sbjct: 1280KVKGKIVECIREGNIKSVAEGQEALSAGAKGMLLSNQPK-QGKTTLAEPHTLSCVEVPHH 1456
Query: 474 ARSITTPKGS-------------EITPED--IKTNATIRMSPANALNGRKPAPVMASFSS 518
A PK S +IT D +K TI+ S A L GRKPAPVMASFSS
Sbjct: 1457APKPPKPKKSAEQERAGSHAPAFDITSMDSKLKAGTTIKFSGAKTLYGRKPAPVMASFSS 1636
Query: 519 RGPNKVQPYILKPDVTAPGVNILAAYSLLASVSNLVTDNRRGFPFNIQQGTSMSCPHVVG 578
RGPNK+QP ILKPDVTAPGVNILAAYSL AS SNL TDNR FPFN+ QGTSMSCPHV G
Sbjct: 1637RGPNKIQPSILKPDVTAPGVNILAAYSLYASASNLKTDNRNNFPFNVLQGTSMSCPHVAG 1816
Query: 579 TAGLIKTLHPNWSPAAIKSAIMTT 602
AGLIKTLHPNWSPAAIKSAIMTT
Sbjct: 1817IAGLIKTLHPNWSPAAIKSAIMTT 1888
>BG580583 weakly similar to GP|9757901|dbj serine protease-like protein
{Arabidopsis thaliana}, partial (13%)
Length = 592
Score = 298 bits (762), Expect(2) = 2e-91
Identities = 147/149 (98%), Positives = 148/149 (98%)
Frame = +3
Query: 190 RKLIGARFFNKAYEAFHGKLPSSQQTARDFVGPGTHTLSTAGGNFVQNATIFGIGNGTIK 249
RKLIGARFFNKAYEAFHGKLPSSQQTARDFVG GTHTLSTAGGNFVQNATIFGIGNGTIK
Sbjct: 144 RKLIGARFFNKAYEAFHGKLPSSQQTARDFVGHGTHTLSTAGGNFVQNATIFGIGNGTIK 323
Query: 250 GGSPRSRVATYKACWSLTDVVDCFGADVLAAIDQAIYDGADLISVSAGGKPNTNPEVIFT 309
GGSPRSRVATYKACWSLTDVVDCFGADVLAAIDQ+IYDGADLISVSAGGKPNTNPEVIFT
Sbjct: 324 GGSPRSRVATYKACWSLTDVVDCFGADVLAAIDQSIYDGADLISVSAGGKPNTNPEVIFT 503
Query: 310 DEISIGAFHALARNILLVASAGNEGPTPG 338
DEISIGAFHALARNILLVASAGNEGPTPG
Sbjct: 504 DEISIGAFHALARNILLVASAGNEGPTPG 590
Score = 57.4 bits (137), Expect(2) = 2e-91
Identities = 32/41 (78%), Positives = 32/41 (78%), Gaps = 9/41 (21%)
Frame = +1
Query: 151 VFGRNRRVLTTEE*VQFH*DGVG---------ILLRKFHVT 182
VFGRNRRVLTTEE*VQFH*DGVG ILLRKFHVT
Sbjct: 22 VFGRNRRVLTTEE*VQFH*DGVGVTFVNLTNSILLRKFHVT 144
>TC85586 similar to GP|20521377|dbj|BAB91889. putative subtilisin-like
protease {Oryza sativa (japonica cultivar-group)},
partial (17%)
Length = 1460
Score = 311 bits (797), Expect = 5e-85
Identities = 157/212 (74%), Positives = 174/212 (82%), Gaps = 1/212 (0%)
Frame = +1
Query: 392 DAKFANVTDVDAQFCRPGTLDPSKVNGKVVACDREGKINSIAEGQEALSAGAVGVIMRNQ 451
DAKF+N T DA+FCRP TLDPSKV GK+VAC REGKI S+AEGQEALSAGA G+ + NQ
Sbjct: 4 DAKFSNATTRDARFCRPRTLDPSKVKGKIVACAREGKIKSVAEGQEALSAGAKGMFLENQ 183
Query: 452 PEVDGKTLLAEPHVVSTINYYDARSITTPKGSEITPED-IKTNATIRMSPANALNGRKPA 510
P+V G TLL+EPHV+ST+ +IT P +T D I++ IR S A L GRKPA
Sbjct: 184 PKVSGNTLLSEPHVLSTVGGNGQAAITAPPRLGVTATDTIESGTKIRFSQAITLIGRKPA 363
Query: 511 PVMASFSSRGPNKVQPYILKPDVTAPGVNILAAYSLLASVSNLVTDNRRGFPFNIQQGTS 570
PVMASFSSRGPN+VQPYILKPDVTAPGVNILAAYSL AS SNL+TDNRRGFPFN+ QGTS
Sbjct: 364 PVMASFSSRGPNQVQPYILKPDVTAPGVNILAAYSLFASASNLLTDNRRGFPFNVMQGTS 543
Query: 571 MSCPHVVGTAGLIKTLHPNWSPAAIKSAIMTT 602
MSCPHV GTAGLIKTLHPNWSPAAIKSAIMTT
Sbjct: 544 MSCPHVAGTAGLIKTLHPNWSPAAIKSAIMTT 639
>TC77557 similar to PIR|JC7519|JC7519 subtilisin-like serine proteinase (EC
3.4.21.-) - Arabidopsis thaliana, partial (88%)
Length = 2682
