

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC130806.3 - phase: 0
(320 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
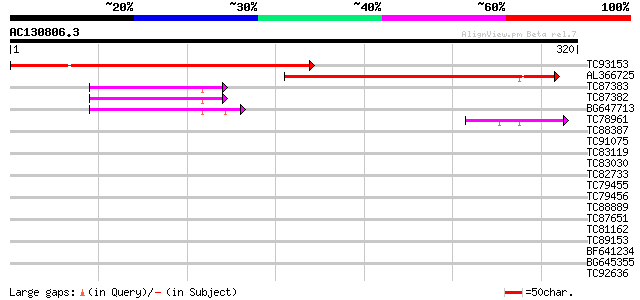
Sequences producing significant alignments: (bits) Value
TC93153 similar to GP|14715220|emb|CAC44106. gag polyprotein {Ci... 269 9e-73
AL366725 250 6e-67
TC87383 similar to GP|19168656|emb|CAD26175. DNA-DIRECTED RNA PO... 52 4e-07
TC87382 similar to EGAD|146423|156195 vitellogenin {Anolis pulch... 50 1e-06
BG647713 homologue to GP|15042313|gb| 232R {Chilo iridescent vir... 49 2e-06
TC78961 similar to GP|18252179|gb|AAL61922.1 unknown protein {Ar... 42 2e-04
TC88387 similar to GP|13357253|gb|AAK20050.1 putative zinc finge... 40 0.001
TC91075 similar to PIR|T48025|T48025 hypothetical protein T12C14... 32 0.25
TC83119 similar to GP|13357253|gb|AAK20050.1 putative zinc finge... 32 0.25
TC83030 weakly similar to GP|18855061|gb|AAL79753.1 putative RNA... 32 0.33
TC82733 similar to GP|10177404|dbj|BAB10535. gene_id:K24M7.12~pi... 32 0.33
TC79455 similar to PIR|T51518|T51518 ubiquitin-fusion degradatio... 31 0.56
TC79456 similar to PIR|T51518|T51518 ubiquitin-fusion degradatio... 31 0.56
TC88889 similar to GP|20340241|gb|AAM19707.1 putative RING zinc ... 30 1.2
TC87651 similar to GP|15292729|gb|AAK92733.1 putative polygalact... 30 1.2
TC81162 weakly similar to PIR|T47673|T47673 hypothetical protein... 30 1.6
TC89153 similar to GP|18855061|gb|AAL79753.1 putative RNA helica... 30 1.6
BF641234 similar to GP|20146108|dbj protein phosphatase 2C {Mese... 29 2.8
BG645355 similar to PIR|G96590|G965 hypothetical protein T24C10.... 29 2.8
TC92636 homologue to GP|15042313|gb|AAK82093.1 232R {Chilo iride... 29 2.8
>TC93153 similar to GP|14715220|emb|CAC44106. gag polyprotein {Cicer
arietinum}, partial (8%)
Length = 516
Score = 269 bits (688), Expect = 9e-73
Identities = 132/172 (76%), Positives = 145/172 (83%)
Frame = +2
Query: 1 MAGSNDAALVAALQAVAQAVGQWPNANAGVTAEMRMLETFMRTHPPTFKGRYDPDGAQTW 60
M GS+DAALVAAL+AVAQAV Q P + G RMLETF+R HPPTFKGRY PDGA W
Sbjct: 2 MTGSSDAALVAALEAVAQAVQQLPKVDTGSDGT-RMLETFLRNHPPTFKGRYAPDGA*KW 178
