

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC133139.3 - phase: 0
(185 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
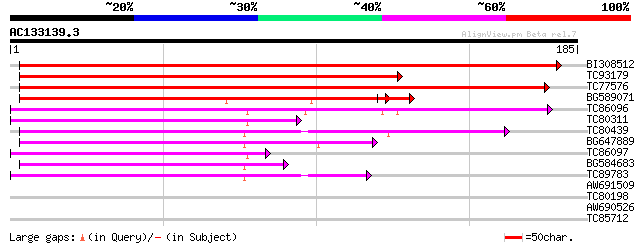
BI308512 similar to SP|O24464|KAD_ Adenylate kinase (EC 2.7.4.3)... 352 4e-98
TC93179 similar to SP|O24464|KAD_PRUAR Adenylate kinase (EC 2.7.... 247 2e-66
TC77576 similar to SP|O04905|UMPK_ARATH Uridylate kinase (EC 2.7... 222 6e-59
BG589071 similar to GP|15928999|gb UMP-CMP kinase {Homo sapiens}... 119 2e-28
TC86096 homologue to SP|Q08480|KADB_ORYSA Adenylate kinase B (EC... 74 3e-14
TC80311 similar to GP|15146328|gb|AAK83647.1 AT5g47840/MCA23_18 ... 73 7e-14
TC80439 similar to GP|6063552|dbj|BAA85412.1 ESTs AU065232(E6085... 69 8e-13
BG647889 similar to PIR|D84790|D84 probable adenylate kinase [im... 66 9e-12
TC86097 homologue to SP|Q08480|KADB_ORYSA Adenylate kinase B (EC... 59 1e-09
BG584683 similar to GP|6063552|dbj ESTs AU065232(E60855) C23624(... 58 2e-09
TC89783 similar to GP|21928121|gb|AAM78088.1 AT3g01820/F28J7_15 ... 54 3e-08
AW691509 similar to GP|6063552|dbj| ESTs AU065232(E60855) C23624... 37 0.006
TC80198 weakly similar to GP|19310480|gb|AAL84974.1 AT5g60340/k9... 31 0.25
AW690526 similar to GP|17473880|gb| putative protein {Arabidopsi... 27 4.6
TC85712 similar to GP|18643341|gb|AAL76272.1 transcription facto... 27 6.0
>BI308512 similar to SP|O24464|KAD_ Adenylate kinase (EC 2.7.4.3) (ATP-AMP
transphosphorylase). [Apricot] {Prunus armeniaca},
partial (77%)
Length = 778
Score = 352 bits (904), Expect = 4e-98
Identities = 175/177 (98%), Positives = 175/177 (98%)
Frame = +2
Query: 4 GGPGSGKGTQCARIVETFGFKHLSAGDLLRKAMVSDSEYGAMILETIREGRIVPSAVTVR 63
GGPGSGKGTQCARIVETFGFKHLSAGDLLRKAMVSDSEYGAMILETIREGRIVPSAVTVR
Sbjct: 179 GGPGSGKGTQCARIVETFGFKHLSAGDLLRKAMVSDSEYGAMILETIREGRIVPSAVTVR 358
Query: 64 LILREMQYGDNRKFLIDGFPRSEENRIAFEHITGTEPDFVLYFDCPEEEMVKRVLSRNQG 123
LILREMQYGDNRKFLIDGFPRSEENRIAFEHITGTEPDFVLYFDCPEEEMVKRVLSRNQG
Sbjct: 359 LILREMQYGDNRKFLIDGFPRSEENRIAFEHITGTEPDFVLYFDCPEEEMVKRVLSRNQG 538
Query: 124 RIDDNIDTIKKRLKVFEALNLPVIDHYARRGRLHRINAVGTEDEIFEQVRPVFAACE 180
RIDDNIDTIKKRLKVFEALNLPVIDHYARRGRLHRINAVGT DEIFE VRPVFAACE
Sbjct: 539 RIDDNIDTIKKRLKVFEALNLPVIDHYARRGRLHRINAVGTRDEIFEPVRPVFAACE 709
>TC93179 similar to SP|O24464|KAD_PRUAR Adenylate kinase (EC 2.7.4.3)
(ATP-AMP transphosphorylase). [Apricot] {Prunus
armeniaca}, partial (55%)
Length = 674
Score = 247 bits (630), Expect = 2e-66
Identities = 121/125 (96%), Positives = 124/125 (98%)
Frame = +3
