

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
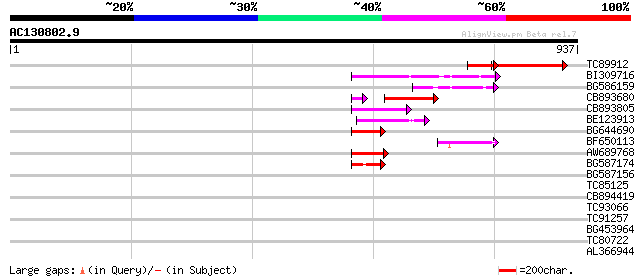
Query= AC130802.9 + phase: 0 /pseudo
(937 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC89912 weakly similar to PIR|B84512|B84512 probable retroelemen... 171 9e-43
BI309716 weakly similar to PIR|G96722|G9 hypothetical protein F2... 122 8e-28
BG586159 weakly similar to PIR|T47841|T4 hypothetical protein T2... 91 2e-18
CB893680 weakly similar to GP|1167523|db ORF(AA 1-1338) {Nicotia... 76 5e-15
CB893805 similar to GP|10177935|d copia-type polyprotein {Arabid... 73 4e-13
BE123913 weakly similar to GP|22093573|d polyprotein {Oryza sati... 67 4e-11
BG644690 weakly similar to GP|18542179|gb putative pol protein {... 60 5e-09
BF650113 weakly similar to GP|4753889|emb| Tpv2-1c {Phaseolus vu... 57 3e-08
AW689768 weakly similar to GP|10177485|d polyprotein {Arabidopsi... 53 5e-07
BG587174 similar to PIR|A47759|A4775 retrovirus-related reverse ... 47 3e-05
BG587156 similar to PIR|G85055|G8 probable polyprotein [imported... 37 0.027
TC85125 weakly similar to SP|P10978|POLX_TOBAC Retrovirus-relate... 34 0.23
CB894419 weakly similar to GP|10177935|db copia-type polyprotein... 32 1.5
TC93066 weakly similar to GP|19920130|gb|AAM08562.1 Putative ret... 31 1.9
TC91257 similar to GP|21434|emb|CAA36616.1|| ORF4 {Solanum tuber... 31 2.5
BG453964 30 3.3
TC80722 similar to GP|19699290|gb|AAL91256.1 At1g15070/F9L1_1 {A... 30 5.6
AL366944 30 5.6
>TC89912 weakly similar to PIR|B84512|B84512 probable retroelement pol
polyprotein [imported] - Arabidopsis thaliana, partial
(10%)
Length = 814
Score = 171 bits (434), Expect = 9e-43
Identities = 93/126 (73%), Positives = 101/126 (79%)
Frame = +3
Query: 796 GNVDTRKSLVGFCVYSL*HDD*LEGKSTIRGGIIYNSSGVHRTC*RCEGGHMVERDDW*V 855
G +K +GFCVYSL HD *LEGKSTIRG II NSSGVH C R E HMVER DW*V
Sbjct: 117 GQCGHKKISIGFCVYSLWHDY*LEGKSTIRGDIINNSSGVHCLCRRGERCHMVERYDW*V 296
Query: 856 RNYSRMCEDTL**PKCHSLGKSSSVS*KDKAH*HSPALCQRHD*NKRDCG*ESGIGGESG 915
RNYSR+CEDTL** KCHSLG+SSSVS*+D AH*HS AL RHD* KRDCG ++GIG ESG
Sbjct: 297 RNYSRICEDTL**SKCHSLGESSSVS*ED*AH*HSLALY*RHD*IKRDCGGKNGIGRESG 476
Query: 916 GCVHQV 921
GCV+QV
Sbjct: 477 GCVYQV 494
Score = 92.4 bits (228), Expect = 7e-19
Identities = 44/51 (86%), Positives = 47/51 (91%)
Frame = +1
Query: 757 QALKWVLRYLNGSLKGGLKYTRAAQDEDALEGYVDADYAGNVDTRKSLVGF 807
QALKWVL+YLN SLK LKYT+AAQ+EDALEGYVDADYAGNVDTRKSL GF
Sbjct: 1 QALKWVLKYLNESLKSSLKYTKAAQEEDALEGYVDADYAGNVDTRKSLSGF 153