Score = 303 bits (777), Expect = 1e-82
Identities = 181/419 (43%), Positives = 253/419 (60%), Gaps = 6/419 (1%)
Frame = +3
Query: 190 RKLIGARFFNKAYEAFHGKLPSSQQTA--RDFVGPGTHTLSTAGGNFVQNATIFGIGNGT 247
+KLIGARFF+K EA G + + ++ RD G GTHT STA G+ V +A++FG +GT
Sbjct: 765 KKLIGARFFSKGVEAMLGPIDETTESKSPRDDDGHGTHTSSTAAGSVVPDASLFGYASGT 944
Query: 248 IKGGSPRSRVATYKACWSLTDVVDCFGADVLAAIDQAIYDGADLISVSAGGKPNTNPEVI 307
+G + R+RVA YK CW CF +D+LAAID+AI D +++S+S GG +
Sbjct: 945 ARGMATRARVAVYKVCWK----GGCFSSDILAAIDKAISDNVNVLSLSLGGGMSD----Y 1100
Query: 308 FTDEISIGAFHALARNILLVASAGNEGPTPGSVTNVAPWVFTVAASTLDRDF-SSVMTIN 366
F D ++IGAF A+ + IL+ SAGN GP+ S++NVAPW+ TV A TLDRDF +SV N
Sbjct: 1101 FRDSVAIGAFSAMEKGILVSCSAGNAGPSAYSLSNVAPWITTVGAGTLDRDFPASVSLGN 1280
Query: 367 NKTLTGASLF-VNLPPNQDFLIIISTDAKFANVTDVDAQFCRPGTLDPSKVNGKVVACDR 425
+G SL+ N P +I + +A A + C GTL P V GK+V CDR
Sbjct: 1281 GLNYSGVSLYRGNALPESPLPLIYAGNATNA----TNGNLCMTGTLSPELVAGKIVLCDR 1448
Query: 426 EGKINSIAEGQEALSAGAVGVIMRNQPEVDGKTLLAEPHVV--STINYYDARSITTPKGS 483
G + +G +AG +G+++ N +G+ L+A+ H++ + + + +I S
Sbjct: 1449 -GMNARVQKGAVVKAAGGLGMVLSN-TAANGEELVADTHLLPATAVGEREGNAIKKYLFS 1622
Query: 484 EITPEDIKTNATIRMSPANALNGRKPAPVMASFSSRGPNKVQPYILKPDVTAPGVNILAA 543
E P T+++ G +P+PV+A+FSSRGPN + P ILKPD+ APGVNILA
Sbjct: 1623 EAKP-------TVKIVFQGTKVGVEPSPVVAAFSSRGPNSITPQILKPDLIAPGVNILAG 1781
Query: 544 YSLLASVSNLVTDNRRGFPFNIQQGTSMSCPHVVGTAGLIKTLHPNWSPAAIKSAIMTT 602
+S + L D RR FNI GTSMSCPHV G A LIK+ HP+WSPAA++SA+MTT
Sbjct: 1782 WSKAVGPTGLAVDERR-VDFNIISGTSMSCPHVSGLAALIKSAHPDWSPAAVRSALMTT 1955
>TC92402 similar to GP|20198252|gb|AAM15483.1 subtilisin-like serine
protease AIR3 {Arabidopsis thaliana}, partial (16%)
Length = 704
Score = 213 bits (543), Expect(2) = 4e-77
Identities = 104/132 (78%), Positives = 115/132 (86%)
Frame = +1
Query: 190 RKLIGARFFNKAYEAFHGKLPSSQQTARDFVGPGTHTLSTAGGNFVQNATIFGIGNGTIK 249
RKLIGARFF+ AYE ++GKLP+SQ+TARDFVG GTHTLSTAGGNFV A+IF IGNGTIK
Sbjct: 307 RKLIGARFFSDAYERYNGKLPTSQRTARDFVGHGTHTLSTAGGNFVPGASIFNIGNGTIK 486
Query: 250 GGSPRSRVATYKACWSLTDVVDCFGADVLAAIDQAIYDGADLISVSAGGKPNTNPEVIFT 309
GGSPR+RVATYK CWSLTD CFGADVL+AIDQAI DG D+I VSAGG +TN E IFT
Sbjct: 487 GGSPRARVATYKVCWSLTDAASCFGADVLSAIDQAIDDGVDIIFVSAGGPSSTNSEEIFT 666
Query: 310 DEISIGAFHALA 321
DE+SIGAFHALA
Sbjct: 667 DEVSIGAFHALA 702
Score = 93.6 bits (231), Expect(2) = 4e-77
Identities = 53/83 (63%), Positives = 61/83 (72%), Gaps = 9/83 (10%)
Frame = +2
Query: 109 AQLANCTQHVLGNSLD*VQMMSTQLGKREGLVKIPL*LTLIQVFGRNRRVLTTEE*VQFH 168
A+ NCT HVLGN LD ++M+STQLGKREGLVKIP *LTLIQVFGRN+ VL TEE QF
Sbjct: 59 AKSINCTLHVLGNFLDCMEMISTQLGKREGLVKIPS*LTLIQVFGRNQGVLATEELAQFQ 238
Query: 169 *DGVGIL---------LRKFHVT 182
+GVG++ LRKF VT
Sbjct: 239 QNGVGVMSVKLTNSEVLRKFLVT 307
>TC85584 weakly similar to PIR|A84473|A84473 probable serine proteinase