Query: 4 GGPGSGKGTQCARIVETFGFKHLSAGDLLRKAMVSDSEYGAMILETIREGRIVPSAVTVR 63
GGPGSGKGTQCARIVETFGFKHLSAGDLLRK MVSDSEYGAMILETIREGRIVPSAVTVR
Sbjct: 273 GGPGSGKGTQCARIVETFGFKHLSAGDLLRKEMVSDSEYGAMILETIREGRIVPSAVTVR 452
Query: 64 LILREMQYGDNRKFLIDGFPRSEENRIAFEHITGTEPDFVLYFDCPEEEMVKRVLSRNQG 123
LILREMQYGDNRKFLIDGFPRSEENRIAFEHITGTEPDFVL+FDCPE+EMVKRVLSRNQG
Sbjct: 453 LILREMQYGDNRKFLIDGFPRSEENRIAFEHITGTEPDFVLFFDCPEKEMVKRVLSRNQG 632
Query: 124 RIDDN 128
RIDD+
Sbjct: 633 RIDDH 647
>TC77576 similar to SP|O04905|UMPK_ARATH Uridylate kinase (EC 2.7.4.-) (UK)
(Uridine monophosphate kinase) (UMP kinase) (UMP/CMP
kinase)., partial (93%)
Length = 1073
Score = 222 bits (566), Expect = 6e-59
Identities = 107/173 (61%), Positives = 135/173 (77%)
Frame = +2
Query: 4 GGPGSGKGTQCARIVETFGFKHLSAGDLLRKAMVSDSEYGAMILETIREGRIVPSAVTVR 63
GGPGSGKGTQCA +VE FGF HLSAGDLLR + S SE G MI I+EG+IVPS VT+R
Sbjct: 296 GGPGSGKGTQCANVVEHFGFTHLSAGDLLRAEIKSGSENGTMIQNMIKEGKIVPSEVTIR 475
Query: 64 LILREMQYGDNRKFLIDGFPRSEENRIAFEHITGTEPDFVLYFDCPEEEMVKRVLSRNQG 123
L+ + ++ N KFLIDGFPR+EENR AFE +TG EP FVLYFDCPEEEM +R+LSRNQG
Sbjct: 476 LLQQAIKDNGNDKFLIDGFPRNEENRAAFERVTGIEPAFVLYFDCPEEEMERRLLSRNQG 655
Query: 124 RIDDNIDTIKKRLKVFEALNLPVIDHYARRGRLHRINAVGTEDEIFEQVRPVF 176
R DDNI+TI+KR KVF +LPVI++Y +G++ +++A +E+FE V+ +F
Sbjct: 656 REDDNIETIRKRFKVFLDSSLPVINYYDAKGKVRKVDAARPVEEVFESVKAIF 814
>BG589071 similar to GP|15928999|gb UMP-CMP kinase {Homo sapiens}, partial
(50%)
Length = 579
Score = 119 bits (299), Expect(2) = 2e-28
Identities = 65/126 (51%), Positives = 81/126 (63%), Gaps = 5/126 (3%)
Frame = +2
Query: 4 GGPGSGKGTQCARIVETFGFKHLSAGDLLRKAMVSDSEYGAMILETIREGRIVPSAVTVR 63
GGPG+GKGTQCA IV FGF HLSAGDLLR+ S SE +I I+EG+IVP +TV
Sbjct: 173 GGPGAGKGTQCANIVRDFGFAHLSAGDLLREERNSGSETAQLIDRYIKEGQIVPVEITVN 352
Query: 64 LILREM---QYGDNRKFLIDGFPRSEENRIAFEHITG--TEPDFVLYFDCPEEEMVKRVL 118
L+ R M Q + FLIDGFPR+++N + + G FVL+FDCPEE M R+L
Sbjct: 353 LLKRAMEQHQKSGSNYFLIDGFPRNQDNLDGWNRVMGNFATVQFVLFFDCPEEVMEARLL 532
Query: 119 SRNQGR 124
R + R
Sbjct: 533 ERGKTR 550
Score = 34.7 bits (78), Expect = 0.022
Identities = 12/34 (35%), Positives = 24/34 (70%)
Frame = +2
Query: 143 NLPVIDHYARRGRLHRINAVGTEDEIFEQVRPVF 176
++PVIDH+A G++ +I A + ++++ V+ VF
Sbjct: 11 SMPVIDHFATLGKVRKITATNSVEQVYSDVQQVF 112
Score = 22.3 bits (46), Expect(2) = 2e-28
Identities = 8/12 (66%), Positives = 11/12 (91%)
Frame = +1
Query: 121 NQGRIDDNIDTI 132
N+GR DDNI++I
Sbjct: 544 NKGRTDDNIESI 579
>TC86096 homologue to SP|Q08480|KADB_ORYSA Adenylate kinase B (EC 2.7.4.3)
