

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC130804.6 + phase: 0 /pseudo
(99 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
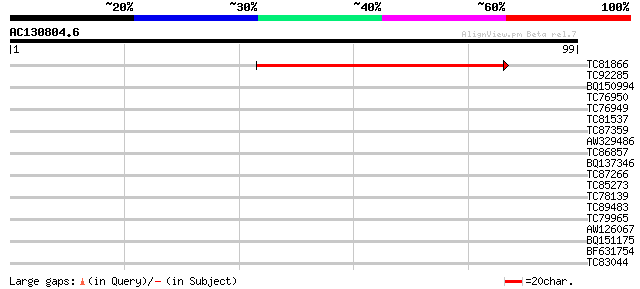
TC81866 similar to GP|11875628|gb|AAG40732.1 XRN3 {Arabidopsis t... 59 2e-10
TC92285 27 0.98
BQ150994 similar to GP|22122173|dbj preproneuropeptide B {Homo s... 27 1.3
TC76950 weakly similar to GP|17380812|gb|AAL36093.1 unknown prot... 26 2.2
TC76949 similar to GP|17380812|gb|AAL36093.1 unknown protein {Ar... 26 2.2
TC81537 similar to GP|7630037|emb|CAB88331.1 putative protein {A... 26 2.9
TC87359 similar to GP|9955544|emb|CAC05429.1 leukotriene-A4 hydr... 26 2.9
AW329486 25 3.7
TC86857 similar to GP|14161088|emb|CAB76907. alpha-fucosidase {C... 25 3.7
BQ137346 homologue to GP|15027095|em hypothetical protein LT.09 ... 25 4.9
TC87266 similar to PIR|T03739|T03739 nitrilase (EC 3.5.5.1) 4B -... 25 4.9
TC85273 weakly similar to GP|8919075|emb|CAA66109.2 specific tis... 25 6.4
TC78139 similar to PIR|S22502|S22502 cysteine proteinase (EC 3.4... 25 6.4
TC89483 similar to GP|18377662|gb|AAL66981.1 unknown protein {Ar... 25 6.4
TC79965 similar to GP|7549635|gb|AAF63820.1| hypothetical protei... 25 6.4
AW126067 similar to GP|7228242|emb cyclic nucleotide gated chann... 25 6.4
BQ151175 24 8.3
BF631754 24 8.3
TC83044 similar to PIR|T48901|T48901 sulfate transporter ATST1 [... 24 8.3
>TC81866 similar to GP|11875628|gb|AAG40732.1 XRN3 {Arabidopsis thaliana},
partial (15%)
Length = 593
Score = 59.3 bits (142), Expect = 2e-10
Identities = 29/44 (65%), Positives = 32/44 (71%)
Frame = +2
Query: 44 IDGVTPRAKMNKQRTRRFRTAKDNEMREAEEERLRKEFEMEANK 87
IDGV PRAKMN+QR+RRFR AKD EE RLR+EFE E K
Sbjct: 428 IDGVAPRAKMNQQRSRRFRAAKDASDAAIEESRLREEFEREGRK 559
>TC92285
Length = 209
Score = 27.3 bits (59), Expect = 0.98
Identities = 17/57 (29%), Positives = 23/57 (39%), Gaps = 11/57 (19%)
Frame = -2
Query: 9 FACH--LLANTLKQKIHETLHAKFNR---------WLIAPPPTLPDIDGVTPRAKMN 54
F CH +LAN + LH R W++ P P LP + +T R N
Sbjct: 178 FVCHPSVLANLSISNLAVLLHPHLRRRIRRLSITTWVVIPSPCLPQVKPLTWRTSPN 8
>BQ150994 similar to GP|22122173|dbj preproneuropeptide B {Homo sapiens},
partial (15%)
Length = 1036
Score = 26.9 bits (58), Expect = 1.3
Identities = 17/51 (33%), Positives = 23/51 (44%)
Frame = +3
Query: 37 PPPTLPDIDGVTPRAKMNKQRTRRFRTAKDNEMREAEEERLRKEFEMEANK 87
PPP + TP ++ TRR R E AE ER R+ + EA +
Sbjct: 243 PPPANTNTKRTTPHGHRQRKHTRRRR-----ETSRAEGEREREGTKKEARR 380
>TC76950 weakly similar to GP|17380812|gb|AAL36093.1 unknown protein
{Arabidopsis thaliana}, partial (87%)
Length = 1338
Score = 26.2 bits (56), Expect = 2.2
Identities = 13/35 (37%), Positives = 19/35 (54%)
Frame = +3
Query: 60 RFRTAKDNEMREAEEERLRKEFEMEANKFFLNKNV 94
R R A D++ + EEE L K E +K+F +V
Sbjct: 624 RDRLAADDKYEDEEEEALEKVIEFFQSKYFKKHSV 728
