

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
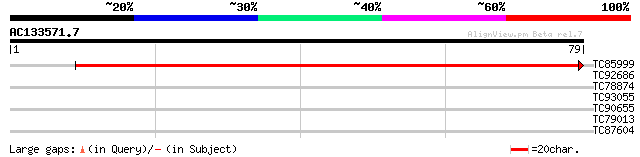
Query= AC133571.7 + phase: 0
(79 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC85999 similar to GP|16209645|gb|AAL14383.1 At2g25110/F13D4.70 ... 145 4e-36
TC92686 weakly similar to PIR|T39560|T39560 dolichyl-phosphate-m... 27 1.9
TC78874 similar to PIR|T49098|T49098 hypothetical protein F4F15.... 26 2.4
TC93055 similar to GP|6625560|gb|AAF19265.1| beta-glucan binding... 26 3.2
TC90655 homologue to GP|17979539|gb|AAL50104.1 At1g65700/F1E22_3... 24 9.3
TC79013 similar to GP|21537154|gb|AAM61495.1 unknown {Arabidopsi... 24 9.3
TC87604 similar to GP|9758199|dbj|BAB08673.1 gb|AAF03497.1~gene_... 24 9.3
>TC85999 similar to GP|16209645|gb|AAL14383.1 At2g25110/F13D4.70
{Arabidopsis thaliana}, partial (88%)
Length = 1244
Score = 145 bits (365), Expect = 4e-36
Identities = 68/70 (97%), Positives = 69/70 (98%)
Frame = +3
Query: 10 EGSGKTWKQDQRFRLQHIDTSGYFYSHDKKYSRIAGGQQEVCGVREKRADNVWLAAEGVY 69
EGSGKTWKQDQRFRLQHIDTSGY +SHDKKYSRIAGGQQEVCGVREKRADNVWLAAEGVY
Sbjct: 723 EGSGKTWKQDQRFRLQHIDTSGYLHSHDKKYSRIAGGQQEVCGVREKRADNVWLAAEGVY 902
Query: 70 LPVTESKQAE 79
LPVTESKQAE
Sbjct: 903 LPVTESKQAE 932
Score = 26.2 bits (56), Expect = 2.4
Identities = 23/70 (32%), Positives = 31/70 (43%), Gaps = 3/70 (4%)
Frame = +3
Query: 1 MNPTPETIGEGSGKTWKQDQRFRLQHIDTSGYFYSHDKKYSRIAGGQQEVC--GVREKRA 58
+ P P T + G T K RLQH+ T + +SH S I+G + C G E
Sbjct: 525 VRPEPGTSAK-QGDTIKSGTIIRLQHMRTRKWLHSH-LHASPISGNLEVSCFGGENESDT 698
Query: 59 DNVW-LAAEG 67
+ W L EG
Sbjct: 699 GDYWRLLIEG 728
>TC92686 weakly similar to PIR|T39560|T39560
dolichyl-phosphate-mannose--protein O-mannosyl
transferase - fission yeast, partial (9%)
Length = 768
Score = 26.6 bits (57), Expect = 1.9
Identities = 13/45 (28%), Positives = 21/45 (45%), Gaps = 6/45 (13%)
Frame = +1
Query: 24 LQHIDTSGYFYSHDKKY------SRIAGGQQEVCGVREKRADNVW 62
L+H +T Y +SH+ Y R++ Q+V G +N W
Sbjct: 43 LRHKETKTYLHSHEDPYPLRYDDGRVSSQGQQVTGYPYNDTNNYW 177
>TC78874 similar to PIR|T49098|T49098 hypothetical protein F4F15.300 -
Arabidopsis thaliana, partial (74%)
Length = 1630
Score = 26.2 bits (56), Expect = 2.4
Identities = 13/43 (30%), Positives = 19/43 (43%)
Frame = +2
Query: 15 TWKQDQRFRLQHIDTSGYFYSHDKKYSRIAGGQQEVCGVREKR 57
+W + RL S SHD KY +A ++C V K+
Sbjct: 881 SWNKIGHKRLLRKSASAMAVSHDGKYLSLASKDGDICVVEVKK 1009
>TC93055 similar to GP|6625560|gb|AAF19265.1| beta-glucan binding protein
{Phaseolus vulgaris}, partial (31%)
Length = 746
Score = 25.8 bits (55), Expect = 3.2
Identities = 9/16 (56%), Positives = 13/16 (81%)
Frame = -2
Query: 47 QQEVCGVREKRADNVW 62
++EVCGVRE R ++W
Sbjct: 139 REEVCGVREDRGMSLW 92
>TC90655 homologue to GP|17979539|gb|AAL50104.1 At1g65700/F1E22_3
{Arabidopsis thaliana}, partial (75%)
Length = 474
Score = 24.3 bits (51), Expect = 9.3
Identities = 12/32 (37%), Positives = 18/32 (55%)
Frame = +3
Query: 35 SHDKKYSRIAGGQQEVCGVREKRADNVWLAAE 66
SH++ +S G QQ V G+ R DN+ + E
Sbjct: 69 SHERVFSTREGVQQLVLGLYIIRGDNISVVGE 164
>TC79013 similar to GP|21537154|gb|AAM61495.1 unknown {Arabidopsis
thaliana}, partial (42%)
Length = 1471
Score = 24.3 bits (51), Expect = 9.3
Identities = 13/41 (31%), Positives = 16/41 (38%)
Frame = +3
Query: 11 GSGKTWKQDQRFRLQHIDTSGYFYSHDKKYSRIAGGQQEVC 51
G+GK W+ D FR Q + H R GG C
Sbjct: 825 GNGKAWRADDAFRPQTLPNLS--KQHSMPNPRFVGGVPWSC 941
>TC87604 similar to GP|9758199|dbj|BAB08673.1
gb|AAF03497.1~gene_id:K17N15.12~similar to unknown
protein {Arabidopsis thaliana}, partial (96%)
Length = 1194
Score = 24.3 bits (51), Expect = 9.3
Identities = 11/24 (45%), Positives = 15/24 (61%), Gaps = 1/24 (4%)
Frame = +2
Query: 13 GKTWKQDQRFRLQH-IDTSGYFYS 35
G+TWK D R R+ H T G +Y+
Sbjct: 521 GRTWKGDGRIRI*HRTHTDG*YYT 592
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.313 0.131 0.398
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 2,563,468
Number of Sequences: 36976
Number of extensions: 27636
Number of successful extensions: 82
Number of sequences better than 10.0: 14
Number of HSP's better than 10.0 without gapping: 80
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 82
length of query: 79
length of database: 9,014,727
effective HSP length: 55
effective length of query: 24
effective length of database: 6,981,047
effective search space: 167545128
effective search space used: 167545128
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 51 (24.3 bits)
Medicago: description of AC133571.7