[imported] - Arabidopsis thaliana, partial (19%)
Length = 1241
Score = 275 bits (704), Expect = 3e-74
Identities = 136/175 (77%), Positives = 151/175 (85%)
Frame = +1
Query: 428 KINSIAEGQEALSAGAVGVIMRNQPEVDGKTLLAEPHVVSTINYYDARSITTPKGSEITP 487
+I S+AEGQEALSAGA GVI+RNQPE++GKTLL+EPHV+STI+Y S TT + +I P
Sbjct: 1 EIKSVAEGQEALSAGAKGVILRNQPEINGKTLLSEPHVLSTISYPGNHSRTTGRSLDIIP 180
Query: 488 EDIKTNATIRMSPANALNGRKPAPVMASFSSRGPNKVQPYILKPDVTAPGVNILAAYSLL 547
DIK+ +RMSPA LN RKPAPVMAS+SSRGPNKVQP ILKPDVTAPGVNILAAYSL
Sbjct: 181 SDIKSGTKLRMSPAKTLNRRKPAPVMASYSSRGPNKVQPSILKPDVTAPGVNILAAYSLF 360
Query: 548 ASVSNLVTDNRRGFPFNIQQGTSMSCPHVVGTAGLIKTLHPNWSPAAIKSAIMTT 602
AS SNL+TD RRGFPFN+ QGTSMSCPHV GTAGLIKTLHPNWSPAAIKSAIMTT
Sbjct: 361 ASASNLITDTRRGFPFNVMQGTSMSCPHVAGTAGLIKTLHPNWSPAAIKSAIMTT 525
>TC85585 similar to GP|22773236|gb|AAN06842.1 Putatvie subtilisin-like
serine protease {Oryza sativa (japonica
cultivar-group)}, partial (21%)
Length = 1353
Score = 273 bits (698), Expect = 1e-73
Identities = 136/138 (98%), Positives = 137/138 (98%)
Frame = +2
Query: 465 VVSTINYYDARSITTPKGSEITPEDIKTNATIRMSPANALNGRKPAPVMASFSSRGPNKV 524
VVSTINY+D RSITTPKGSEITPEDIKTNATIRMSPANALNGRKPAPVMASFSSRGPNKV
Sbjct: 5 VVSTINYHDPRSITTPKGSEITPEDIKTNATIRMSPANALNGRKPAPVMASFSSRGPNKV 184
Query: 525 QPYILKPDVTAPGVNILAAYSLLASVSNLVTDNRRGFPFNIQQGTSMSCPHVVGTAGLIK 584
QPYILKPDVTAPGVNILAAYSLLASVSNLVTDNRRGFPFNIQQGTSMSCPHVVGTAGLIK
Sbjct: 185 QPYILKPDVTAPGVNILAAYSLLASVSNLVTDNRRGFPFNIQQGTSMSCPHVVGTAGLIK 364
Query: 585 TLHPNWSPAAIKSAIMTT 602
TLHPNWSPAAIKSAIMTT
Sbjct: 365 TLHPNWSPAAIKSAIMTT 418
>TC85588 similar to GP|20198169|gb|AAM15440.1 subtilisin-like serine
protease AIR3 {Arabidopsis thaliana}, partial (7%)
Length = 572
Score = 270 bits (689), Expect = 2e-72
Identities = 140/217 (64%), Positives = 159/217 (72%)
Frame = +1
Query: 251 GSPRSRVATYKACWSLTDVVDCFGADVLAAIDQAIYDGADLISVSAGGKPNTNPEVIFTD 310
GSPR+RVATYK CWSL D+ DCFGADVLAAIDQAI DG D+IS+S G PE IFTD
Sbjct: 1 GSPRARVATYKVCWSLLDLEDCFGADVLAAIDQAISDGVDIISLSLAGHSLVYPEDIFTD 180
Query: 311 EISIGAFHALARNILLVASAGNEGPTPGSVTNVAPWVFTVAASTLDRDFSSVMTINNKTL 370
E+SIGAFHAL+RNILLVASAGNEGPT GSV NV PWVFT+AASTLDRDFSS +TI N+T+
Sbjct: 181 EVSIGAFHALSRNILLVASAGNEGPTGGSVVNVTPWVFTIAASTLDRDFSSTITIGNQTI 360
Query: 371 TGASLFVNLPPNQDFLIIISTDAKFANVTDVDAQFCRPGTLDPSKVNGKVVACDREGKIN 430
GASLFVNLPPNQ F +I+STD K AN T+ DAQFC+PGTLDPSKV G
Sbjct: 361 RGASLFVNLPPNQAFPLIVSTDGKLANATNHDAQFCKPGTLDPSKVKG------------ 504
Query: 431 SIAEGQEALSAGAVGVIMRNQPEVDGKTLLAEPHVVS 467
+++ NQP+ GKT LAEPH +S
Sbjct: 505 ---------------MLLSNQPK-QGKTTLAEPHTLS 567
>TC86329 similar to PIR|T07171|T07171 subtilisin-like proteinase (EC 3.4.21.-)
1 - tomato, partial (91%)
Length = 2671
Score = 166 bits (419), Expect(2) = 4e-70
Identities = 109/258 (42%), Positives = 147/258 (56%), Gaps = 4/258 (1%)
Frame = +3