Query: 61 LKEIERVFRVMQYTEDQKVRFGTHQLAEEGDDWWVSLLPTLKQDGAAVTWAVFRREFLSR 120
LKEIER+FRVMQ E QKV+FGTH LAEE DDWW+SLLP L+QD A VTWA+FR+EFL R
Sbjct: 179 LKEIERIFRVMQCFETQKVQFGTHMLAEEADDWWISLLPVLEQDDAVVTWAMFRKEFLGR 358
Query: 121 YFPEDVRGKKEIEFLELKQGNMSMTEYAAKFVELAKFYPHYAAETTEFLKCI 172
YFPEDVRGKKEIEFLELKQG+MS+TEYAAKFVELA FYPHY+AET EF KCI
Sbjct: 359 YFPEDVRGKKEIEFLELKQGDMSVTEYAAKFVELATFYPHYSAETAEFSKCI 514
>AL366725
Length = 485
Score = 250 bits (638), Expect = 6e-67
Identities = 120/160 (75%), Positives = 132/160 (82%), Gaps = 5/160 (3%)
Frame = +2
Query: 156 KFYPHYAAETTEFLKCIKFENGLRADIKRAIGYQQIRVFSELVNSCRIYEDDTKAHYKIV 215
KFYPHYAAET EF KCIKFENGLR DIKRAIGYQQ+RVF +LVN+CRIYE+DTKAH K+V
Sbjct: 2 KFYPHYAAETAEFSKCIKFENGLRPDIKRAIGYQQLRVFPDLVNTCRIYEEDTKAHDKVV 181
Query: 216 NERKTKGQQSRPKPYSAPADKGKQRMVEDRRPKKKDALAEIICFNCGEKGHKSNACPSEI 275
NERKTKGQ SRPKPYSAPADKGKQRMV+DRRPKKKDA AEI+CFN GEKGHKSN CP EI
Sbjct: 182 NERKTKGQ*SRPKPYSAPADKGKQRMVDDRRPKKKDAPAEIVCFNYGEKGHKSNVCPKEI 361
Query: 276 KRCFRCGKKGH-----KSNAEIICFNIQHEGRASHRGRSP 310
K+C RC KKGH K N +I+CFN EG + + P
Sbjct: 362 KKCVRCDKKGHIVADCKRN-DIVCFNCNEEGHIGSQCKQP 478
>TC87383 similar to GP|19168656|emb|CAD26175. DNA-DIRECTED RNA POLYMERASE II
{Encephalitozoon cuniculi}, partial (0%)
Length = 1247
Score = 51.6 bits (122), Expect = 4e-07
Identities = 24/80 (30%), Positives = 42/80 (52%), Gaps = 2/80 (2%)
Frame = -2
Query: 46 PTFKGRYDPDGAQTWLKEIERVFRVMQYTEDQKVRFGTHQLAEEGDDWWVSLLPTLKQDG 105
P F+G PD WL+ +ER+F+ + E+QKV+ +L + WW ++ K++G
Sbjct: 724 PDFEGNLQPDDLLDWLQIMERLFKYKEVLEEQKVKIVAAKLKKLASIWWENVKRRRKREG 545
Query: 106 AA--VTWAVFRREFLSRYFP 123
+ TW R++ +Y P
Sbjct: 544 KSKIKTWEKMRQKLTRKYLP 485
>TC87382 similar to EGAD|146423|156195 vitellogenin {Anolis pulchellus},
partial (7%)
Length = 2304
Score = 50.1 bits (118), Expect = 1e-06
Identities = 26/80 (32%), Positives = 40/80 (49%), Gaps = 2/80 (2%)
Frame = +2
Query: 46 PTFKGRYDPDGAQTWLKEIERVFRVMQYTEDQKVRFGTHQLAEEGDDWWVSLLPTLKQDG 105
P F+G D WL+ IERVF + E+QKV+ +L + WW +L K++G
Sbjct: 629 PDFEGNLQLDDFLDWLQTIERVFEYKEVPEEQKVKIVAAKLKKHALIWWENLKRRRKREG 808
Query: 106 AA--VTWAVFRREFLSRYFP 123
+ TW R++ +Y P
