

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC130807.10 - phase: 0 /pseudo
(530 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
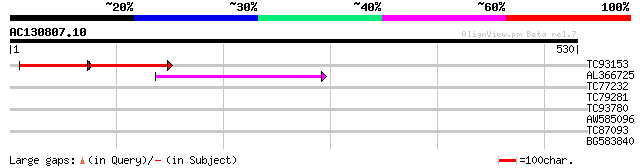
TC93153 similar to GP|14715220|emb|CAC44106. gag polyprotein {Ci... 95 5e-32
AL366725 88 7e-18
TC77232 similar to SP|O04130|SERA_ARATH D-3-phosphoglycerate deh... 30 1.8
TC79281 similar to GP|12597851|gb|AAG60161.1 hypothetical protei... 30 1.8
TC93780 similar to GP|16648903|gb|AAL24303.1 alpha glucosidase-l... 30 3.0
AW585096 similar to GP|11125147|emb MHC class I chain-related pr... 29 5.1
TC87093 homologue to GP|10177817|dbj|BAB11183. gene_id:MKD15.14~... 29 5.1
BG583840 GP|20146556|gb ATP synthase alpha subunit {Clethra barb... 28 6.7
>TC93153 similar to GP|14715220|emb|CAC44106. gag polyprotein {Cicer
arietinum}, partial (8%)
Length = 516
Score = 95.1 bits (235), Expect(2) = 5e-32
Identities = 50/78 (64%), Positives = 56/78 (71%)
Frame = +3
Query: 75 GKFCYLCWKRMELQ*LGRCSGENF*TGTSRKMSEVRKRLNFWS*SRVTCLLLSMLQSLWN 134
G C L W RM L * G CSG +F G ++MS VRKRLNFWS*+RVTCL SMLQSLWN
Sbjct: 279 GLVCCLFWSRMMLW*PGPCSGRSFWAGIFQRMSGVRKRLNFWS*NRVTCLSQSMLQSLWN 458
Query: 135 LQSSTLTIQRRQLNSPNV 152
L STL RRQL+SP+V
Sbjct: 459 LPHSTLITVRRQLSSPSV 512
Score = 61.2 bits (147), Expect(2) = 5e-32
Identities = 37/68 (54%), Positives = 44/68 (64%)
Frame = +1
Query: 10 W*WRSEDAGDLFEESSPNIQGKV*SRWCPEVA*GSGEDFQGHAVL*SAKGAVRDAYVG*G 69
W* ++DA D EESS NIQG V S WC EVA G ++ HA+ * +GAV DA+V *
Sbjct: 85 W**WNQDARDFLEESSTNIQG*VCS*WCLEVAEGD*KNIPSHAMF*DTEGAVWDAHVS*R 264
Query: 70 GR*LVGKF 77
GR*LV F
Sbjct: 265 GR*LVD*F 288
>AL366725
Length = 485
Score = 88.2 bits (217), Expect = 7e-18
Identities = 72/160 (45%), Positives = 90/160 (56%)
Frame = +3
Query: 137 SSTLTIQRRQLNSPNV*SLRIA*ELTSREP*GINRLEFSLIW*VVAGSMRRTQKIIIR** 196
SS T+ R L+S N S R+ * TSR N EFSLI *+ A S RR +++ R *
Sbjct: 3 SSIRTMLLRLLSSRNASSSRMV*GPTSRGQSDTNSSEFSLI**IPAESTRRILRLMTRL* 182
Query: 197 VSEVVRDSRVVLNRTVPLLTRESKD*MMRGGPIGEMLLLR*FVTSVAIKATRVMFVPKMR 256
+S R +RV L+ VPLL R +++ M G MLL R V+++A KATRVMFV K
Sbjct: 183 MSGRPRANRVALSLIVPLLIRANREWSMIGVLRRRMLLRRLSVSTMARKATRVMFVLKRS 362
Query: 257 RSASGVVRRVTH*LIVRGETLFAITTMKRAISVLSAHSRR 296
R+ SGV RRV *LI L A KR I V +A S R
Sbjct: 363 RNVSGVTRRVIL*LIASEMILCASIATKRVILVPNASSLR 482
>TC77232 similar to SP|O04130|SERA_ARATH D-3-phosphoglycerate dehydrogenase
chloroplast precursor (EC 1.1.1.95) (3-PGDH). [Mouse-ear
cress], partial (87%)
Length = 2474
Score = 30.4 bits (67), Expect = 1.8
Identities = 20/49 (40%), Positives = 35/49 (70%)
Frame = +3
Query: 320 EVLVSLIVLL*LLL*ILVLLIVSLLLNVHISWVWFYLI*KEKWLLKLQL 368
++L++L+VLL*+LL LLI+ LLLN+ + ++ +L+ K +L L+L
Sbjct: 603 KLLLNLVVLL*MLL----LLILLLLLNMGLLFLLLWLVTFLKLMLPLKL 737