Query: 349 TVAASTLDRDFSSVMTINN-KTLTGASLFVN-LPPNQDFLIIISTDAKFANVT-DVDAQF 405
TV A T+DRDF + +T+ N G SL+ LPPN ++ + ANV+ D
Sbjct: 1086 TVGARTIDRDFPAYITLGNGNRYNGVSLYNGKLPPNSPLPLVYA-----ANVSQDSSDNL 1250
Query: 406 CRPGTLDPSKVNGKVVACDREGKINSIAEGQEALS-AGAVGVIMRNQPEVDGKTLLAEPH 464
C +L PSKV+GK+V CDR G N AE + AG +G+I+ N + G+ L+A+
Sbjct: 1251 CSTDSLIPSKVSGKIVICDRGG--NPRAEKSLVVKRAGGIGMILANNQDY-GEELVADSF 1421
Query: 465 VVSTINYYDARSITTPKGSEITPEDIKTNATIRMSPANALNGRKPAPVMASFSSRGPNKV 524
++ + S K + P N T +++ G +P+PV+A+FSSRGPN +
Sbjct: 1422 LLPAAALGEKASNEIKKYASSAP-----NPTAKIAFGGTRFGVQPSPVVAAFSSRGPNIL 1586
Query: 525 QPYILKPDVTAPGVNILAAYSLLASVSNLVTDNRRGFPFNIQQGTSMSCPHVVGTAGLIK 584
P ILKPD+ APGVNILA +S + L D R FNI GTSMSCPHV G A L+K
Sbjct: 1587 TPKILKPDLIAPGVNILAGWSGKVGPTGLSVDTRH-VSFNIISGTSMSCPHVSGLAALLK 1763
Query: 585 TLHPNWSPAAIKSAIMTT 602
HP WSPAAI+SA+MTT
Sbjct: 1764 GAHPEWSPAAIRSALMTT 1817
Score = 118 bits (295), Expect(2) = 4e-70
Identities = 67/164 (40%), Positives = 98/164 (58%), Gaps = 6/164 (3%)
Frame = +2
Query: 190 RKLIGARFFNKAYEAFHGKLPSSQQTA--RDFVGPGTHTLSTAGGNFVQNATIFGIGNGT 247
+KL+GARFF K YEA G + + ++ RD G G+HT +TA G+ V A++FG +GT
Sbjct: 617 KKLVGARFFAKGYEAAFGPIDENTESKSPRDDDGHGSHTSTTAAGSAVAGASLFGFASGT 796
Query: 248 IKGGSPRSRVATYKACWSLTDVVDCFGADVLAAIDQAI----YDGADLISVSAGGKPNTN 303
KG + ++RVA YK CW + CF +D+ AAID+AI A +S GG +
Sbjct: 797 AKGMATQARVAAYKVCW----LGGCFTSDIAAAIDKAIDTQARTAAYKVSRLGGGLTD-- 958
Query: 304 PEVIFTDEISIGAFHALARNILLVASAGNEGPTPGSVTNVAPWV 347
+ D +++G F A+ IL+ +SAGN GP+ S+ NVAPW+
Sbjct: 959 ---YYKDTVAMGTFAAIEHGILVSSSAGNGGPSKASLANVAPWI 1081
>TC87377 weakly similar to PIR|T07171|T07171 subtilisin-like proteinase (EC
3.4.21.-) 1 - tomato, partial (54%)
Length = 2071
Score = 186 bits (473), Expect(2) = 2e-59
Identities = 116/290 (40%), Positives = 171/290 (58%), Gaps = 2/290 (0%)
Frame = +2
Query: 255 SRVATYKACWSLTDVVDCFGADVLAAIDQAIYDGADLISVSAGGKPNTNPEVIFTDEISI 314
+RVA YK CW + CF +D+ A +D+AI DG +++S+S GG + D I+I
Sbjct: 11 ARVAAYKVCW----LSGCFTSDIAAGMDKAIEDGVNILSMSIGGSIMD----YYRDIIAI 166
Query: 315 GAFHALARNILLVASAGNEGPTPGSVTNVAPWVFTVAASTLDRDFSSVMTI-NNKTLTGA 373
GAF A++ IL+ +SAGN GP+ S++NVAPW+ TV A T+DRDF S +T+ N KT TGA
Sbjct: 167 GAFTAMSHGILVSSSAGNGGPSAESLSNVAPWITTVGAGTIDRDFPSYITLGNGKTYTGA 346
Query: 374 SLFVNLPPNQDFLIIISTDAKFANVTDVDAQF-CRPGTLDPSKVNGKVVACDREGKINSI 432
SL+ P + L ++ NV++ + C P +L SKV GK+V C+R G + +
Sbjct: 347 SLYNGKPSSDSLLPVVYA----GNVSESSVGYLCIPDSLTSSKVLGKIVICERGGN-SRV 511
Query: 433 AEGQEALSAGAVGVIMRNQPEVDGKTLLAEPHVVSTINYYDARSITTPKGSEITPEDIKT 492
+G +AG VG+I+ N E G+ L+A+ H++ +S T K T ++ +
Sbjct: 512 EKGLVVKNAGGVGMILVNN-EAYGEELIADSHLLPAA-ALGQKSSTVLKDYVFTTKNPRA 685
Query: 493 NATIRMSPANALNGRKPAPVMASFSSRGPNKVQPYILKPDVTAPGVNILA 542
+ +P+PV+A+FSSRGPN + P ILKPD+ APGVNILA
Sbjct: 686 KLVFGGTHLQV----QPSPVVAAFSSRGPNSLTPKILKPDLIAPGVNILA 823
Score = 61.6 bits (148), Expect(2) = 2e-59