Sbjct: 809 KSKIKTWDKMRQKLTRKYLP 868
>BG647713 homologue to GP|15042313|gb| 232R {Chilo iridescent virus}, partial
(1%)
Length = 726
Score = 49.3 bits (116), Expect = 2e-06
Identities = 28/95 (29%), Positives = 46/95 (47%), Gaps = 7/95 (7%)
Frame = +2
Query: 46 PTFKGRYDPDGAQTWLKEIERVFRVMQYTEDQKVRFGTHQLAEEGDDWWVSLLPTLKQDG 105
P F G+ PD WL+ IERVF+ + E+QKV+ +L + WW +L +G
Sbjct: 347 PDF*GKLQPDEFVDWLQTIERVFKYKEVAEEQKVKIVAAKLKKHASIWWKNLKRKRNCEG 526
Query: 106 AA--VTWAVFRREFLSR-----YFPEDVRGKKEIE 133
+ TW R++ + Y+ ++ KK I+
Sbjct: 527 KSKIKTWDKMRQKLTRKYLHPHYYQDNFTQKKNIQ 631
>TC78961 similar to GP|18252179|gb|AAL61922.1 unknown protein {Arabidopsis
thaliana}, partial (71%)
Length = 974
Score = 42.4 bits (98), Expect = 2e-04
Identities = 22/65 (33%), Positives = 35/65 (53%), Gaps = 7/65 (10%)
Frame = +2
Query: 258 CFNCGEKGHKSNACPSEI--KRCFRCGKKGH-----KSNAEIICFNIQHEGRASHRGRSP 310
CFNCG GH + C + +C+RCG++GH K++ + + + + R+ R RSP
Sbjct: 359 CFNCGIDGHWARDCKAGDWKNKCYRCGERGHIEKNCKNSPKKLSRHARSVSRSPGRSRSP 538
Query: 311 ANRSS 315
A S
Sbjct: 539 ARSRS 553
Score = 30.4 bits (67), Expect = 0.95
Identities = 17/58 (29%), Positives = 24/58 (41%)
Frame = +2
Query: 258 CFNCGEKGHKSNACPSEIKRCFRCGKKGHKSNAEIICFNIQHEGRASHRGRSPANRSS 315
C+ CGE+GH C + K+ R + +S R+ R RSPA S
Sbjct: 425 CYRCGERGHIEKNCKNSPKKLSRHARSVSRSPG---------RSRSPARSRSPARSRS 571
>TC88387 similar to GP|13357253|gb|AAK20050.1 putative zinc finger protein
{Oryza sativa (japonica cultivar-group)}, partial (96%)
Length = 1286
Score = 40.4 bits (93), Expect = 0.001
Identities = 15/32 (46%), Positives = 23/32 (71%)
Frame = +1
Query: 257 ICFNCGEKGHKSNACPSEIKRCFRCGKKGHKS 288
+C+NC E GH +++CP+E C CGK GH++
Sbjct: 535 LCWNCKEPGHMASSCPNE-GICHTCGKAGHRA 627
Score = 35.8 bits (81), Expect = 0.023
Identities = 26/97 (26%), Positives = 43/97 (43%), Gaps = 19/97 (19%)
Frame = +1
Query: 224 QSRPKPYS-APADKGKQRMVEDRR-----------PKKKDALA----EIICFNCGEKGHK 267
+SR K S +P D+ + R DRR P ++D+ + +C NC GH
Sbjct: 274 RSRSKSRSRSPMDRSRSRSPVDRRIRSERFSHREAPYRRDSRRGFSQDNLCKNCKRPGHY 453
Query: 268 SNACPSEIKRCFRCGKKGH---KSNAEIICFNIQHEG 301
CP+ + C C GH + + + +C+N + G
Sbjct: 454 VRECPN-VAVCHNCSLPGHIASECSTKSLCWNCKEPG 561