>TC76949 similar to GP|17380812|gb|AAL36093.1 unknown protein {Arabidopsis
thaliana}, partial (68%)
Length = 997
Score = 26.2 bits (56), Expect = 2.2
Identities = 13/35 (37%), Positives = 19/35 (54%)
Frame = +2
Query: 60 RFRTAKDNEMREAEEERLRKEFEMEANKFFLNKNV 94
R R A D++ + EEE L K E +K+F +V
Sbjct: 512 RDRLAADDKYEDEEEEALEKVIEFFQSKYFKKHSV 616
>TC81537 similar to GP|7630037|emb|CAB88331.1 putative protein {Arabidopsis
thaliana}, partial (34%)
Length = 751
Score = 25.8 bits (55), Expect = 2.9
Identities = 12/25 (48%), Positives = 16/25 (64%)
Frame = +1
Query: 60 RFRTAKDNEMREAEEERLRKEFEME 84
R+R K EM EEE L K+FE++
Sbjct: 415 RYRKKKIEEMGVTEEEYLAKQFEIK 489
>TC87359 similar to GP|9955544|emb|CAC05429.1 leukotriene-A4 hydrolase-like
protein {Arabidopsis thaliana}, partial (84%)
Length = 2022
Score = 25.8 bits (55), Expect = 2.9
Identities = 12/35 (34%), Positives = 18/35 (51%)
Frame = -1
Query: 64 AKDNEMREAEEERLRKEFEMEANKFFLNKNVKCQN 98
A+D +E E FE+E FL K +KC++
Sbjct: 1395 ARDVCFKEVNECFCINRFELEGGNIFLQKLIKCRS 1291
>AW329486
Length = 358
Score = 25.4 bits (54), Expect = 3.7
Identities = 13/23 (56%), Positives = 15/23 (64%)
Frame = +1
Query: 62 RTAKDNEMREAEEERLRKEFEME 84
R + E REA EER+RK FE E
Sbjct: 37 RLQEMKEEREAFEERVRKLFEQE 105
>TC86857 similar to GP|14161088|emb|CAB76907. alpha-fucosidase {Cicer
arietinum}, partial (41%)
Length = 860
Score = 25.4 bits (54), Expect = 3.7
Identities = 16/49 (32%), Positives = 23/49 (46%), Gaps = 6/49 (12%)
Frame = -1
Query: 37 PPPTLPDIDGV---TPRAKMN---KQRTRRFRTAKDNEMREAEEERLRK 79
PPP P IDG+ P KM +T AK NE + +R+++
Sbjct: 248 PPPAAPKIDGII*NVPPGKMGFPFVSKTCSETCAKANEKQMVNAKRIKE 102
>BQ137346 homologue to GP|15027095|em hypothetical protein LT.09 {Leishmania
major}, partial (0%)
Length = 1101
Score = 25.0 bits (53), Expect = 4.9
Identities = 14/33 (42%), Positives = 17/33 (51%)
Frame = +1
Query: 48 TPRAKMNKQRTRRFRTAKDNEMREAEEERLRKE 80
TPR + NKQRT T K + + E R KE
Sbjct: 337 TPRRRRNKQRT----TGKQPKTKAGENARKTKE 423
>TC87266 similar to PIR|T03739|T03739 nitrilase (EC 3.5.5.1) 4B - common
tobacco, partial (97%)
Length = 1382
Score = 25.0 bits (53), Expect = 4.9
Identities = 15/48 (31%), Positives = 20/48 (41%)
Frame = -1
Query: 7 NKFACHLLANTLKQKIHETLHAKFNRWLIAPPPTLPDIDGVTPRAKMN 54
N+ C +L T K + +RW+I PP L TP K N
Sbjct: 845 NRIRCKVLF*TFKNIVR-------SRWIILPPTELVCRQNKTPTFKCN 723
>TC85273 weakly similar to GP|8919075|emb|CAA66109.2 specific tissue protein
2 {Cicer arietinum}, partial (45%)
Length = 839
Score = 24.6 bits (52), Expect = 6.4
Identities = 14/43 (32%), Positives = 22/43 (50%), Gaps = 6/43 (13%)
Frame = +2
Query: 50 RAKMNKQRTRRFRTA------KDNEMREAEEERLRKEFEMEAN 86
RAK K T+ F +DN++ E++R+ K+FE N
Sbjct: 350 RAKRKKGVTKNFEPRPNVSKYEDNDINAEEKKRVVKDFEPRPN 478
>TC78139 similar to PIR|S22502|S22502 cysteine proteinase (EC 3.4.22.-) -