(ATP-AMP transphosphorylase). [Rice] {Oryza sativa},
partial (97%)
Length = 1117
Score = 74.3 bits (181), Expect = 3e-14
Identities = 57/210 (27%), Positives = 95/210 (45%), Gaps = 33/210 (15%)
Frame = +2
Query: 1 MLSGGPGSGKGTQCARIVETFGFKHLSAGDLLRKAMVSDSEYGAMILETIREGRIVPSAV 60
+L G PGSGKGTQ I + + HL+ GD+LR A+ + + G E + +G +V +
Sbjct: 164 ILVGPPGSGKGTQSPIIKDEYCLCHLATGDMLRAAVAAKTPLGIKAKEAMEKGELVSDDL 343
Query: 61 TVRLILREMQYGDNRK-FLIDGFPRSEENRIAFEHI---TGTEPDFVLYFDCPEEEMVKR 116
V +I M+ +K F++DGFPR+ + + G + D VL F + + +R
Sbjct: 344 VVGIIDEAMKKPSCQKGFILDGFPRTVVQAQKLDEMLQKQGAKIDKVLNFAIDDAILEER 523
Query: 117 VLSR----NQGRI-------------------------DDNIDTIKKRLKVFEALNLPVI 147
+ R + GR DD +K RL+ F PVI
Sbjct: 524 ITGRWIHPSSGRTYHTKFAPPKVSGVDDVTGEPLIQRKDDTAAVLKSRLESFHRQTEPVI 703
Query: 148 DHYARRGRLHRINAVGTEDEIFEQVRPVFA 177
D+Y+++G + ++A E+ +V V +
Sbjct: 704 DYYSKKGIVASLHAEKPPKEVTSEVEKVLS 793
>TC80311 similar to GP|15146328|gb|AAK83647.1 AT5g47840/MCA23_18
{Arabidopsis thaliana}, partial (43%)
Length = 656
Score = 72.8 bits (177), Expect = 7e-14
Identities = 37/97 (38%), Positives = 57/97 (58%), Gaps = 2/97 (2%)
Frame = +3
Query: 1 MLSGGPGSGKGTQCARIVETFGFKHLSAGDLLRKAMVSDSEYGAMILETIREGRIVPSAV 60
M+SG P SGKGTQC I +G H++AGDLLR + + SE G E + +G++VP +
Sbjct: 285 MISGAPASGKGTQCHLITNKYGLVHVAAGDLLRAEIETGSENGKRAKEYMEKGQLVPDEI 464
Query: 61 TVRLILREMQYGDNRK--FLIDGFPRSEENRIAFEHI 95
V ++ + D+ K +L+DG+PRS A + +
Sbjct: 465 VVMMVKDRLLKPDSVKNGWLLDGYPRSLSQATALKEL 575
>TC80439 similar to GP|6063552|dbj|BAA85412.1 ESTs AU065232(E60855)
C23624(S1554) AU078241(E60855) correspond to a region
of the predicted gene.~, partial (71%)
Length = 1172
Score = 69.3 bits (168), Expect = 8e-13
Identities = 48/203 (23%), Positives = 84/203 (40%), Gaps = 43/203 (21%)
Frame = +2
Query: 4 GGPGSGKGTQCARIVETFGFKHLSAGDLLRKAMVSDSEYGAMILETIREGRIVPSAVTVR 63
G PG GKGT +R+ G H++ GDL+R + S+ + + E + +G++V + +
Sbjct: 140 GCPGVGKGTYASRLCNLLGVPHIATGDLVRHELSSNGPLSSQLSEIVNQGQLVSDEIIMS 319
Query: 64 LILREMQYGDNR---KFLIDGFPRSEENRIAFEHITGTEPDFVLYFDCPEEEMVKRVLSR 120
L+ + + G + F++DGFPR+ + E + T+ D V+ EE ++ + L R
Sbjct: 320 LLSKRLAEGQAKGESGFILDGFPRTIKQAEILEGV--TDIDLVVNLKIREEALLAKCLGR 493
Query: 121 NQ----------------------------------------GRIDDNIDTIKKRLKVFE 140
R DD +K+RL+++
Sbjct: 494 RTCSQCGGNFNVASINIKGENGCPGISMDPLLPPEHCMSKLITRSDDTEPVVKERLRIYN 673
Query: 141 ALNLPVIDHYARRGRLHRINAVG 163
L+ PV Y RG+L G
Sbjct: 674 ELSQPVEGFYRSRGKLMEFELPG 742
>BG647889 similar to PIR|D84790|D84 probable adenylate kinase [imported] -
Arabidopsis thaliana, partial (65%)