Score = 35.4 bits (80), Expect = 0.030
Identities = 18/56 (32%), Positives = 27/56 (48%), Gaps = 9/56 (16%)
Frame = +1
Query: 255 EIICFNCGEKGHKSNAC------PSEIKRCFRCGKKGH---KSNAEIICFNIQHEG 301
E IC CG+ GH++ C P +++ C C K+GH + E C N + G
Sbjct: 586 EGICHTCGKAGHRARECTVPQKPPGDLRLCNNCYKQGHIAVECTNEKACNNCRKTG 753
Score = 29.6 bits (65), Expect = 1.6
Identities = 11/35 (31%), Positives = 20/35 (56%), Gaps = 1/35 (2%)
Frame = +1
Query: 255 EIICFNCGEKGHKSNAC-PSEIKRCFRCGKKGHKS 288
+++C +C + GH S C + C CG +GH++
Sbjct: 904 DVVCRSCQQFGHMSRDCMGGPLMICQNCGGRGHQA 1008
Score = 29.6 bits (65), Expect = 1.6
Identities = 10/18 (55%), Positives = 14/18 (77%)
Frame = +1
Query: 256 IICFNCGEKGHKSNACPS 273
+IC NCG +GH++ CPS
Sbjct: 970 MICQNCGGRGHQAYECPS 1023
>TC91075 similar to PIR|T48025|T48025 hypothetical protein T12C14.30 -
Arabidopsis thaliana, partial (29%)
Length = 686
Score = 32.3 bits (72), Expect = 0.25
Identities = 15/40 (37%), Positives = 22/40 (54%)
Frame = +2
Query: 236 KGKQRMVEDRRPKKKDALAEIICFNCGEKGHKSNACPSEI 275
+ K + E K+K+ E+IC CGE GH + CPS +
Sbjct: 329 EAKSKATERNWKKRKEE--EMICKLCGESGHFTQGCPSTL 442
>TC83119 similar to GP|13357253|gb|AAK20050.1 putative zinc finger protein
{Oryza sativa (japonica cultivar-group)}, partial (16%)
Length = 421
Score = 32.3 bits (72), Expect = 0.25
Identities = 15/64 (23%), Positives = 28/64 (43%)
Frame = +3
Query: 219 KTKGQQSRPKPYSAPADKGKQRMVEDRRPKKKDALAEIICFNCGEKGHKSNACPSEIKRC 278
+++ + P+ +D+ R RR + + +C NC GH + CP+ + C
Sbjct: 231 RSRSRSRSPRIRKIRSDRHSYRDAPYRRDSSRGFSRDNLCKNCKRPGHYARECPN-VAVC 407
Query: 279 FRCG 282
CG
Sbjct: 408 HNCG 419
>TC83030 weakly similar to GP|18855061|gb|AAL79753.1 putative RNA helicase
{Oryza sativa}, partial (7%)
Length = 624
Score = 32.0 bits (71), Expect = 0.33
Identities = 19/61 (31%), Positives = 27/61 (44%), Gaps = 4/61 (6%)
Frame = +1
Query: 218 RKTKGQQSRPKPYSAPADK----GKQRMVEDRRPKKKDALAEIICFNCGEKGHKSNACPS 273
R K S KP + D G+Q P + A CF CGE GH+++ CP+
Sbjct: 43 RSYKSGNSWSKPERSSRDDWLIGGRQSSRSSSSPNRSFAGT---CFTCGESGHRASDCPN 213
Query: 274 E 274
+
Sbjct: 214 K 216
>TC82733 similar to GP|10177404|dbj|BAB10535.