>BI309716 weakly similar to PIR|G96722|G9 hypothetical protein F20P5.25
[imported] - Arabidopsis thaliana, partial (10%)
Length = 744
Score = 122 bits (305), Expect = 8e-28
Identities = 82/246 (33%), Positives = 133/246 (53%)
Frame = +2
Query: 565 EESLYGLKQSPREWYRRFDEFLLKTGFVRSGYDSCVYMLKKNEKVILYLLLYVDDILMAS 624
++S+YGLKQ+ R+WY + E L+ G+++S D ++ K + LL+YVDDI++A
Sbjct: 32 QKSIYGLKQASRQWYSKLSESLISFGYLQSSSDFSLFT-KFKDSSFTTLLVYVDDIVLAG 208
Query: 625 SSKDEIMKLKERLNGEFEMKDLGPAKRVLGIDIKRNRDKGELFLSQLGYLKKGGERFRMS 684
+ EI +K L F++KDLG + LG+++ R+ K + L+Q Y + E
Sbjct: 209 NDISEIQHVKCFLIDRFKIKDLGSLRYFLGLEVARS--KQGILLNQRKYTLELLEDSGNL 382
Query: 685 NSKTVSTPLGHHTKLSIQQCPQSEDEKQLMEGTPYASGVGSIMYGMVCSRPDLAYAVSIV 744
K+ TP KL P DE T Y +G ++Y + +RPD+++AV +
Sbjct: 383 AVKSTLTPYDISLKLHNSDSPLYNDE------TQYRRLIGKLIY-LTTTRPDISFAVQQL 541
Query: 745 SRFMANPGIVHWQALKWVLRYLNGSLKGGLKYTRAAQDEDALEGYVDADYAGNVDTRKSL 804
S+F++ P VH+QA VL+YL + GL Y +A L + D+D+A TRKS+
Sbjct: 542 SQFVSKPQQVHYQAAIRVLQYLKTAPAKGLFY--SATSNLKLSSFADSDWATCPTTRKSV 715
Query: 805 VGFCVY 810
G+ V+
Sbjct: 716 TGYWVF 733
>BG586159 weakly similar to PIR|T47841|T4 hypothetical protein T2O9.150 -
Arabidopsis thaliana, partial (11%)
Length = 732
Score = 91.3 bits (225), Expect = 2e-18
Identities = 55/143 (38%), Positives = 81/143 (56%), Gaps = 1/143 (0%)
Frame = +1
Query: 666 LFLSQLGYLKKGGERFRMSNSKTVSTPLGHHTKLSIQQCPQSEDEKQL-MEGTPYASGVG 724
+++ Q Y+ ERF M S P+ KL +DE + ++ T Y VG
Sbjct: 28 IYICQRKYVTDLLERFGMEKSNLSRNPIAPRCKLI-------KDENGVKVDATKYKQIVG 186
Query: 725 SIMYGMVCSRPDLAYAVSIVSRFMANPGIVHWQALKWVLRYLNGSLKGGLKYTRAAQDED 784
+MY + +RPDL Y +S++SRFM P +H A+K VLRYLNG++ G+ Y R ++
Sbjct: 187 CLMY-LAATRPDLMYVLSLISRFMNCPTELHMHAVKRVLRYLNGTINLGIMYKRNGSEK- 360
Query: 785 ALEGYVDADYAGNVDTRKSLVGF 807
LE Y D+DYAG++D RKS G+
Sbjct: 361 -LEAYTDSDYAGDLDDRKSTSGY 426
>CB893680 weakly similar to GP|1167523|db ORF(AA 1-1338) {Nicotiana tabacum},
partial (7%)
Length = 780
Score = 75.9 bits (185), Expect(2) = 5e-15
Identities = 40/89 (44%), Positives = 58/89 (64%)
Frame = -3
Query: 620 ILMASSSKDEIMKLKERLNGEFEMKDLGPAKRVLGIDIKRNRDKGELFLSQLGYLKKGGE 679
+L+ S+ DEI LK R + E +MKDLGPAK+++G+ I ++ KG L LSQ+ Y+ + +
Sbjct: 271 LLVVGSNIDEIKNLKTRFSKEIDMKDLGPAKKIIGMQIMIDKQKGVL*LSQVEYITRVLQ 92
Query: 680 RFRMSNSKTVSTPLGHHTKLSIQQCPQSE 708
F M N+ VST L H LS +Q PQ+E
Sbjct: 91 IFNMGNAILVSTTLASHFCLSHEQSPQTE 5
Score = 23.9 bits (50), Expect(2) = 5e-15