>TC79281 similar to GP|12597851|gb|AAG60161.1 hypothetical protein
{Arabidopsis thaliana}, partial (57%)
Length = 1860
Score = 30.4 bits (67), Expect = 1.8
Identities = 12/30 (40%), Positives = 21/30 (70%)
Frame = +2
Query: 1 LFNSNRNWSW*WRSEDAGDLFEESSPNIQG 30
L +S +NW++*+R ++ G E +P+IQG
Sbjct: 995 LTSSTKNWAF*YRKKEVGSSGEGGTPSIQG 1084
>TC93780 similar to GP|16648903|gb|AAL24303.1 alpha glucosidase-like protein
{Arabidopsis thaliana}, partial (18%)
Length = 825
Score = 29.6 bits (65), Expect = 3.0
Identities = 14/27 (51%), Positives = 17/27 (62%)
Frame = +3
Query: 3 NSNRNWSW*WRSEDAGDLFEESSPNIQ 29
NS + WSW WR G L ++SPNIQ
Sbjct: 225 NSTQRWSWFWRVLKVGML--KNSPNIQ 299
>AW585096 similar to GP|11125147|emb MHC class I chain-related protein A
{Homo sapiens}, partial (5%)
Length = 231
Score = 28.9 bits (63), Expect = 5.1
Identities = 14/34 (41%), Positives = 25/34 (73%)
Frame = +3
Query: 317 DLSEVLVSLIVLL*LLL*ILVLLIVSLLLNVHIS 350
++ +L++L++LL LLL L LLI+ L+L + I+
Sbjct: 111 EIMRLLLNLLLLLLLLLLFLTLLIILLILTIDIT 212
>TC87093 homologue to GP|10177817|dbj|BAB11183. gene_id:MKD15.14~unknown
protein {Arabidopsis thaliana}, partial (49%)
Length = 1313
Score = 28.9 bits (63), Expect = 5.1
Identities = 24/70 (34%), Positives = 41/70 (58%)
Frame = -1
Query: 276 TLFAITTMKRAISVLSAHSRRRLEPVEKCLH*PVRRLLMRTDLSEVLVSLIVLL*LLL*I 335
T+F I+T + IS+ + ++++ K + *P R L +++ L+V+L LLL
Sbjct: 818 TIFIISTTR--ISIPTRQ*SQQIQMTRKIIP*PRRGSFTHRALLLLVMLLVVVLLLLL-- 651
Query: 336 LVLLIVSLLL 345
LVLL++ LLL
Sbjct: 650 LVLLLLLLLL 621
>BG583840 GP|20146556|gb ATP synthase alpha subunit {Clethra barbinervis},
partial (17%)
Length = 585
Score = 28.5 bits (62), Expect = 6.7
Identities = 16/36 (44%), Positives = 28/36 (77%), Gaps = 1/36 (2%)
Frame = +2
Query: 321 VLVSLIVLL*LLL*ILVLLIVSLLLNVHI-SWVWFY 355
+L+ L++LL LLL +L+LL++ LLL ++I ++W Y
Sbjct: 2 LLLLLLLLLLLLLLLLLLLLLLLLLLLYIYIYMWLY 109
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.370 0.167 0.608
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 14,559,029
Number of Sequences: 36976
Number of extensions: 203931
Number of successful extensions: 2952
Number of sequences better than 10.0: 16
Number of HSP's better than 10.0 without gapping: 1179
Number of HSP's successfully gapped in prelim test: 89
Number of HSP's that attempted gapping in prelim test: 1621
Number of HSP's gapped (non-prelim): 1384
length of query: 530
length of database: 9,014,727
effective HSP length: 101
effective length of query: 429
effective length of database: 5,280,151
effective search space: 2265184779
effective search space used: 2265184779
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 14 ( 7.5 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 36 (21.8 bits)
S2: 61 (28.1 bits)
Medicago: description of AC130807.10