Identities = 27/46 (58%), Positives = 34/46 (73%)
Frame = +1
Query: 557 NRRGFPFNIQQGTSMSCPHVVGTAGLIKTLHPNWSPAAIKSAIMTT 602
++R FNI GTSMSCPH G A ++K +P WSPAAI+SA+MTT
Sbjct: 865 DKRHVNFNIISGTSMSCPHASGLAAIVKGAYPEWSPAAIRSALMTT 1002
>BE249159 weakly similar to GP|20198252|gb subtilisin-like serine protease
AIR3 {Arabidopsis thaliana}, partial (24%)
Length = 661
Score = 213 bits (541), Expect = 2e-55
Identities = 116/209 (55%), Positives = 144/209 (68%), Gaps = 3/209 (1%)
Frame = +1
Query: 180 HVTVYLNGLP--RKLIGARFFNKAYEAFHGKLPSSQQTARDFVGPGTHTLSTAGGNFVQN 237
H T+ L+ RKLIGAR F K YEA+ GKL +S TARD +G G+HTLSTAGGNFVQ
Sbjct: 49 HPTIILSNCSTSRKLIGARAFYKGYEAYVGKLDASFYTARDTIGHGSHTLSTAGGNFVQG 228
Query: 238 ATIFGIGNGTIKGGSPRSRVATYKACWSLTDVVDCFGADVLAAIDQAIYDGADLISVSAG 297
+++G GNGT KGGSP++ VA YK CW C ADVLA + AI DG D++SVS G
Sbjct: 229 VSVYGNGNGTAKGGSPKAHVAAYKVCWK----GGCSDADVLAGFEAAISDGVDVLSVSLG 396
Query: 298 GKPNTNPEVIFTDEISIGAFHALARNILLVASAGNEGPTPGSVTNVAPWVFTVAASTLDR 357
K + +FTD ISIG+FHA+A I++VASAGN GP G+V+NVAPW+FTVAAST+DR
Sbjct: 397 MKTHN----LFTDSISIGSFHAVANGIVVVASAGNSGPYFGTVSNVAPWLFTVAASTIDR 564
Query: 358 DFSSVMTI-NNKTLTGASLFVNLPPNQDF 385
DF+S +T+ +NK G SL P F
Sbjct: 565 DFASYVTLGDNKHFKGTSLSSKDLPTHKF 651
>CB892799 weakly similar to PIR|T05768|T05 subtilisin-like proteinase (EC
3.4.21.-) - Arabidopsis thaliana, partial (20%)
Length = 875
Score = 192 bits (488), Expect = 3e-49
Identities = 122/297 (41%), Positives = 175/297 (58%), Gaps = 4/297 (1%)
Frame = +2
Query: 310 DEISIGAFHALARNILLVASAGNEGPTPGSVTNVAPWVFTVAASTLDRDFSSVMTINNKT 369
D ISIG+FHA R + +V+SAGNEG GS TN+APW+ TVAA + DRDF+S + + N
Sbjct: 2 DAISIGSFHAANRGLFVVSSAGNEGNL-GSATNLAPWMLTVAAGSTDRDFTSDIILGNGA 178
Query: 370 -LTGASLFVNLPPNQDFLIIISTDAKFANVTDVDAQFCRPGTLDPSKVNGKVVACDREGK 428
+TG SL + N II +++A T + +C +L+ +K GKV+ C +
Sbjct: 179 KITGESLSL-FEMNASTRIISASEAFAGYFTPYQSSYCLESSLNKTKTKGKVLVCRHVER 355
Query: 429 I--NSIAEGQEALSAGAVGVIMRNQPEVDGKTLLAEPHVVSTINYYDARSITTPKGSEIT 486
+ +A+ + AG VG+I+ ++ + D +A P V+ + + KG +I
Sbjct: 356 STESKVAKSKIVKEAGGVGMILIDETDQD----VAIPFVIPSA------IVGKKKGQKIL 505
Query: 487 PEDIKTNATI-RMSPANALNGRKPAPVMASFSSRGPNKVQPYILKPDVTAPGVNILAAYS 545
T + ++ A + G + AP +A+FSSRGPN + P ILKPD+TAPG+NILAA+S
Sbjct: 506 SYLKTTRKPMSKILRAKTVIGAQSAPRVAAFSSRGPNALNPEILKPDITAPGLNILAAWS 685
Query: 546 LLASVSNLVTDNRRGFPFNIQQGTSMSCPHVVGTAGLIKTLHPNWSPAAIKSAIMTT 602
+A G FNI GTSM+CPHV G A L+K +HP+WSP+AIKSAIMTT
Sbjct: 686 PVA-----------GNMFNILSGTSMACPHVTGIATLVKAVHPSWSPSAIKSAIMTT 823
>BI307979 weakly similar to GP|20521377|db putative subtilisin-like protease
{Oryza sativa (japonica cultivar-group)}, partial (18%)
Length = 777
Score = 170 bits (431), Expect = 1e-42
Identities = 101/268 (37%), Positives = 155/268 (57%), Gaps = 1/268 (0%)
Frame = +3
Query: 278 LAAIDQAIYDGADLISVSAGGKPNTNPEVIFTDEISIGAFHALARNILLVASAGNEGPTP 337