gene_id:K24M7.12~pir||S42136~similar to unknown protein
{Arabidopsis thaliana}, partial (57%)
Length = 710
Score = 32.0 bits (71), Expect = 0.33
Identities = 13/37 (35%), Positives = 20/37 (53%), Gaps = 7/37 (18%)
Frame = +3
Query: 259 FNCGEKGHKSNACPSEIK-------RCFRCGKKGHKS 288
+NCG+ GH CP ++ +CF C ++GH S
Sbjct: 435 YNCGDNGHSLANCPHPLQEGGTMFAQCFVCKEQGHLS 545
Score = 31.6 bits (70), Expect = 0.43
Identities = 22/93 (23%), Positives = 36/93 (38%), Gaps = 23/93 (24%)
Frame = +3
Query: 217 ERKTKGQQSRPKPYSAPADKGKQRMVEDRRP-------KKKDALAEI-----------IC 258
++K K ++ +P S P + V +P K D +A+ IC
Sbjct: 180 KKKNKFKRKKPDSNSKPRTGKRPLRVPGMKPGDSCFICKGLDHIAKFCTQKAEWEKNKIC 359
Query: 259 FNCGEKGHKSNACP-----SEIKRCFRCGKKGH 286
C +GH++ CP + K + CG GH
Sbjct: 360 LRCRRRGHRAQNCPDGGSKEDFKY*YNCGDNGH 458
Score = 28.5 bits (62), Expect = 3.6
Identities = 20/76 (26%), Positives = 32/76 (41%), Gaps = 5/76 (6%)
Frame = +3
Query: 218 RKTKGQQSRPKPYSAPADKGKQRMVEDRRPKKKDALAEIICFNCGEKGHKSNACPSEI-- 275
+K K + R KP S + +R + K D+ CF C H + C +
Sbjct: 177 KKKKNKFKRKKPDSNSKPRTGKRPLRVPGMKPGDS-----CFICKGLDHIAKFCTQKAEW 341
Query: 276 ---KRCFRCGKKGHKS 288
K C RC ++GH++
Sbjct: 342 EKNKICLRCRRRGHRA 389
>TC79455 similar to PIR|T51518|T51518 ubiquitin-fusion degradation
protein-like - Arabidopsis thaliana, partial (8%)
Length = 627
Score = 31.2 bits (69), Expect = 0.56
Identities = 13/21 (61%), Positives = 17/21 (80%)
Frame = -2
Query: 112 VFRREFLSRYFPEDVRGKKEI 132
+F R+FLSR FPE VR KK++
Sbjct: 68 IFSRKFLSRNFPEKVRKKKDL 6
>TC79456 similar to PIR|T51518|T51518 ubiquitin-fusion degradation
protein-like - Arabidopsis thaliana, partial (31%)
Length = 1392
Score = 31.2 bits (69), Expect = 0.56
Identities = 13/21 (61%), Positives = 17/21 (80%)
Frame = -2
Query: 112 VFRREFLSRYFPEDVRGKKEI 132
+F R+FLSR FPE VR KK++
Sbjct: 71 IFSRKFLSRNFPEKVRKKKDL 9
>TC88889 similar to GP|20340241|gb|AAM19707.1 putative RING zinc finger
protein-like protein {Thellungiella halophila}, partial
(46%)
Length = 911
Score = 30.0 bits (66), Expect = 1.2
Identities = 21/73 (28%), Positives = 31/73 (41%), Gaps = 9/73 (12%)
Frame = +1
Query: 224 QSRPKPYS---APADKGKQRMVEDRRPKKKDALAEIICFNCGEKGHKSNAC------PSE 274
Q+ P+ Y +P+D VE+ R + + +C +C E H + C SE
Sbjct: 370 QNIPESYEIHLSPSDD----FVEEFRSRTPTLRFDSVCNSCKEPEHDCSVCLTQFEPESE 537
Query: 275 IKRCFRCGKKGHK 287
I C CG HK
Sbjct: 538 INYCISCGHVFHK 576
>TC87651 similar to GP|15292729|gb|AAK92733.1 putative polygalacturonase PG1
{Arabidopsis thaliana}, partial (35%)
Length = 822
Score = 30.0 bits (66), Expect = 1.2
Identities = 14/41 (34%), Positives = 19/41 (46%)
Frame = +3
Query: 54 PDGAQTWLKEIERVFRVMQYTEDQKVRFGTHQLAEEGDDWW 94
PDG +W + R ++ Y DQ GT + GD WW
Sbjct: 570 PDGPDSWPEADSRNQWLVFYKLDQMSLNGTGTIEGNGDQWW 692