Identities = 10/27 (37%), Positives = 15/27 (55%)
Frame = -2
Query: 565 EESLYGLKQSPREWYRRFDEFLLKTGF 591
++S+YGLKQ PR+ + GF
Sbjct: 371 KKSMYGLKQGPRQCI*SLKALCTRKGF 291
>CB893805 similar to GP|10177935|d copia-type polyprotein {Arabidopsis
thaliana}, partial (14%)
Length = 778
Score = 73.2 bits (178), Expect = 4e-13
Identities = 34/100 (34%), Positives = 58/100 (58%)
Frame = +3
Query: 565 EESLYGLKQSPREWYRRFDEFLLKTGFVRSGYDSCVYMLKKNEKVILYLLLYVDDILMAS 624
+ +LYGLKQ+PR WY R + + K GF + Y+ +++ IL + LYVDD++
Sbjct: 468 KRALYGLKQAPRAWYSRIEAYFTKEGFEKCPYEHTLFVKLSEGGKILIISLYVDDLIFIG 647
Query: 625 SSKDEIMKLKERLNGEFEMKDLGPAKRVLGIDIKRNRDKG 664
+ ++ + K+ + EF M DLG LG+++ +N +KG
Sbjct: 648 NDENMFEEFKKSMKKEFNMSDLGKMHYFLGVEVTQN-EKG 764
>BE123913 weakly similar to GP|22093573|d polyprotein {Oryza sativa (japonica
cultivar-group)}, partial (8%)
Length = 503
Score = 66.6 bits (161), Expect = 4e-11
Identities = 42/122 (34%), Positives = 65/122 (52%)
Frame = +1
Query: 573 QSPREWYRRFDEFLLKTGFVRSGYDSCVYMLKKNEKVILYLLLYVDDILMASSSKDEIMK 632
QSPR+W+ RF + K G+++ D +++ + L++YVDDI + I +
Sbjct: 1 QSPRDWFDRFT*VVKKFGYIQCQTDHAMFIKHSSTVKKAILIVYVDDIFLTGDHGK*IKR 180
Query: 633 LKERLNGEFEMKDLGPAKRVLGIDIKRNRDKGELFLSQLGYLKKGGERFRMSNSKTVSTP 692
LK L EFE+KDLG K LG+++ R + KG +SQ Y+ + RM KT+ P
Sbjct: 181 LKNLLAEEFEIKDLGNLKYFLGMEVARWK-KGS-SISQRKYVLDLLKETRMIGCKTIRDP 354
Query: 693 LG 694
G
Sbjct: 355 YG 360
>BG644690 weakly similar to GP|18542179|gb putative pol protein {Zea mays},
partial (22%)
Length = 629
Score = 59.7 bits (143), Expect = 5e-09
Identities = 28/55 (50%), Positives = 40/55 (71%)
Frame = -2
Query: 566 ESLYGLKQSPREWYRRFDEFLLKTGFVRSGYDSCVYMLKKNEKVILYLLLYVDDI 620
++LYGLKQ+PR WY R +FLLK GF R D+ +++LK+ E +L + +YVDDI
Sbjct: 163 KTLYGLKQAPRAWYERLSKFLLKNGFKRGKIDNTLFLLKR-E*ELLIIQVYVDDI 2
>BF650113 weakly similar to GP|4753889|emb| Tpv2-1c {Phaseolus vulgaris},
partial (13%)
Length = 494
Score = 57.4 bits (137), Expect = 3e-08
Identities = 37/108 (34%), Positives = 61/108 (56%), Gaps = 8/108 (7%)
Frame = +1
Query: 708 EDEKQLMEGTPYASGVG------SIMYGM-VCSRPDLAYAVSIVSRFMANPGIVHWQALK 760
E E+ L +G P S +G + G+ +C RPD+ Y+VS++S+FM +P H A
Sbjct: 22 EQEQGLEKGEPQESRIGVVRS*IEVQVGINLC*RPDICYSVSVISKFMHDPRKPHLIAAN 201
Query: 761 WVLRYLNGSLKGGLKYTRAAQDE-DALEGYVDADYAGNVDTRKSLVGF 807
+LRY+ G+++ GL + A+ E L Y D+D+ G+ R+S G+
Sbjct: 202 RILRYVRGTMEYGLLFPYGAKSEVYELICYSDSDWCGD---RRSTSGY 336
>AW689768 weakly similar to GP|10177485|d polyprotein {Arabidopsis thaliana},