Length = 793
Score = 65.9 bits (159), Expect = 9e-12
Identities = 34/127 (26%), Positives = 67/127 (51%), Gaps = 10/127 (7%)
Frame = +3
Query: 4 GGPGSGKGTQCARIVETFGFKHLSAGDLLRKAMVSDSEYGAMILETIREGRIVPSAVTVR 63
G PG GKGT +R+ G H++ GDL+R+ + S + + E +++G++V + +
Sbjct: 183 GCPGVGKGTYASRLSNLLGVPHIATGDLVREELSSSGPLSSQLSEIVKQGQLVSDEIIIN 362
Query: 64 LILREMQYGDNR---KFLIDGFPRSEENRIAFEHITGTE-------PDFVLYFDCPEEEM 113
L+ + + G+ + F++DGFPR+ + ++ I +E D V+ PE +
Sbjct: 363 LLSKRLDDGEAKGELGFILDGFPRTIKQAVSISSIW*SEILEGVADIDLVINLRLPENVL 542
Query: 114 VKRVLSR 120
+++ L R
Sbjct: 543 LEKCLGR 563
>TC86097 homologue to SP|Q08480|KADB_ORYSA Adenylate kinase B (EC 2.7.4.3)
(ATP-AMP transphosphorylase). [Rice] {Oryza sativa},
partial (84%)
Length = 699
Score = 58.5 bits (140), Expect = 1e-09
Identities = 32/86 (37%), Positives = 50/86 (57%), Gaps = 1/86 (1%)
Frame = +3
Query: 1 MLSGGPGSGKGTQCARIVETFGFKHLSAGDLLRKAMVSDSEYGAMILETIREGRIVPSAV 60
+L G PGSGKGTQ I + + HL+ GD+LR A+ + + G E + +G +V +
Sbjct: 138 ILVGPPGSGKGTQSPIIKDEYCLCHLATGDMLRAAVAAKTPLGIKAKEAMEKGELVSDDL 317
Query: 61 TVRLILREMQYGDNRK-FLIDGFPRS 85
V +I M+ +K F++DGFPR+
Sbjct: 318 VVGIIDEAMKKPSCQKGFILDGFPRT 395
>BG584683 similar to GP|6063552|dbj ESTs AU065232(E60855) C23624(S1554)
AU078241(E60855) correspond to a region of the predicted
gene.~, partial (44%)
Length = 719
Score = 58.2 bits (139), Expect = 2e-09
Identities = 26/91 (28%), Positives = 51/91 (55%), Gaps = 3/91 (3%)
Frame = +1
Query: 4 GGPGSGKGTQCARIVETFGFKHLSAGDLLRKAMVSDSEYGAMILETIREGRIVPSAVTVR 63
G PG GKGT +R+ G H++ GDL+R + S+ + + E + +G++V + +
Sbjct: 148 GCPGVGKGTYASRLCNLLGVPHIATGDLVRHELSSNGPLSSQLSEIVNQGQLVSDEIIMS 327
Query: 64 LILREMQYGDNR---KFLIDGFPRSEENRIA 91
L+ + + G + F++DGFPR+ + ++
Sbjct: 328 LLSKRLAEGQAKGESGFILDGFPRTIKQAVS 420
>TC89783 similar to GP|21928121|gb|AAM78088.1 AT3g01820/F28J7_15
{Arabidopsis thaliana}, partial (55%)
Length = 909
Score = 54.3 bits (129), Expect = 3e-08
Identities = 32/121 (26%), Positives = 57/121 (46%), Gaps = 3/121 (2%)
Frame = +2
Query: 1 MLSGGPGSGKGTQCARIVETFGFKHLSAGDLLRKAMVSDSEYGAMILETIREGRIVPSAV 60
++ G PG+ + R+ + H+S LL + + S I + G+IVP +
Sbjct: 224 VMIGEPGAKRHVFAERLSKLLEVPHISMASLLHQELNPRSSLYQQIAGALDHGKIVPEEI 403
Query: 61 TVRLILREMQYGDNR---KFLIDGFPRSEENRIAFEHITGTEPDFVLYFDCPEEEMVKRV 117
L+ + ++ G +R F++DG PR+ H+ D V+ F CP+EE++K
Sbjct: 404 IFALLSKRLEDGYDRGETGFILDGIPRTRMQAEILNHV--AHVDLVVNFKCPQEELMKNK 577
Query: 118 L 118
L
Sbjct: 578 L 580
>AW691509 similar to GP|6063552|dbj| ESTs AU065232(E60855) C23624(S1554)