L AID AI DG D+I++S G E D I+ GA A+ +NI++V SAGN GP+P
Sbjct: 3 LKAIDDAIEDGVDVINLSIGFPAPLKYE---DDVIAKGALQAVRKNIVVVCSAGNAGPSP 173
Query: 338 GSVTNVAPWVFTVAASTLDRDFSSVMTINN-KTLTGASLFVNLPPNQDFLIIISTDAKFA 396
S++N +PW+ TV AST+DR F + + ++N T+ G S+ N +++++D ++A
Sbjct: 174 HSLSNPSPWIITVGASTVDRTFLAPIKLSNGTTIEGRSITPLRMGNSFCPLVLASDVEYA 353
Query: 397 NVTDVDAQFCRPGTLDPSKVNGKVVACDREGKINSIAEGQEALSAGAVGVIMRNQPEVDG 456
+ ++ +C TLDPSKV GK+V C R G+ + + E AG VG+I+ N +V
Sbjct: 354 GILSANSSYCLDNTLDPSKVKGKIVLCMR-GQGGRVKKSLEVQRAGGVGLILGNN-KVYA 527
Query: 457 KTLLAEPHVVSTINYYDARSITTPKGSEITPEDIKTNATIRMSPANALNGRKPAPVMASF 516
+ ++P+ + T ++ + +P N ++ P + KPAP MA F
Sbjct: 528 NDVPSDPYFIPTTGVTYENTLKLVQYIHSSP-----NPMAQLLPGRTVLDTKPAPSMAIF 692
Query: 517 SSRGPNKVQPYILKPDVTAPGVNILAAY 544
SSRGPN + P ILKPD+TAPGV+I AA+
Sbjct: 693 SSRGPNIIDPNILKPDITAPGVDIFAAW 776
>TC83163 weakly similar to GP|6721520|dbj|BAA89562.1 EST AU029428(E30359)
corresponds to a region of the predicted gene.~Similar
to antifreeze-like, partial (29%)
Length = 990
Score = 160 bits (404), Expect = 2e-39
Identities = 116/322 (36%), Positives = 167/322 (51%), Gaps = 18/322 (5%)
Frame = +2
Query: 45 YIVYLGAHVHGPTPSSVDLETATYSHYDLLGSILGSHEEAEEAIIYSYNKQINGFAAILE 104
Y+VY+G++++G VD + H DLL SI+ S E A+I+ Y+ NGF+A+L
Sbjct: 95 YVVYMGSNING-----VDGQIPESVHLDLLSSIIPSEESERIALIHHYSHAFNGFSAMLT 259
Query: 105 EEEAAQLANCTQHVLGNSLD*VQMMSTQ--------LGKR-EGLVKIPL*LTLIQVFGRN 155
+ EA+ LA V +++ +T+ LG R G++K I +
Sbjct: 260 QSEASALAGNDGVVSVFEDPFLELHTTRSWDFLESDLGMRPHGILKHQHSSNDIII---- 427
Query: 156 RRVLTT---EE*VQFH*DGVGILLRKFHVTVY------LNGLPRKLIGARFFNKAYEAFH 206
V+ T E F +G+G + ++ + RKLIGAR++NK
Sbjct: 428 -GVIDTGIWPESPSFKDEGIGKIPSRWKGVCMEAHDFKKSNCNRKLIGARYYNKK----- 589
Query: 207 GKLPSSQQTARDFVGPGTHTLSTAGGNFVQNATIFGIGNGTIKGGSPRSRVATYKACWSL 266
+ + RDF G GTHT STA G V NA+ +G+ GT +GGSP +R+A YKAC
Sbjct: 590 ----DPKGSPRDFNGHGTHTASTAAGVIVNNASYYGLAKGTARGGSPSARIAAYKAC--- 748
Query: 267 TDVVDCFGADVLAAIDQAIYDGADLISVSAGGKPNTNPEVIFTDEISIGAFHALARNILL 326
C G +L AID AI DG D+IS+S G E + +D I+IGAFHA R +++
Sbjct: 749 -SGEGCSGGTLLKAIDDAIKDGVDIISISIGFSSEFLSEYL-SDPIAIGAFHAEQRGVMV 922
Query: 327 VASAGNEGPTPGSVTNVAPWVF 348
V SAGNEGP +V N PW+F
Sbjct: 923 VCSAGNEGPDHYTVVNTTPWIF 988
>TC81705 weakly similar to GP|6721520|dbj|BAA89562.1 EST AU029428(E30359)
corresponds to a region of the predicted gene.~Similar
to antifreeze-like, partial (33%)
Length = 1443
Score = 158 bits (400), Expect = 5e-39
Identities = 88/194 (45%), Positives = 119/194 (60%), Gaps = 4/194 (2%)
Frame = +1
Query: 190 RKLIGARFFNK--AYEAFHGKLPSSQQTARDFVGPGTHTLSTAGGNFVQNATIFGIGNGT 247
RKLIGAR++N + + + ++ + RD VG GTHT STA G V NA +G+ GT
Sbjct: 547 RKLIGARYYNTQDTFGSNKTHIGGAKGSPRDTVGHGTHTASTAAGVNVNNANYYGLAKGT 726
Query: 248 IKGGSPRSRVATYKACWSLTDVVDCFGADVLAAIDQAIYDGADLISVSAGGKPNTNPEVI 307