kidney bean, complete
Length = 1720
Score = 24.6 bits (52), Expect = 6.4
Identities = 9/33 (27%), Positives = 18/33 (54%)
Frame = +3
Query: 56 QRTRRFRTAKDNEMREAEEERLRKEFEMEANKF 88
++ +RF K N M ++ K ++++ NKF
Sbjct: 594 EKQKRFNVFKSNVMHVHNTNKMDKPYKLKLNKF 692
>TC89483 similar to GP|18377662|gb|AAL66981.1 unknown protein {Arabidopsis
thaliana}, partial (35%)
Length = 1711
Score = 24.6 bits (52), Expect = 6.4
Identities = 13/39 (33%), Positives = 20/39 (50%), Gaps = 4/39 (10%)
Frame = +1
Query: 50 RAKMNKQRTRRFRTAKDNEM----REAEEERLRKEFEME 84
R + R R + K+ + REAE +R+RKE E +
Sbjct: 130 RKSRERDRERELKREKERVLERYEREAERDRIRKEREQK 246
>TC79965 similar to GP|7549635|gb|AAF63820.1| hypothetical protein
{Arabidopsis thaliana}, partial (15%)
Length = 742
Score = 24.6 bits (52), Expect = 6.4
Identities = 13/34 (38%), Positives = 17/34 (49%), Gaps = 1/34 (2%)
Frame = +1
Query: 35 IAPPPTLPDIDGVTPRA-KMNKQRTRRFRTAKDN 67
+APPP P + R K+RTRR + DN
Sbjct: 178 VAPPPVKPSTRRIPRRTLTRRKRRTRRKLSGDDN 279
>AW126067 similar to GP|7228242|emb cyclic nucleotide gated channel (CNGC4)
like protein {Arabidopsis thaliana}, partial (21%)
Length = 451
Score = 24.6 bits (52), Expect = 6.4
Identities = 12/23 (52%), Positives = 14/23 (60%)
Frame = +2
Query: 55 KQRTRRFRTAKDNEMREAEEERL 77
KQR RR+ K E R EEE+L
Sbjct: 125 KQRIRRYEQYKWQENRGVEEEKL 193
>BQ151175
Length = 211
Score = 24.3 bits (51), Expect = 8.3
Identities = 12/28 (42%), Positives = 15/28 (52%)
Frame = -2
Query: 11 CHLLANTLKQKIHETLHAKFNRWLIAPP 38
C+ TLK KI K NR++IA P
Sbjct: 168 CYTKVLTLKMKIQRWCRFKPNRYMIAIP 85
>BF631754
Length = 552
Score = 24.3 bits (51), Expect = 8.3
Identities = 9/30 (30%), Positives = 15/30 (50%)
Frame = +2
Query: 13 LLANTLKQKIHETLHAKFNRWLIAPPPTLP 42
++ NT++ IH F+ W+ PP P
Sbjct: 344 VVPNTMEDPIHHEDSGFFSSWMSLNPPKFP 433
>TC83044 similar to PIR|T48901|T48901 sulfate transporter ATST1 [imported] -
Arabidopsis thaliana, partial (50%)
Length = 1278
Score = 24.3 bits (51), Expect = 8.3
Identities = 10/20 (50%), Positives = 12/20 (60%)
Frame = -1
Query: 20 QKIHETLHAKFNRWLIAPPP 39
+KI +T H F WLI PP
Sbjct: 423 KKILKTKHQFFRSWLILKPP 364
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.321 0.134 0.398
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 2,996,738
Number of Sequences: 36976
Number of extensions: 30903
Number of successful extensions: 207
Number of sequences better than 10.0: 40
Number of HSP's better than 10.0 without gapping: 207
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 207
length of query: 99
length of database: 9,014,727
effective HSP length: 75
effective length of query: 24
effective length of database: 6,241,527
effective search space: 149796648
effective search space used: 149796648
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 50 (23.9 bits)
Medicago: description of AC130804.6