partial (9%)
Length = 675
Score = 53.1 bits (126), Expect = 5e-07
Identities = 27/61 (44%), Positives = 37/61 (60%)
Frame = +1
Query: 566 ESLYGLKQSPREWYRRFDEFLLKTGFVRSGYDSCVYMLKKNEKVILYLLLYVDDILMASS 625
+SLYGLKQ+PR WY ++ GF +S D + + +N I YL +YVDDIL+ S
Sbjct: 490 KSLYGLKQAPRAWYEXLTSAQIQFGFTKSRCDPSLLIYNQNGACI-YLXIYVDDILITGS 666
Query: 626 S 626
S
Sbjct: 667 S 669
>BG587174 similar to PIR|A47759|A4775 retrovirus-related reverse
transcriptase homolog - rape retrotransposon copia-like
(fragment), partial (84%)
Length = 249
Score = 47.4 bits (111), Expect = 3e-05
Identities = 25/58 (43%), Positives = 38/58 (65%), Gaps = 2/58 (3%)
Frame = -1
Query: 566 ESLYGLKQSPREWYRRFDEFLLKTGFVRSGYDSCVYMLKKNEKV--ILYLLLYVDDIL 621
+SLYGLKQSPR+WY+RFD + ++ + +G V ++ +YL+LYVDD+L
Sbjct: 168 KSLYGLKQSPRQWYKRFDSY--RSSWATTGVLMTVVST*TR*RMSRYIYLVLYVDDML 1
>BG587156 similar to PIR|G85055|G8 probable polyprotein [imported] -
Arabidopsis thaliana, partial (17%)
Length = 618
Score = 37.4 bits (85), Expect = 0.027
Identities = 16/39 (41%), Positives = 24/39 (61%)
Frame = -1
Query: 565 EESLYGLKQSPREWYRRFDEFLLKTGFVRSGYDSCVYML 603
++++YGLKQSPR WY + L GF +S D ++ L
Sbjct: 129 KKAIYGLKQSPRAWYNKLSTTLNGRGFRKSELDHTLFTL 13
>TC85125 weakly similar to SP|P10978|POLX_TOBAC Retrovirus-related Pol
polyprotein from transposon TNT 1-94 [Contains: Protease
(EC 3.4.23.-);, partial (7%)
Length = 705
Score = 34.3 bits (77), Expect = 0.23
Identities = 17/49 (34%), Positives = 29/49 (58%)
Frame = +1
Query: 759 LKWVLRYLNGSLKGGLKYTRAAQDEDALEGYVDADYAGNVDTRKSLVGF 807
+K ++RY+ G+ + + E + GYVD+D+AG+ D RKS G+
Sbjct: 1 VKRIMRYIKGTSGVAVCF---GGSELTVRGYVDSDFAGDHDKRKSTTGY 138
>CB894419 weakly similar to GP|10177935|db copia-type polyprotein
{Arabidopsis thaliana}, partial (2%)
Length = 170
Score = 31.6 bits (70), Expect = 1.5
Identities = 16/50 (32%), Positives = 29/50 (58%)
Frame = +3
Query: 679 ERFRMSNSKTVSTPLGHHTKLSIQQCPQSEDEKQLMEGTPYASGVGSIMY 728
++F+M +SK +STP+ KL+ E + + ++ T Y S +GS+ Y
Sbjct: 27 KKFKMEHSKPISTPVEEKLKLT------RESDGKRVDSTHYKSLIGSLRY 158
>TC93066 weakly similar to GP|19920130|gb|AAM08562.1 Putative retroelement
{Oryza sativa} [Oryza sativa (japonica cultivar-group)],
partial (10%)
Length = 823
Score = 31.2 bits (69), Expect = 1.9
Identities = 35/121 (28%), Positives = 53/121 (42%)
Frame = +3
Query: 105 AFGSLGSF*DSYSWGRFLFSFYH**LL*ESVGLCFEKQK*HL*KVQRMAHSHRKSNGN*T 164
+F LG+F + W L+ YH** E +GL F K* +Q + +S S+