+GGSP +R+A YK C C G+ +L A+D AI DG D+IS+S G + +
Sbjct: 727 ARGGSPSTRIAAYKTC----SEEGCSGSTILKAMDDAIKDGVDIISISIGLSSLMQSDYL 894
Query: 308 FTDEISIGAFHALARNILLVASAGNEGPTPGSVTNVAPWVFTVAASTLDRDFSSVMTI-N 366
D I+IGAFHA R + V SAGN+GP P +V N APW+FTVAAS +DR+F S + + N
Sbjct: 895 -NDPIAIGAFHAEQRGVTAVCSAGNDGPDPNTVVNTAPWIFTVAASNIDRNFQSTIVLGN 1071
Query: 367 NKTLTGASL-FVNL 379
K+ GA + F NL
Sbjct: 1072GKSFQGAGINFSNL 1113
>BE124129 weakly similar to GP|9757901|dbj serine protease-like protein
{Arabidopsis thaliana}, partial (22%)
Length = 582
Score = 151 bits (381), Expect = 8e-37
Identities = 88/194 (45%), Positives = 121/194 (62%), Gaps = 2/194 (1%)
Frame = +2
Query: 387 IIISTDAKFANVTDVDAQFCRPGTLDPSKVNGKVVACDREGKINSIAEGQEALSAGAVGV 446
II +TDAK A+ T+ DA C+ GTLDP KV GK+V C R G + +G++AL AGAVG+
Sbjct: 29 IIKATDAKLASATNEDAVLCQNGTLDPKKVKGKIVLCLR-GINARVDKGEQALLAGAVGM 205
Query: 447 IMRNQPEVDGKTLLAEPHVV--STINYYDARSITTPKGSEITPEDIKTNATIRMSPANAL 504
++ N + G ++A+PHV+ S IN+ D + S +P T+ T ++
Sbjct: 206 VLAND-KTTGNEIIADPHVLPASHINFSDGVEVFHYVNSSKSPVAYITHPTTKLHT---- 370
Query: 505 NGRKPAPVMASFSSRGPNKVQPYILKPDVTAPGVNILAAYSLLASVSNLVTDNRRGFPFN 564
KPAP MA+FSS+GPN + P ILKPD+TAPGV+++AAY+ +N DNRR FN
Sbjct: 371 ---KPAPFMAAFSSKGPNTIIPEILKPDITAPGVSVIAAYTEAEGPTNQEFDNRR-IQFN 538
Query: 565 IQQGTSMSCPHVVG 578
GTSMSCPH+ G
Sbjct: 539 SVSGTSMSCPHISG 580
>TC89220 similar to PIR|T05768|T05768 subtilisin-like proteinase (EC
3.4.21.-) - Arabidopsis thaliana, partial (39%)
Length = 1149
Score = 147 bits (372), Expect = 9e-36
Identities = 80/184 (43%), Positives = 115/184 (62%), Gaps = 8/184 (4%)
Frame = +3
Query: 190 RKLIGARFFNKAYEAFHGKL----PSSQ----QTARDFVGPGTHTLSTAGGNFVQNATIF 241
RKLIGAR+F+K +E G P ++ ++ RD G GTHT STA G + A +
Sbjct: 564 RKLIGARYFSKGHEVGAGSAGPLNPINETVEFRSPRDADGHGTHTASTAAGRYAFQANMS 743
Query: 242 GIGNGTIKGGSPRSRVATYKACWSLTDVVDCFGADVLAAIDQAIYDGADLISVSAGGKPN 301
G +G KG +P++R+A YK CW + CF +D+LAA D A+ DG D+IS+S GG
Sbjct: 744 GYASGIAKGVAPKARLAVYKVCWKNSG---CFDSDILAAFDAAVNDGVDVISISIGGGDG 914
Query: 302 TNPEVIFTDEISIGAFHALARNILLVASAGNEGPTPGSVTNVAPWVFTVAASTLDRDFSS 361
+ D I+IG++ A++R + + +SAGN+GP+ SVTN+APW+ TV A T+DRDF S
Sbjct: 915 I-ASPYYLDPIAIGSYGAVSRGVFVSSSAGNDGPSGMSVTNLAPWLTTVGAGTIDRDFPS 1091
Query: 362 VMTI 365
+ I
Sbjct: 1092QIII 1103
>AW329591 weakly similar to GP|15010692|gb| At2g04160/T16B23.1 {Arabidopsis
thaliana}, partial (22%)
Length = 592
Score = 141 bits (356), Expect = 6e-34
Identities = 82/200 (41%), Positives = 120/200 (60%), Gaps = 3/200 (1%)
Frame = +2
Query: 348 FTVAASTLDRDFSSVMTINNK-TLTGASLFVNLPPNQDFLIIISTDAKFANVTDVDAQFC 406
F+VAAST+DR+F S + + +K + G SL LP + + ++ S DAK N T DA+ C
Sbjct: 2 FSVAASTIDRNFVSYLQLGDKHIIMGTSLSTGLPNEKFYSLVSSVDAKVGNATIEDAKIC 181
Query: 407 RPGTLDPSKVNGKVVACDREGKINSIAEGQEALSAGAVGVIMRNQPEVDGKTLLAEPHVV 466
+ G+LDP+KV GK++ C + +EA+S G++G+++ N + G ++A H++