AU078241(E60855) correspond to a region of the predicted
gene.~, partial (16%)
Length = 620
Score = 36.6 bits (83), Expect = 0.006
Identities = 15/43 (34%), Positives = 24/43 (54%)
Frame = +1
Query: 4 GGPGSGKGTQCARIVETFGFKHLSAGDLLRKAMVSDSEYGAMI 46
G PG GKGT +R+ G H++ GDL+R + S+ + +
Sbjct: 199 GCPGVGKGTYASRLCNLLGVPHIATGDLVRHELSSNGPLSSQV 327
>TC80198 weakly similar to GP|19310480|gb|AAL84974.1 AT5g60340/k9b18_30
{Arabidopsis thaliana}, partial (89%)
Length = 762
Score = 31.2 bits (69), Expect = 0.25
Identities = 12/39 (30%), Positives = 23/39 (58%)
Frame = +2
Query: 1 MLSGGPGSGKGTQCARIVETFGFKHLSAGDLLRKAMVSD 39
+++G PG+GK T + E H++ GDL+++ + D
Sbjct: 254 LVTGTPGTGKTTTSTALAEATQLNHINIGDLVKEKNLHD 370
>AW690526 similar to GP|17473880|gb| putative protein {Arabidopsis thaliana},
partial (5%)
Length = 557
Score = 26.9 bits (58), Expect = 4.6
Identities = 22/76 (28%), Positives = 33/76 (42%), Gaps = 10/76 (13%)
Frame = +1
Query: 21 FGFKHLSAGDLLRKAMVS----DSEYGAMILETIREGRIVPSAVTVRLILREMQ----YG 72
F F H L+ ++ + YG+++ + R RI V R+ILR Q G
Sbjct: 58 FNFSHAQYFQSLQGVIILS*ALEKLYGSLLYQGYRYSRIPVELVVSRIILRRTQNRETKG 237
Query: 73 DN--RKFLIDGFPRSE 86
+N R F + PR E
Sbjct: 238 ENLGRDFQLSVLPREE 285
>TC85712 similar to GP|18643341|gb|AAL76272.1 transcription factor EIL1
{Vigna radiata}, partial (90%)
Length = 2456
Score = 26.6 bits (57), Expect = 6.0
Identities = 22/86 (25%), Positives = 37/86 (42%), Gaps = 3/86 (3%)
Frame = +2
Query: 98 TEPDFVLYFDCPEEEMVKRVLSRNQGRIDDNIDTIKKRLKVFEALNLPVI---DHYARRG 154
TEPD V+ D +EEM L R R + +K++ K E ++ ARR
Sbjct: 359 TEPDSVVEDDYSDEEMDVDELERRMWRDKMRLKRLKEQTKAKEGIDAAKARQSQEQARRK 538
Query: 155 RLHRINAVGTEDEIFEQVRPVFAACE 180
++ R +D I + + + C+
Sbjct: 539 KMSR-----AQDGILKYMLKMMEVCK 601
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.323 0.142 0.410
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 4,385,479
Number of Sequences: 36976
Number of extensions: 45554
Number of successful extensions: 193
Number of sequences better than 10.0: 30
Number of HSP's better than 10.0 without gapping: 183
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 184
length of query: 185
length of database: 9,014,727
effective HSP length: 90
effective length of query: 95
effective length of database: 5,686,887
effective search space: 540254265
effective search space used: 540254265
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.5 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 55 (25.8 bits)
Medicago: description of AC133139.3