>TC81162 weakly similar to PIR|T47673|T47673 hypothetical protein T26I12.220
- Arabidopsis thaliana, partial (9%)
Length = 756
Score = 29.6 bits (65), Expect = 1.6
Identities = 17/69 (24%), Positives = 26/69 (37%)
Frame = +2
Query: 205 EDDTKAHYKIVNERKTKGQQSRPKPYSAPADKGKQRMVEDRRPKKKDALAEIICFNCGEK 264
E A + E+ G++S P GK VE +C+ C +K
Sbjct: 269 EKSVIAEKSVAGEKSGAGEKSFA--VEQPGAGGKPSSVEQPSSVASGKRKNRMCYGCRQK 442
Query: 265 GHKSNACPS 273
GH + CP+
Sbjct: 443 GHNLSECPN 469
>TC89153 similar to GP|18855061|gb|AAL79753.1 putative RNA helicase {Oryza
sativa}, partial (3%)
Length = 737
Score = 29.6 bits (65), Expect = 1.6
Identities = 8/16 (50%), Positives = 14/16 (87%)
Frame = +3
Query: 258 CFNCGEKGHKSNACPS 273
CF+CG+ GH+++ CP+
Sbjct: 147 CFSCGQPGHRASDCPN 194
>BF641234 similar to GP|20146108|dbj protein phosphatase 2C {Mesembryanthemum
crystallinum}, partial (13%)
Length = 591
Score = 28.9 bits (63), Expect = 2.8
Identities = 19/93 (20%), Positives = 37/93 (39%), Gaps = 1/93 (1%)
Frame = -3
Query: 99 PTLKQDGAAVTWAVFRREFLSRYFPEDVRGKKEIEFLELKQGNMSMTEYAAKFVELAKFY 158
P + W R E S +R ++ F + N + ++ +
Sbjct: 439 PYISTSSGTELWKEIRLETFSNILSRYIR---QLHFRKRSVANKRFSSLKQFTPSISSIF 269
Query: 159 PHYAAETTEF-LKCIKFENGLRADIKRAIGYQQ 190
++ +T++ + KFENG D K+ +GYQ+
Sbjct: 268 LNHKIQTSKIQTQ*PKFENGFNPDCKKELGYQK 170
>BG645355 similar to PIR|G96590|G965 hypothetical protein T24C10.5 [imported]
- Arabidopsis thaliana, partial (5%)
Length = 627
Score = 28.9 bits (63), Expect = 2.8
Identities = 12/45 (26%), Positives = 20/45 (43%), Gaps = 13/45 (28%)
Frame = -1
Query: 258 CFNCGEKGHKSNACPS-------------EIKRCFRCGKKGHKSN 289
C+ C + GH ++ CPS C++C + GH +N
Sbjct: 501 CYKCQQPGHWASNCPSMSAANRVSGGSGGASGNCYKCNQPGHWAN 367
>TC92636 homologue to GP|15042313|gb|AAK82093.1 232R {Chilo iridescent
virus}, partial (1%)
Length = 772
Score = 28.9 bits (63), Expect = 2.8
Identities = 12/23 (52%), Positives = 14/23 (60%)
Frame = +3
Query: 46 PTFKGRYDPDGAQTWLKEIERVF 68
P F+G PD WL+ IERVF
Sbjct: 438 PDFEGELQPDEFVDWLQAIERVF 506
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.319 0.133 0.403
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 9,634,205
Number of Sequences: 36976
Number of extensions: 129331
Number of successful extensions: 652
Number of sequences better than 10.0: 48
Number of HSP's better than 10.0 without gapping: 608
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 642
length of query: 320
length of database: 9,014,727
effective HSP length: 96
effective length of query: 224
effective length of database: 5,465,031
effective search space: 1224166944
effective search space used: 1224166944
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 58 (26.9 bits)
Medicago: description of AC130806.3