Sbjct: 182 KVGSLDPNKVKGKILFCLLRELDGLVYAEEEAISGGSIGLVLGNDKQ-RGNDIMAYAHLL 358
Query: 467 ST--INYYDARSITTPKGSEITPEDIKTNATIRMSPANALNGRKPAPVMASFSSRGPNKV 524
T INY D + + + TP T A + G KPAPV+AS SSRGPN +
Sbjct: 359 PTSHINYTDGEYVHSYIKATKTPMAYMTKAKTEV-------GVKPAPVIASLSSRGPNPI 517
Query: 525 QPYILKPDVTAPGVNILAAY 544
QP ILKPD+TAPGV+IL AY
Sbjct: 518 QPIILKPDITAPGVDILYAY 577
>TC89137 similar to GP|4200338|emb|CAA76726.1 P69C protein {Lycopersicon
esculentum}, partial (37%)
Length = 1230
Score = 136 bits (343), Expect = 2e-32
Identities = 86/208 (41%), Positives = 114/208 (54%), Gaps = 2/208 (0%)
Frame = +3
Query: 397 NVTDVDAQFCRPGTLDPSKVNGKVVACDREGKINSIAEGQEALSAGAVGVIMRNQPEVDG 456
N +D FC P + V GK+V C++ G + +A+GQ AG +I+ N D
Sbjct: 21 NTSDDSIAFCGPIAMKKVDVKGKIVVCEQGGFVGRVAKGQAVKDAGGAAMILLNSEGEDF 200
Query: 457 KTLLAEPHVVSTIN--YYDARSITTPKGSEITPEDIKTNATIRMSPANALNGRKPAPVMA 514
+ A+ HV+ ++ Y +I S TP ATI + G AP +A
Sbjct: 201 NPI-ADVHVLPAVHVSYSAGLNIQDYINSTSTPM-----ATILFK--GTVIGNPNAPQVA 356
Query: 515 SFSSRGPNKVQPYILKPDVTAPGVNILAAYSLLASVSNLVTDNRRGFPFNIQQGTSMSCP 574
SFSSRGP+K P ILKPD+ PG+NILA + + S+ N + FNI GTSMSCP
Sbjct: 357 SFSSRGPSKASPGILKPDILGPGLNILAGWPI--SLDNSTSS------FNIIAGTSMSCP 512
Query: 575 HVVGTAGLIKTLHPNWSPAAIKSAIMTT 602
H+ G A L+K HP+WSPAAIKSAIMTT
Sbjct: 513 HLSGIAALLKNSHPDWSPAAIKSAIMTT 596
>BI262965 similar to PIR|A84473|A84 probable serine proteinase [imported] -
Arabidopsis thaliana, partial (16%)
Length = 394
Score = 129 bits (324), Expect = 3e-30
Identities = 65/130 (50%), Positives = 87/130 (66%)
Frame = +3
Query: 238 ATIFGIGNGTIKGGSPRSRVATYKACWSLTDVVDCFGADVLAAIDQAIYDGADLISVSAG 297
+T+ G GT +G +P++R+A YK CW TD CF +D+LA IDQAI DG D++S+S G
Sbjct: 6 STLLGYATGTARGMAPQARIAVYKVCW--TD--GCFASDILAGIDQAIQDGVDVLSLSLG 173
Query: 298 GKPNTNPEVIFTDEISIGAFHALARNILLVASAGNEGPTPGSVTNVAPWVFTVAASTLDR 357
G +T + D I+IGAF A+ R I + SAGN GP GS++NVAPW+ TV A TLDR
Sbjct: 174 GSSSTP---YYFDTIAIGAFAAVERGIFVSCSAGNTGPRSGSLSNVAPWIMTVGAGTLDR 344
Query: 358 DFSSVMTINN 367
DF + T+ N
Sbjct: 345 DFPAYATLGN 374
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.323 0.138 0.412
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 17,222,572
Number of Sequences: 36976
Number of extensions: 231328
Number of successful extensions: 1331
Number of sequences better than 10.0: 91
Number of HSP's better than 10.0 without gapping: 1242
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1265
length of query: 611
length of database: 9,014,727
effective HSP length: 102
effective length of query: 509
effective length of database: 5,243,175
effective search space: 2668776075
effective search space used: 2668776075
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.5 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (22.0 bits)
S2: 61 (28.1 bits)
Medicago: description of AC133779.1