Sbjct: 108 SF*PLGTFKSYFLWRTPLYDDYH**FSSEGLGLFFAV*K*DFSHIQEVENSC*NSDREEC 287
Query: 165 KRFKN*QWPGVCFRAV**VLQVERNQEA*NRTKNTATKWPCGTHE*DTFGACEVYDSRSW 224
+ N V +**VL N +K + TK C T++ D+ +Y + W
Sbjct: 288 EEAHNR*LIRVL***L**VLHKSWYC*TQNHSKESPTKRCCRTNDQDST*ESSMYALKCW 467
Query: 225 V 225
V
Sbjct: 468 V 470
>TC91257 similar to GP|21434|emb|CAA36616.1|| ORF4 {Solanum tuberosum},
partial (8%)
Length = 854
Score = 30.8 bits (68), Expect = 2.5
Identities = 14/30 (46%), Positives = 22/30 (72%)
Frame = +3
Query: 719 YASGVGSIMYGMVCSRPDLAYAVSIVSRFM 748
Y VG + Y + +RPD++YAVS+VS+F+
Sbjct: 762 YRRLVGKLNY-LTMTRPDISYAVSVVSQFL 848
>BG453964
Length = 647
Score = 30.4 bits (67), Expect = 3.3
Identities = 17/50 (34%), Positives = 31/50 (62%)
Frame = -2
Query: 611 LYLLLYVDDILMASSSKDEIMKLKERLNGEFEMKDLGPAKRVLGIDIKRN 660
+YLL+Y+D+IL+ S ++ L++ L+ F+ KDL K GI + ++
Sbjct: 202 IYLLVYIDEILL-SIVIIPLVCLRQHLSNHFQTKDLDLFKYFSGIVVAQS 56
>TC80722 similar to GP|19699290|gb|AAL91256.1 At1g15070/F9L1_1 {Arabidopsis
thaliana}, partial (16%)
Length = 1001
Score = 29.6 bits (65), Expect = 5.6
Identities = 12/43 (27%), Positives = 21/43 (47%)
Frame = +3
Query: 731 VCSRPDLAYAVSIVSRFMANPGIVHWQALKWVLRYLNGSLKGG 773
VC + L + V + R ++ +Q W+L Y+N S + G
Sbjct: 663 VCQKESLGFLVKVKQRSCLPNYLIRFQNFWWILHYMNNSTR*G 791
>AL366944
Length = 275
Score = 29.6 bits (65), Expect = 5.6
Identities = 15/38 (39%), Positives = 22/38 (57%)
Frame = -3
Query: 734 RPDLAYAVSIVSRFMANPGIVHWQALKWVLRYLNGSLK 771
RP AVS+VS+F + H L W+L+Y+ G+ K
Sbjct: 117 RPKHFIAVSLVSQF*NSSSQRHKDVLIWMLKYIEGARK 4
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.356 0.161 0.595
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 28,415,973
Number of Sequences: 36976
Number of extensions: 414619
Number of successful extensions: 3350
Number of sequences better than 10.0: 36
Number of HSP's better than 10.0 without gapping: 981
Number of HSP's successfully gapped in prelim test: 104
Number of HSP's that attempted gapping in prelim test: 2295
Number of HSP's gapped (non-prelim): 1185
length of query: 937
length of database: 9,014,727
effective HSP length: 105
effective length of query: 832
effective length of database: 5,132,247
effective search space: 4270029504
effective search space used: 4270029504
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 14 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 37 (21.6 bits)
S2: 63 (28.9 bits)
Medicago: description of AC130802.9


